Gene
KWMTBOMO10589
Pre Gene Modal
BGIBMGA003915
Annotation
PREDICTED:_LOW_QUALITY_PROTEIN:_monocarboxylate_transporter_14-like_[Bombyx_mori]
Location in the cell
PlasmaMembrane Reliability : 4.676
Sequence
CDS
ATGGACACGGAACGTACATATGCAAATGGAGTCGCCGGTAATGAGAAGTATCTCAAGTACGAGCATAAAGTTCCGTCTGAAGAGGAGAAGACTACGTCTCCTTTATTAGAGAAGGACAATATAGAGTTTATTGATGGTGTAGATTTGAAGAAAGATGAGGGAACTCAAGATTCAGATAGAAAAGATGACGGAAATAACTTGACGGTTGGTTACTATAAAGACGGGAAAGAAGATGATGCGACATCAACTAATGAAGGTGTAAAGTTCTTTGGGCTAGGAGATGCAGATAGTATATGTTCTAGTGGACCACCGCCGGAGTCAACTCCGCAAATACCAGATGGCGGTTGGGGCTGGGTGGTTGTGGCTGCTTCATTTCTAGTAGCGACAGTAGCAGACGGCCTGGCATTTTCTTACGGGCTGATAAATGACAAGTTCGTCCTTTACTTCGAGACCAGCGAAGCGAAAACTTCGCTTATCGGAAGTTTGTTTATATCCGTTCCATTGATCGCTGGGCCAATTATGAGTGCTTTAGTCGACAGATACGGATGCAAGAATATGACTATAGTTGGCGGGGTTGCGTCTACAATCGGATTCGTTGCTGCATCATTCAGCAATTCCGTTGAAGTCTTATATGTTACTTATGGTATAATGGCTGGATTAGGTATGGGACTATTGTATGTTACAGCTGTCGTATCAATAGCGTTTTGGTTTGAAAAGCGCCGTAACTTAGCCGTCGGGCTCGGCTCGTGCGGAGTTGGCTTTGGAACATTTATTTACTCTCCCTTGACGACCTACTTACTTGATGAATACGGTTGGAGAGGTGCCTTACTACTACTCGCTGGCACGGTACTAAACGTGTGCGTGTGTGGTATGCTCATGAGGGACCCTGAATGGCTGATTCTGGAACAAAAAAAGCAAAGACAGTTGAGTAAGGCAAAGAGAGCTTCCAGTTCTATGTCGATCTCGGCTAGATCTGCAAAGTCTGGTGGAGCGGAGTCCATGTACCCTGGAGTCGAGGAAATTAAGACAATGATAAAGAGTGGTGAGACACCGGAATATATCCTAACGGCATTAGTCGCGTCGATTGCCGAAGCGGAAAGCTTGGAGGCTGCTACAAAAGCTAATGCGGATCTTTCCAACCATAAAGTTAGCTCAGTGATAAATTTACCTACGTTTTTGAAGCAAAGCGAAAAGGTACCAGCAGAAGTATTAGACCAGCTTATATCAAACAAGCGACTTTATAACATAGTATTACAGAATTATCCATCATTACTTGCACTAAGGAGTAATTCCGAACAGAAACTACCCACAGAACCGCCCCCGTCCAAAGCGAAACCAGTGAAAATGTCCATGAAGCTGAAACTCAGTAAGAAAGAGAAGAAAGACTTTGCATCTAAGCTGGACGTTGTGAAGGAGAAGTTACTGCAACCGATACCAGAAAACAAGGTCATAATACCGAAAGCTGAACGGCAGGCTTGGTTCTCGAGACAGATCCAAACCGATCATCATTATTTCAGAGAGATAAGGGTGCACAGGAATTCGATAATGCATCGCGGGGCAATGATGAACATTGCCAAGTACAAACTCCGTGCGTCGTCTTGTCCAGATATTTATAGAAATTCTATGTGGTCCGTTGAAGACCACGAGGAAAAGACATGGGGACGACGACTAATAGACACGATCAACAAAACATTTGATTTCAACATGTTCACGGAGTTCCATTTCCTAATGATGAACTTATCCACGCTGGTGCTGTTCATATGGTTCATAGTACCTTACTTCTACATATCCAGTTTCATGCAGATGCGGGGCTACTCCGAAGTAGACGGCTCTTGGTTACTGAGCACTTTTGGCATTGCTACCATAATTGGAATAGTGGGCCTCGGTTGGATGGGCGATCTGCCCTGGGTGAACGTGACCAAGACTTACGCGATATGCCTCGTATTCTGCGGCATATCGATAGTAATGTTCCCGGTACTGATACGAGTATTGGATCCGGCATCGCCGGCCAGTTTCTACGTTCTAGCCATCAACGCGGCGATTTTTGGATTGATGTTTTCCAGTTCATATTCTTACACGCCCAGTATATTGGTGGAGCTGATTGCGCTTGAAAGGTTTACTATGGCGTATGGCCTGGTGCTACTCAGTCAGGGCGTTGGCCATCTGATTGGTCCCCCTATAGCTGGAGCTCTCAAAGATTCGACCGGCCAATGGGACGCGGCGTTCTACTTGGCTGGTATATGGGTGATCGTTTCCGGTGTCCTTGTTGGCGTCATACCTTACACCCAGAACTTCAGGATGTGCGGCAACGCGCCCCTGGCCAAGGACGCGGCCGGCGAGCCTGACCCTGGAATCAAAATTATTATAGCACATTAA
Protein
MDTERTYANGVAGNEKYLKYEHKVPSEEEKTTSPLLEKDNIEFIDGVDLKKDEGTQDSDRKDDGNNLTVGYYKDGKEDDATSTNEGVKFFGLGDADSICSSGPPPESTPQIPDGGWGWVVVAASFLVATVADGLAFSYGLINDKFVLYFETSEAKTSLIGSLFISVPLIAGPIMSALVDRYGCKNMTIVGGVASTIGFVAASFSNSVEVLYVTYGIMAGLGMGLLYVTAVVSIAFWFEKRRNLAVGLGSCGVGFGTFIYSPLTTYLLDEYGWRGALLLLAGTVLNVCVCGMLMRDPEWLILEQKKQRQLSKAKRASSSMSISARSAKSGGAESMYPGVEEIKTMIKSGETPEYILTALVASIAEAESLEAATKANADLSNHKVSSVINLPTFLKQSEKVPAEVLDQLISNKRLYNIVLQNYPSLLALRSNSEQKLPTEPPPSKAKPVKMSMKLKLSKKEKKDFASKLDVVKEKLLQPIPENKVIIPKAERQAWFSRQIQTDHHYFREIRVHRNSIMHRGAMMNIAKYKLRASSCPDIYRNSMWSVEDHEEKTWGRRLIDTINKTFDFNMFTEFHFLMMNLSTLVLFIWFIVPYFYISSFMQMRGYSEVDGSWLLSTFGIATIIGIVGLGWMGDLPWVNVTKTYAICLVFCGISIVMFPVLIRVLDPASPASFYVLAINAAIFGLMFSSSYSYTPSILVELIALERFTMAYGLVLLSQGVGHLIGPPIAGALKDSTGQWDAAFYLAGIWVIVSGVLVGVIPYTQNFRMCGNAPLAKDAAGEPDPGIKIIIAH
Summary
Uniprot
H9J328
A0A2W1BRL4
A0A2A4IUF1
A0A2H1V1L2
A0A3S2NAV0
A0A194PK99
+ More
A0A194RIL6 A0A212EL73 A0A0N0P9U3 A0A0L7KU96 A0A194QG55 A0A2A4JUI4 A0A2W1BN40 A0A0N1INZ7 A0A2H1VWU9 H9JCA9 A0A3S2NL29 H9JCF9 A0A212FI86 E9JCL6 A0A0M9A4Z2 F4WBF5 A0A1B6JKQ8 A0A195B559 A0A151JP46 A0A151X9S7 A0A151K1W4 A0A2J7QWR1 A0A1B6MQ51 A0A1B6FAH8 E2B035 A0A2A3EMT5 A0A310SCB8 A0A232EY82 K7ISI1 A0A1B6CQY0 A0A1B6DDB5 E0VFL1 A0A1B6FEK1 A0A3L8DBS4 A0A026WKB9 A0A0L7L314 A0A2P8Y960 A0A087ZZF8 A0A0J7NFB9 E2BIZ7 A0A1Y1K255 D6WW59 J9JX09 A0A154PIA3 A0A194PIB8 A0A2H1VBZ1 A0A195CUZ7 H9J327 A0A2W1C0X9 A0A224X9P2 A0A194RJQ5 N6U8D4 T1HW00 A0A1B6D3S6 A0A212EL79 N6U5M4 A0A1W4W461 A0A2J7QWT6 A0A310S6P6 A0A087ZZF7 A0A154PI87 A0A224XKF4 A0A1B6C2J0 A0A0A9XLI7 E2B036 A0A026WLG7 T1JKY7 A0A1Y1M0L3 A0A1B6GX81 U5ETA5 E0VFK5 A0A1W4W469 A0A1L8DE17 A0A1L8DE24 F4WBF6 A0A1L8DE12 A0A1J1IHT6 E2BIZ9 A0A1L8DEE9 A0A336M5C5 A0A2S2QFW9 A0A0L7QNA7 A0A158NQA4 U5EMI5 A0A195CTG3 T1IP78 A0A067QTW1 A0A195B5U1 E9JCL7 A0A151JNR2 A0A2J7QWQ4 D6WW60 A0A162CFD9 A0A0P5EBE7
A0A194RIL6 A0A212EL73 A0A0N0P9U3 A0A0L7KU96 A0A194QG55 A0A2A4JUI4 A0A2W1BN40 A0A0N1INZ7 A0A2H1VWU9 H9JCA9 A0A3S2NL29 H9JCF9 A0A212FI86 E9JCL6 A0A0M9A4Z2 F4WBF5 A0A1B6JKQ8 A0A195B559 A0A151JP46 A0A151X9S7 A0A151K1W4 A0A2J7QWR1 A0A1B6MQ51 A0A1B6FAH8 E2B035 A0A2A3EMT5 A0A310SCB8 A0A232EY82 K7ISI1 A0A1B6CQY0 A0A1B6DDB5 E0VFL1 A0A1B6FEK1 A0A3L8DBS4 A0A026WKB9 A0A0L7L314 A0A2P8Y960 A0A087ZZF8 A0A0J7NFB9 E2BIZ7 A0A1Y1K255 D6WW59 J9JX09 A0A154PIA3 A0A194PIB8 A0A2H1VBZ1 A0A195CUZ7 H9J327 A0A2W1C0X9 A0A224X9P2 A0A194RJQ5 N6U8D4 T1HW00 A0A1B6D3S6 A0A212EL79 N6U5M4 A0A1W4W461 A0A2J7QWT6 A0A310S6P6 A0A087ZZF7 A0A154PI87 A0A224XKF4 A0A1B6C2J0 A0A0A9XLI7 E2B036 A0A026WLG7 T1JKY7 A0A1Y1M0L3 A0A1B6GX81 U5ETA5 E0VFK5 A0A1W4W469 A0A1L8DE17 A0A1L8DE24 F4WBF6 A0A1L8DE12 A0A1J1IHT6 E2BIZ9 A0A1L8DEE9 A0A336M5C5 A0A2S2QFW9 A0A0L7QNA7 A0A158NQA4 U5EMI5 A0A195CTG3 T1IP78 A0A067QTW1 A0A195B5U1 E9JCL7 A0A151JNR2 A0A2J7QWQ4 D6WW60 A0A162CFD9 A0A0P5EBE7
Pubmed
EMBL
BABH01037649
KZ149921
PZC77732.1
NWSH01007033
PCG63126.1
ODYU01000260
+ More
SOQ34727.1 RSAL01000304 RVE42716.1 KQ459603 KPI93154.1 KQ460124 KPJ17668.1 AGBW02014130 OWR42229.1 KQ459435 KPJ01099.1 JTDY01005603 KOB66842.1 KQ458981 KPJ04472.1 NWSH01000554 PCG75685.1 KZ150013 PZC75044.1 KQ460882 KPJ11474.1 ODYU01004928 SOQ45305.1 BABH01031609 RSAL01000317 RVE42585.1 BABH01031610 AGBW02008415 OWR53453.1 GL771849 EFZ09427.1 KQ435732 KOX77337.1 GL888062 EGI68511.1 GECU01028486 GECU01022863 GECU01016348 GECU01008024 JAS79220.1 JAS84843.1 JAS91358.1 JAS99682.1 KQ976595 KYM79626.1 KQ978820 KYN28133.1 KQ982365 KYQ57127.1 KQ981189 KYN45223.1 NEVH01009422 PNF33023.1 GEBQ01017028 GEBQ01016920 GEBQ01012615 GEBQ01001956 JAT22949.1 JAT23057.1 JAT27362.1 JAT38021.1 GECZ01022618 GECZ01008634 JAS47151.1 JAS61135.1 GL444372 EFN60961.1 KZ288212 PBC32804.1 KQ779356 OAD51997.1 NNAY01001665 OXU23277.1 GEDC01021494 JAS15804.1 GEDC01013652 JAS23646.1 DS235115 EEB12167.1 GECZ01021235 JAS48534.1 QOIP01000010 RLU17786.1 KK107167 EZA56497.1 JTDY01003280 KOB69810.1 PYGN01000790 PSN40793.1 LBMM01005754 KMQ91230.1 GL448543 EFN84342.1 GEZM01099141 JAV53466.1 KQ971361 EFA08671.2 ABLF02030750 KQ434924 KZC11585.1 KPI93156.1 ODYU01001747 SOQ38363.1 KQ977279 KYN03989.1 BABH01037651 PZC77733.1 GFTR01007370 JAW09056.1 KPJ17669.1 APGK01038976 APGK01038977 APGK01038978 APGK01038979 KB740966 ENN76911.1 ACPB03014175 GEDC01030362 GEDC01016970 JAS06936.1 JAS20328.1 OWR42228.1 APGK01038973 KB631578 ENN76910.1 ERL84404.1 PNF33024.1 OAD51996.1 KZC11586.1 GFTR01007803 JAW08623.1 GEDC01029848 JAS07450.1 GBHO01025619 GDHC01000641 JAG17985.1 JAQ17988.1 EFN60962.1 EZA56496.1 RLU17785.1 JH431785 GEZM01042994 GEZM01042993 JAV79402.1 GECZ01002733 JAS67036.1 GANO01004466 JAB55405.1 EEB12161.1 GFDF01009383 JAV04701.1 GFDF01009382 JAV04702.1 EGI68512.1 GFDF01009381 JAV04703.1 CVRI01000053 CRK99823.1 EFN84344.1 GFDF01009380 JAV04704.1 UFQS01000367 UFQT01000367 SSX03211.1 SSX23577.1 GGMS01007197 MBY76400.1 KQ414855 KOC60127.1 ADTU01022996 GANO01004465 JAB55406.1 KYN03988.1 JH431250 KK853278 KDR09062.1 KYM79627.1 EFZ09435.1 KYN28134.1 PNF33018.1 EFA08184.1 LRGB01000915 KZS15182.1 GDIP01143974 JAJ79428.1
SOQ34727.1 RSAL01000304 RVE42716.1 KQ459603 KPI93154.1 KQ460124 KPJ17668.1 AGBW02014130 OWR42229.1 KQ459435 KPJ01099.1 JTDY01005603 KOB66842.1 KQ458981 KPJ04472.1 NWSH01000554 PCG75685.1 KZ150013 PZC75044.1 KQ460882 KPJ11474.1 ODYU01004928 SOQ45305.1 BABH01031609 RSAL01000317 RVE42585.1 BABH01031610 AGBW02008415 OWR53453.1 GL771849 EFZ09427.1 KQ435732 KOX77337.1 GL888062 EGI68511.1 GECU01028486 GECU01022863 GECU01016348 GECU01008024 JAS79220.1 JAS84843.1 JAS91358.1 JAS99682.1 KQ976595 KYM79626.1 KQ978820 KYN28133.1 KQ982365 KYQ57127.1 KQ981189 KYN45223.1 NEVH01009422 PNF33023.1 GEBQ01017028 GEBQ01016920 GEBQ01012615 GEBQ01001956 JAT22949.1 JAT23057.1 JAT27362.1 JAT38021.1 GECZ01022618 GECZ01008634 JAS47151.1 JAS61135.1 GL444372 EFN60961.1 KZ288212 PBC32804.1 KQ779356 OAD51997.1 NNAY01001665 OXU23277.1 GEDC01021494 JAS15804.1 GEDC01013652 JAS23646.1 DS235115 EEB12167.1 GECZ01021235 JAS48534.1 QOIP01000010 RLU17786.1 KK107167 EZA56497.1 JTDY01003280 KOB69810.1 PYGN01000790 PSN40793.1 LBMM01005754 KMQ91230.1 GL448543 EFN84342.1 GEZM01099141 JAV53466.1 KQ971361 EFA08671.2 ABLF02030750 KQ434924 KZC11585.1 KPI93156.1 ODYU01001747 SOQ38363.1 KQ977279 KYN03989.1 BABH01037651 PZC77733.1 GFTR01007370 JAW09056.1 KPJ17669.1 APGK01038976 APGK01038977 APGK01038978 APGK01038979 KB740966 ENN76911.1 ACPB03014175 GEDC01030362 GEDC01016970 JAS06936.1 JAS20328.1 OWR42228.1 APGK01038973 KB631578 ENN76910.1 ERL84404.1 PNF33024.1 OAD51996.1 KZC11586.1 GFTR01007803 JAW08623.1 GEDC01029848 JAS07450.1 GBHO01025619 GDHC01000641 JAG17985.1 JAQ17988.1 EFN60962.1 EZA56496.1 RLU17785.1 JH431785 GEZM01042994 GEZM01042993 JAV79402.1 GECZ01002733 JAS67036.1 GANO01004466 JAB55405.1 EEB12161.1 GFDF01009383 JAV04701.1 GFDF01009382 JAV04702.1 EGI68512.1 GFDF01009381 JAV04703.1 CVRI01000053 CRK99823.1 EFN84344.1 GFDF01009380 JAV04704.1 UFQS01000367 UFQT01000367 SSX03211.1 SSX23577.1 GGMS01007197 MBY76400.1 KQ414855 KOC60127.1 ADTU01022996 GANO01004465 JAB55406.1 KYN03988.1 JH431250 KK853278 KDR09062.1 KYM79627.1 EFZ09435.1 KYN28134.1 PNF33018.1 EFA08184.1 LRGB01000915 KZS15182.1 GDIP01143974 JAJ79428.1
Proteomes
UP000005204
UP000218220
UP000283053
UP000053268
UP000053240
UP000007151
+ More
UP000037510 UP000053105 UP000007755 UP000078540 UP000078492 UP000075809 UP000078541 UP000235965 UP000000311 UP000242457 UP000215335 UP000002358 UP000009046 UP000279307 UP000053097 UP000245037 UP000005203 UP000036403 UP000008237 UP000007266 UP000007819 UP000076502 UP000078542 UP000019118 UP000015103 UP000030742 UP000192223 UP000183832 UP000053825 UP000005205 UP000027135 UP000076858
UP000037510 UP000053105 UP000007755 UP000078540 UP000078492 UP000075809 UP000078541 UP000235965 UP000000311 UP000242457 UP000215335 UP000002358 UP000009046 UP000279307 UP000053097 UP000245037 UP000005203 UP000036403 UP000008237 UP000007266 UP000007819 UP000076502 UP000078542 UP000019118 UP000015103 UP000030742 UP000192223 UP000183832 UP000053825 UP000005205 UP000027135 UP000076858
Pfam
PF07690 MFS_1
SUPFAM
SSF103473
SSF103473
CDD
ProteinModelPortal
H9J328
A0A2W1BRL4
A0A2A4IUF1
A0A2H1V1L2
A0A3S2NAV0
A0A194PK99
+ More
A0A194RIL6 A0A212EL73 A0A0N0P9U3 A0A0L7KU96 A0A194QG55 A0A2A4JUI4 A0A2W1BN40 A0A0N1INZ7 A0A2H1VWU9 H9JCA9 A0A3S2NL29 H9JCF9 A0A212FI86 E9JCL6 A0A0M9A4Z2 F4WBF5 A0A1B6JKQ8 A0A195B559 A0A151JP46 A0A151X9S7 A0A151K1W4 A0A2J7QWR1 A0A1B6MQ51 A0A1B6FAH8 E2B035 A0A2A3EMT5 A0A310SCB8 A0A232EY82 K7ISI1 A0A1B6CQY0 A0A1B6DDB5 E0VFL1 A0A1B6FEK1 A0A3L8DBS4 A0A026WKB9 A0A0L7L314 A0A2P8Y960 A0A087ZZF8 A0A0J7NFB9 E2BIZ7 A0A1Y1K255 D6WW59 J9JX09 A0A154PIA3 A0A194PIB8 A0A2H1VBZ1 A0A195CUZ7 H9J327 A0A2W1C0X9 A0A224X9P2 A0A194RJQ5 N6U8D4 T1HW00 A0A1B6D3S6 A0A212EL79 N6U5M4 A0A1W4W461 A0A2J7QWT6 A0A310S6P6 A0A087ZZF7 A0A154PI87 A0A224XKF4 A0A1B6C2J0 A0A0A9XLI7 E2B036 A0A026WLG7 T1JKY7 A0A1Y1M0L3 A0A1B6GX81 U5ETA5 E0VFK5 A0A1W4W469 A0A1L8DE17 A0A1L8DE24 F4WBF6 A0A1L8DE12 A0A1J1IHT6 E2BIZ9 A0A1L8DEE9 A0A336M5C5 A0A2S2QFW9 A0A0L7QNA7 A0A158NQA4 U5EMI5 A0A195CTG3 T1IP78 A0A067QTW1 A0A195B5U1 E9JCL7 A0A151JNR2 A0A2J7QWQ4 D6WW60 A0A162CFD9 A0A0P5EBE7
A0A194RIL6 A0A212EL73 A0A0N0P9U3 A0A0L7KU96 A0A194QG55 A0A2A4JUI4 A0A2W1BN40 A0A0N1INZ7 A0A2H1VWU9 H9JCA9 A0A3S2NL29 H9JCF9 A0A212FI86 E9JCL6 A0A0M9A4Z2 F4WBF5 A0A1B6JKQ8 A0A195B559 A0A151JP46 A0A151X9S7 A0A151K1W4 A0A2J7QWR1 A0A1B6MQ51 A0A1B6FAH8 E2B035 A0A2A3EMT5 A0A310SCB8 A0A232EY82 K7ISI1 A0A1B6CQY0 A0A1B6DDB5 E0VFL1 A0A1B6FEK1 A0A3L8DBS4 A0A026WKB9 A0A0L7L314 A0A2P8Y960 A0A087ZZF8 A0A0J7NFB9 E2BIZ7 A0A1Y1K255 D6WW59 J9JX09 A0A154PIA3 A0A194PIB8 A0A2H1VBZ1 A0A195CUZ7 H9J327 A0A2W1C0X9 A0A224X9P2 A0A194RJQ5 N6U8D4 T1HW00 A0A1B6D3S6 A0A212EL79 N6U5M4 A0A1W4W461 A0A2J7QWT6 A0A310S6P6 A0A087ZZF7 A0A154PI87 A0A224XKF4 A0A1B6C2J0 A0A0A9XLI7 E2B036 A0A026WLG7 T1JKY7 A0A1Y1M0L3 A0A1B6GX81 U5ETA5 E0VFK5 A0A1W4W469 A0A1L8DE17 A0A1L8DE24 F4WBF6 A0A1L8DE12 A0A1J1IHT6 E2BIZ9 A0A1L8DEE9 A0A336M5C5 A0A2S2QFW9 A0A0L7QNA7 A0A158NQA4 U5EMI5 A0A195CTG3 T1IP78 A0A067QTW1 A0A195B5U1 E9JCL7 A0A151JNR2 A0A2J7QWQ4 D6WW60 A0A162CFD9 A0A0P5EBE7
Ontologies
GO
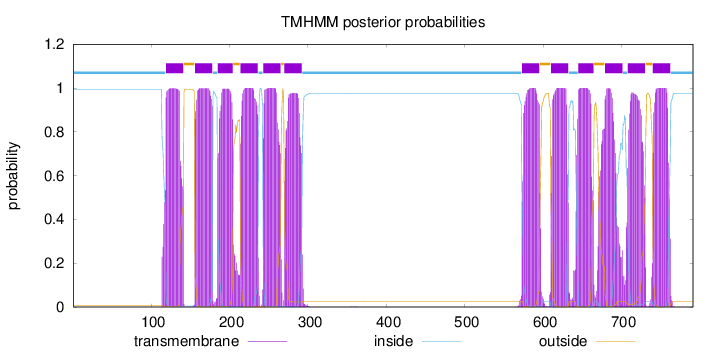
Topology
Length:
791
Number of predicted TMHs:
12
Exp number of AAs in TMHs:
261.39234
Exp number, first 60 AAs:
0
Total prob of N-in:
0.99477
inside
1 - 118
TMhelix
119 - 141
outside
142 - 155
TMhelix
156 - 178
inside
179 - 184
TMhelix
185 - 204
outside
205 - 213
TMhelix
214 - 236
inside
237 - 242
TMhelix
243 - 265
outside
266 - 269
TMhelix
270 - 292
inside
293 - 572
TMhelix
573 - 595
outside
596 - 609
TMhelix
610 - 632
inside
633 - 644
TMhelix
645 - 664
outside
665 - 678
TMhelix
679 - 701
inside
702 - 707
TMhelix
708 - 730
outside
731 - 739
TMhelix
740 - 762
inside
763 - 791
Population Genetic Test Statistics
Pi
181.61114
Theta
166.353423
Tajima's D
-0.75779
CLR
171.82485
CSRT
0.190840457977101
Interpretation
Uncertain
