Gene
KWMTBOMO10620 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA003945
Annotation
cytochrome_P450_9a20_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 2.05 PlasmaMembrane Reliability : 1.502
Sequence
CDS
ATGATTCTCCTCATTTGGGCCGTCGTGCTCATTGCCGCCTTCGTGCTGTTTTACAAACAAGCCTATTCTTTGTTCAGCAAGCACGGAGTGAAGGGCTTCACTCCGCTGCCTTTCTTTGGCAACATGGGGCGTATTGTAATAAAAATGGATCATTTCTCGGACCATATACAAAGTTTATACGATTCATTCCCAGAGGAAAGGTTCGTCGGAAGATACGAATTTCTAAACCCAATGGTTATAATTCGTGACATTGAGTTACTGAAGAAAATCACGGTTAAGGACTTCGAGCATTTCCTGGACCATCGGACTATCATCAACAAGGACACCGACCCGTTCTTCGGAAGGAGCTTGTTCTTCTTGAGAGATCAAGACTGGAAAGACATGCGCTCAACGCTTTCGCCGGCGTTCACCAGTTCCAAAATGAAGCTCATGATGCCCTTCATCGTTGAAGTGGGAGAACAGATGAATAAAGCGCTAAAGCAGAGGATACAAGAAGCAGGGGTTGGCTATGTGGACATCGACTCCAAAGACCTGACCACTAGATACGCCAACGATGTGATCGCGTCCTGCGCCTTCGGTCTGAAGGTCGACTCCATCACCGAGGAGAACAACCAGTTCTACGCGATGGGGAAAGCCGCTTCCACTTTCAACTTCAGGCAGCTTCTAATTTTCTTTGGTCTCGCATCTGTCCCGAAGCTTGTGAAGATTCTACGCATTACGCTGTTTCAAAAAGAAATCAAAACCTTCTTCAGGGAGCTGATCTTGGGTACCATGAAGAACAGGGAAGCACAAAACATTATCAGACCTGACATGATCCATCTACTCATGGAAGCTAAGAAAGGCAAGTTGAGGCACGATGAGAAATCCACAAAAGACAGCGACGCTGGCTTTGCTACCGTGGAAGAGTCCTCAGTCGGCAAAAAGGATATTAACCGAGTGTGGACAGACGATGACTTGGTCGCCCAAGCGGTTCTGTTTTTCGTCGCCGGCTTCGAAACAGTATCGTCAGCGATGACATTCCTGCTTCACGAGTTGGCTTTGAACCCTGAAGTGCAGGAGAAGCTGGTGGAAGAAATCCGAGAAAACGAGAAAAACAACAACGGAAAATTCGACTACAACTCCATTCAGAACATGGTGTATTTAGATATGGTGGTGTCAGAGGTCTTAAGATTGTGGCCACCTGTTATTGCCTTAGATAGAATGTGCGTTAAAGACTACAACTTGGGTAAACCAAATGACAAAAGCAAAGAAGATTTTATTATACGCAAAGATGTGGCGGTGGGTATACCAGTATGGGGCTTACATCGAGATCCCGAATTCTTCCCGAACCCCCTGAAATTCGATCCCGAAAGGTTCTCAGAGGAGAACAAACACAACATCAAACCCTTCAGTTACATGCCATTTGGTCTGGGACCTAGGAATTGTATAGGTTCAAGGTTCGCTTTGTGCGAAGTCAAGGTGATGACTTACCAACTCCTCCAGCACATGGAGATATCTCCGTGCGAAAAGACCTGCATTCCCTCGAAACTCAGCAAGGAGACTTTCAACCTTCGCCTTGAAGGCGGTCATTGGATCAGGCTGAAGATTAGAAATTGA
Protein
MILLIWAVVLIAAFVLFYKQAYSLFSKHGVKGFTPLPFFGNMGRIVIKMDHFSDHIQSLYDSFPEERFVGRYEFLNPMVIIRDIELLKKITVKDFEHFLDHRTIINKDTDPFFGRSLFFLRDQDWKDMRSTLSPAFTSSKMKLMMPFIVEVGEQMNKALKQRIQEAGVGYVDIDSKDLTTRYANDVIASCAFGLKVDSITEENNQFYAMGKAASTFNFRQLLIFFGLASVPKLVKILRITLFQKEIKTFFRELILGTMKNREAQNIIRPDMIHLLMEAKKGKLRHDEKSTKDSDAGFATVEESSVGKKDINRVWTDDDLVAQAVLFFVAGFETVSSAMTFLLHELALNPEVQEKLVEEIRENEKNNNGKFDYNSIQNMVYLDMVVSEVLRLWPPVIALDRMCVKDYNLGKPNDKSKEDFIIRKDVAVGIPVWGLHRDPEFFPNPLKFDPERFSEENKHNIKPFSYMPFGLGPRNCIGSRFALCEVKVMTYQLLQHMEISPCEKTCIPSKLSKETFNLRLEGGHWIRLKIRN
Summary
Similarity
Belongs to the cytochrome P450 family.
Uniprot
H9J358
A3RKJ5
D0FZ10
L0N4J7
L0N797
L0N725
+ More
B6VEL8 Q1KLD7 H9J356 L0N6I2 A5HMQ7 A5HKM0 A3RIC2 A4UU25 B6V7E2 L0N4K3 A5HKL9 Q5YJK0 C6H0J5 A0A068F0Y7 Q4ZJH3 Q5XLI9 A0A2W1BL52 B7TYK9 B2CPL0 A0A385LJM0 A0A068EU83 A0A1W5ZJN7 A0A0K8TUK1 A0A0P0EW60 B2CPL1 A4KAE0 A9Z0U8 V5JDC8 A0A2W1BKQ5 A0A0C5CGW0 A0A248QFE0 A0A2A4JDS7 Q6UEI0 A0A2A4JDS0 X5DAZ5 A0A3G5EAT6 A0A2D1QTX7 A0A2D1QTX0 A0A248QJ18 B5MF67 A0A385LJV6 Q7YZX4 D2JLK7 A0A3S2M2T6 A0A068ETV5 A9Z0U9 A0A2W1BPX7 B6ECI4 A0A0K0YD31 C9E6N0 A4KAE1 A0A194QJQ1 Q6RW46 A0A3S2NVP0 X5D4G9 A0A068EXP4 A0A0K8TUK8 A0A194R3L1 A0A194QDY1 A0A1W5ZJJ8 A0A194QDQ3 A5HKL6 A5HKL8 A4ZWB1 D0VY60 A5HKL7 A0A286MXM7 A0A2D1QTX1 D2JLJ7 A0A1V0D9E5 A0A385LJP1 A0A0L7L0K4 A0A194R4E5 A0A068EUU6 L0N5G5 A0A385LJT9 A0A248QEG8 A0A068ETV9 A0A248QEG5 A0A212F932 Q9U9C1 A0A2W1BR50 Q2XSW0 E5L8C3 A0A2A4JJX9 A0A2A4JEN6 A0A068EVL9 A0A2A4JJV4 A0A2A4JJD6 A0A2W1BSJ6 W0FSV4 A0A068EWN1 A0A0K0YD28 A0A1V0D9E7
B6VEL8 Q1KLD7 H9J356 L0N6I2 A5HMQ7 A5HKM0 A3RIC2 A4UU25 B6V7E2 L0N4K3 A5HKL9 Q5YJK0 C6H0J5 A0A068F0Y7 Q4ZJH3 Q5XLI9 A0A2W1BL52 B7TYK9 B2CPL0 A0A385LJM0 A0A068EU83 A0A1W5ZJN7 A0A0K8TUK1 A0A0P0EW60 B2CPL1 A4KAE0 A9Z0U8 V5JDC8 A0A2W1BKQ5 A0A0C5CGW0 A0A248QFE0 A0A2A4JDS7 Q6UEI0 A0A2A4JDS0 X5DAZ5 A0A3G5EAT6 A0A2D1QTX7 A0A2D1QTX0 A0A248QJ18 B5MF67 A0A385LJV6 Q7YZX4 D2JLK7 A0A3S2M2T6 A0A068ETV5 A9Z0U9 A0A2W1BPX7 B6ECI4 A0A0K0YD31 C9E6N0 A4KAE1 A0A194QJQ1 Q6RW46 A0A3S2NVP0 X5D4G9 A0A068EXP4 A0A0K8TUK8 A0A194R3L1 A0A194QDY1 A0A1W5ZJJ8 A0A194QDQ3 A5HKL6 A5HKL8 A4ZWB1 D0VY60 A5HKL7 A0A286MXM7 A0A2D1QTX1 D2JLJ7 A0A1V0D9E5 A0A385LJP1 A0A0L7L0K4 A0A194R4E5 A0A068EUU6 L0N5G5 A0A385LJT9 A0A248QEG8 A0A068ETV9 A0A248QEG5 A0A212F932 Q9U9C1 A0A2W1BR50 Q2XSW0 E5L8C3 A0A2A4JJX9 A0A2A4JEN6 A0A068EVL9 A0A2A4JJV4 A0A2A4JJD6 A0A2W1BSJ6 W0FSV4 A0A068EWN1 A0A0K0YD28 A0A1V0D9E7
Pubmed
EMBL
BABH01036637
BABH01036638
EF421989
ABO07439.1
AB498807
BAI47532.1
+ More
AK289297 BAM73824.1 AK289298 BAM73825.1 AK289299 BAM73826.1 FJ378716 ACJ05915.1 DQ473438 ABF14737.1 BABH01036639 BABH01036640 AK289300 BAM73827.1 EF542811 ABQ18318.1 EF535808 ABQ08710.1 EF415298 ABN71369.1 EF488994 ABP02071.1 FJ265741 ACJ04711.1 AK289302 BAM73829.1 EF535807 ABQ08709.1 AY390260 AAR26518.1 FN421128 CAZ65619.1 KM016753 AID54905.1 DQ003275 AAY21809.1 AY753201 AAV28704.1 KZ149989 PZC75618.1 FJ416332 ACJ37388.1 EU541247 ACB30272.1 MG793355 AYA28074.1 KM016752 AID54904.1 KY436738 ARI68318.1 GCVX01000068 JAI18162.1 KR676343 ALJ30298.1 EU541248 ACB30273.2 DQ788839 ABH09252.1 EU327673 ABY47595.1 JX486677 AGI65181.1 PZC75619.1 KP001132 AJN91177.1 KX443437 ASO98012.1 NWSH01001758 PCG70237.1 AY371318 AAQ73544.1 PCG70235.1 KF701145 AHW57315.1 MH001162 AYW35335.1 KY348420 ATP15901.1 KY348419 ATP15900.1 KX443436 ASO98011.1 AB381883 BAG71410.1 MG793354 AYA28073.1 AF525031 AAP80766.1 GQ915323 ACZ97417.2 RSAL01000054 RVE50006.1 KM016750 AID54902.1 EU327674 ABY47596.1 PZC75615.1 FJ227150 ACI43222.1 KR095603 AKS48891.1 GQ465040 ACV88722.1 DQ788840 ABH09253.1 KQ459144 KPJ03681.1 AY487948 AAR37015.1 RVE50007.1 KF701146 AHW57316.1 KJ671577 AID55429.1 GCVX01000063 JAI18167.1 KQ460779 KPJ12393.1 KPJ03682.1 KY436739 ARI68319.1 KPJ03683.1 EF535804 ABQ08706.1 EF535806 ABQ08708.2 BABH01036652 BABH01036653 EF491004 AK343209 ABP65279.1 BAM73893.1 AB500115 BAI49180.1 EF535805 ABQ08707.1 MF684345 ASX93979.1 KY348418 ATP15899.1 GQ915313 ACZ97407.2 KY212055 ARA91619.1 MG793356 AYA28075.1 JTDY01003828 KOB68945.1 KPJ12394.1 KJ671578 AID55430.1 AK289301 BAM73828.1 MG793353 AYA28072.1 KX443440 ASO98015.1 KM016755 AID54907.1 KX443439 ASO98014.1 AGBW02009651 OWR50238.1 AF172279 AAD51036.1 PZC75617.1 DQ256408 ABB69055.1 HQ340240 KX443438 ADP55210.1 ASO98013.1 NWSH01001322 PCG71702.1 PCG70236.1 KM016751 AID54903.1 PCG71703.1 PCG71704.1 PZC75616.1 KC832920 AHF45921.1 KJ671579 AID55431.1 KR095604 AKS48892.1 KY212054 ARA91618.1
AK289297 BAM73824.1 AK289298 BAM73825.1 AK289299 BAM73826.1 FJ378716 ACJ05915.1 DQ473438 ABF14737.1 BABH01036639 BABH01036640 AK289300 BAM73827.1 EF542811 ABQ18318.1 EF535808 ABQ08710.1 EF415298 ABN71369.1 EF488994 ABP02071.1 FJ265741 ACJ04711.1 AK289302 BAM73829.1 EF535807 ABQ08709.1 AY390260 AAR26518.1 FN421128 CAZ65619.1 KM016753 AID54905.1 DQ003275 AAY21809.1 AY753201 AAV28704.1 KZ149989 PZC75618.1 FJ416332 ACJ37388.1 EU541247 ACB30272.1 MG793355 AYA28074.1 KM016752 AID54904.1 KY436738 ARI68318.1 GCVX01000068 JAI18162.1 KR676343 ALJ30298.1 EU541248 ACB30273.2 DQ788839 ABH09252.1 EU327673 ABY47595.1 JX486677 AGI65181.1 PZC75619.1 KP001132 AJN91177.1 KX443437 ASO98012.1 NWSH01001758 PCG70237.1 AY371318 AAQ73544.1 PCG70235.1 KF701145 AHW57315.1 MH001162 AYW35335.1 KY348420 ATP15901.1 KY348419 ATP15900.1 KX443436 ASO98011.1 AB381883 BAG71410.1 MG793354 AYA28073.1 AF525031 AAP80766.1 GQ915323 ACZ97417.2 RSAL01000054 RVE50006.1 KM016750 AID54902.1 EU327674 ABY47596.1 PZC75615.1 FJ227150 ACI43222.1 KR095603 AKS48891.1 GQ465040 ACV88722.1 DQ788840 ABH09253.1 KQ459144 KPJ03681.1 AY487948 AAR37015.1 RVE50007.1 KF701146 AHW57316.1 KJ671577 AID55429.1 GCVX01000063 JAI18167.1 KQ460779 KPJ12393.1 KPJ03682.1 KY436739 ARI68319.1 KPJ03683.1 EF535804 ABQ08706.1 EF535806 ABQ08708.2 BABH01036652 BABH01036653 EF491004 AK343209 ABP65279.1 BAM73893.1 AB500115 BAI49180.1 EF535805 ABQ08707.1 MF684345 ASX93979.1 KY348418 ATP15899.1 GQ915313 ACZ97407.2 KY212055 ARA91619.1 MG793356 AYA28075.1 JTDY01003828 KOB68945.1 KPJ12394.1 KJ671578 AID55430.1 AK289301 BAM73828.1 MG793353 AYA28072.1 KX443440 ASO98015.1 KM016755 AID54907.1 KX443439 ASO98014.1 AGBW02009651 OWR50238.1 AF172279 AAD51036.1 PZC75617.1 DQ256408 ABB69055.1 HQ340240 KX443438 ADP55210.1 ASO98013.1 NWSH01001322 PCG71702.1 PCG70236.1 KM016751 AID54903.1 PCG71703.1 PCG71704.1 PZC75616.1 KC832920 AHF45921.1 KJ671579 AID55431.1 KR095604 AKS48892.1 KY212054 ARA91618.1
Proteomes
PRIDE
Pfam
PF00067 p450
Interpro
SUPFAM
SSF48264
SSF48264
Gene 3D
ProteinModelPortal
H9J358
A3RKJ5
D0FZ10
L0N4J7
L0N797
L0N725
+ More
B6VEL8 Q1KLD7 H9J356 L0N6I2 A5HMQ7 A5HKM0 A3RIC2 A4UU25 B6V7E2 L0N4K3 A5HKL9 Q5YJK0 C6H0J5 A0A068F0Y7 Q4ZJH3 Q5XLI9 A0A2W1BL52 B7TYK9 B2CPL0 A0A385LJM0 A0A068EU83 A0A1W5ZJN7 A0A0K8TUK1 A0A0P0EW60 B2CPL1 A4KAE0 A9Z0U8 V5JDC8 A0A2W1BKQ5 A0A0C5CGW0 A0A248QFE0 A0A2A4JDS7 Q6UEI0 A0A2A4JDS0 X5DAZ5 A0A3G5EAT6 A0A2D1QTX7 A0A2D1QTX0 A0A248QJ18 B5MF67 A0A385LJV6 Q7YZX4 D2JLK7 A0A3S2M2T6 A0A068ETV5 A9Z0U9 A0A2W1BPX7 B6ECI4 A0A0K0YD31 C9E6N0 A4KAE1 A0A194QJQ1 Q6RW46 A0A3S2NVP0 X5D4G9 A0A068EXP4 A0A0K8TUK8 A0A194R3L1 A0A194QDY1 A0A1W5ZJJ8 A0A194QDQ3 A5HKL6 A5HKL8 A4ZWB1 D0VY60 A5HKL7 A0A286MXM7 A0A2D1QTX1 D2JLJ7 A0A1V0D9E5 A0A385LJP1 A0A0L7L0K4 A0A194R4E5 A0A068EUU6 L0N5G5 A0A385LJT9 A0A248QEG8 A0A068ETV9 A0A248QEG5 A0A212F932 Q9U9C1 A0A2W1BR50 Q2XSW0 E5L8C3 A0A2A4JJX9 A0A2A4JEN6 A0A068EVL9 A0A2A4JJV4 A0A2A4JJD6 A0A2W1BSJ6 W0FSV4 A0A068EWN1 A0A0K0YD28 A0A1V0D9E7
B6VEL8 Q1KLD7 H9J356 L0N6I2 A5HMQ7 A5HKM0 A3RIC2 A4UU25 B6V7E2 L0N4K3 A5HKL9 Q5YJK0 C6H0J5 A0A068F0Y7 Q4ZJH3 Q5XLI9 A0A2W1BL52 B7TYK9 B2CPL0 A0A385LJM0 A0A068EU83 A0A1W5ZJN7 A0A0K8TUK1 A0A0P0EW60 B2CPL1 A4KAE0 A9Z0U8 V5JDC8 A0A2W1BKQ5 A0A0C5CGW0 A0A248QFE0 A0A2A4JDS7 Q6UEI0 A0A2A4JDS0 X5DAZ5 A0A3G5EAT6 A0A2D1QTX7 A0A2D1QTX0 A0A248QJ18 B5MF67 A0A385LJV6 Q7YZX4 D2JLK7 A0A3S2M2T6 A0A068ETV5 A9Z0U9 A0A2W1BPX7 B6ECI4 A0A0K0YD31 C9E6N0 A4KAE1 A0A194QJQ1 Q6RW46 A0A3S2NVP0 X5D4G9 A0A068EXP4 A0A0K8TUK8 A0A194R3L1 A0A194QDY1 A0A1W5ZJJ8 A0A194QDQ3 A5HKL6 A5HKL8 A4ZWB1 D0VY60 A5HKL7 A0A286MXM7 A0A2D1QTX1 D2JLJ7 A0A1V0D9E5 A0A385LJP1 A0A0L7L0K4 A0A194R4E5 A0A068EUU6 L0N5G5 A0A385LJT9 A0A248QEG8 A0A068ETV9 A0A248QEG5 A0A212F932 Q9U9C1 A0A2W1BR50 Q2XSW0 E5L8C3 A0A2A4JJX9 A0A2A4JEN6 A0A068EVL9 A0A2A4JJV4 A0A2A4JJD6 A0A2W1BSJ6 W0FSV4 A0A068EWN1 A0A0K0YD28 A0A1V0D9E7
PDB
6MA7
E-value=7.2895e-71,
Score=680
Ontologies
KEGG
GO
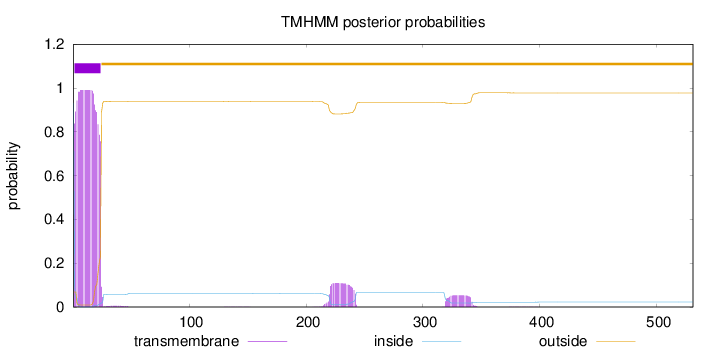
Topology
Length:
531
Number of predicted TMHs:
1
Exp number of AAs in TMHs:
25.39055
Exp number, first 60 AAs:
21.82008
Total prob of N-in:
0.93192
POSSIBLE N-term signal
sequence
inside
1 - 1
TMhelix
2 - 24
outside
25 - 531
Population Genetic Test Statistics
Pi
199.855508
Theta
176.83155
Tajima's D
0.911777
CLR
0.16289
CSRT
0.630518474076296
Interpretation
Uncertain
