Gene
KWMTBOMO10356 Validated by peptides from experiments
Annotation
PREDICTED:_uncharacterized_protein_LOC106142516_isoform_X1_[Amyelois_transitella]
Location in the cell
Nuclear Reliability : 2.357
Sequence
CDS
ATGGAATTGACTCGAGAAAATTCAAGAGCGATGATTTATTATGACTTTCGAAGTGGTTTAACACAAAAACAGTGTGTTGACCGGATGATTTCTGCATTTGGTGATGAAGCCCCATCCAAAACCACAATTTATCGCTGGTTTGCTGAGTTTCAACGTGGACGTGTCAAGCTTAGTGATGATCCCCGTCAAGGACGTCTAAAAACTGCAGCCACCCAAGAAAACGTTGATGCTGTGCGTAAGCTGATTGAGGAAGATCGACATGTGACATACCGCGAAATTCAGGCAGCTTTAGACATTGGCATGAGTCAAATACAAATAATTTTGCAGGAACAATTAGGTGTAAAAAAGTTGTTTTCCCGATGGATACCGCATTCGCTCTGTGAAGAGCAAAAAGCGGCTCGCGTTACTTGTACAATGGATGAACACACCTCAACGGACACAGTGCAAGAAAGCTCAAATCCAACATTACATGTTGAAGCACCAACAGATTGTGACTTTAACTTAACCCCTGTACAAATACAACAGCAATATTTTGAAAACTTAGGGGATTGTAACTGGATTAGATATGAAAAGTGCAATAGAAATCATTTATTGCAAAACTTGGCATTTGGACTATTTGGTCCTCCGATTACTGAAGGGGAGCTGATAGAAACGGAAAAAATGAATGCAATTAATCTTCAAGGATACAATTCCAGACAAAAAGAAAAAGTTGACTTAGTCTATGCAAAAATATGTGAACAAGGTAAATATTCATTTGAAGATGAAATTTACATGTCGAATTTACTGGTTGTTTGTGCACGTCCAAAGCCCAAGAATTTTTTAAACATTCAACCGTCGAAATGTTGGATTGATCTACACGAACGGAGTGACTCCGAAATATGGCTCGTACCTGTCTTTTGCATAAGAAAATGTATACCTATAGAAATGAATGCGAAACCATGCAAAATATATATTGATGACAATGCCCGGGTGTACCATAGTTGGAATGCTTACTTGACAGAAAACACATTGGACAAATGTGTTCTAATAGTCCCTGAGAACGGAGAATATACCGGAACACTTACTGAGGACGGAACAGACTTCACGGTGAATCTAACAGTGGCGCCGTCACCTGCCCTCGGACTGAAGTCACGCGTGTTGAGTTCGGTGGACACAGCGACCACGGTGGTCTCGCTGGGCGCCGTGGGAGTGGTCGGGGTGGCCGCGCTAACACCGGTGGCGCCCGTGCTGCTCGCAAGCGCGGCCACCGTCTCCGTCACCGCGGGAATCTACGGGCTGATCAGGTCTTCGTTGCATTTACACGACCGAAGAACGCACGAACAGACCATAAACGTGCGCAATTCTGAAGCTCGTAACAGTTGGTTAAACGTAGCGACCTCGGGCGTGGGTCTCACGGCTGGAGCGGCCAGCAATCTCCTGTCTAAATCAGCAGCGGCCGGCACCAACCTCACTACGTCGGGGAGGTTCTTGACTAAGTCAGTCGACGTGCTGCGACACGTGAACCTTTTGACCGGAGCTGCGAGCTTTACTAATGGCTTTGTGTATCTTATAATACAGTACAGGAAACACGGTCAGAAGCCGAGTCGTATTGAGATATTCCAATTCGCCATGTCGACGTTGTTCTTCTTCAACGCTGTCGTATCGAATCAGACGGCGCAAGAGATCATTGAGCAGAGTCAGGCGAACAAGATAAATGAGTTCCGCGATACGCTCCGGAGTAATCGGCACAGGAAAATTTTCGATAAGCTTTCCGCCGAAGCTAGACGAGTGCGGGGCACCGTTCAAGGTAACACCGAAGTCATCCAGGGCATCAAGAATATCGCTAACAAAGACGAATACTTTGCGCAAGTTCTGAGCATCAACAAGGACGTGAACCAGCACAGGCTCAGGATATCGATGACGGCCGACGGCCGGGTCATGTTGAACAACCAACACGGTTACAACCCGTCCGAGTTACACTCGCTCGGCTCGGAAGGGAGATCGCAAATATTCACCGAAATCGGCCCAGCCCCCCCGGGGCCTGCTAATGTGTCCACACGTCTGAACTCGCCCGTCAACAGAGTGGACAACTACTTCGGAGTAGACGACGAAGAAGGCCATCTATTAGTCGGCATCCGTCCAGAAGAGATACTCCAAATCGGCACATATTTCGTGCGTATCAGCTCGTCGGGCACCGATGAAATAGTTTCGATGCTCCAAAATCTCTCCGAAGAGGTCTATACGAATTTGATGACCCTATCCTTTAATCTGCTGTCGAAACTTGTGCCAAGCGAAATTGCCAATCTGAAATTGCTCGATCCGGAGCGAGATTTGATTTCGCAGGTAGTTGAATTTGTTTTCAATTACATGAAACAGCGGAGACCGTTCGGGGAAGCGAGACACGACGATAACGGAATCGTTGAAGTAGTAAAAGACTTCTTTCACGATGGCCGAGTTAATCAAGACACGATATTGCGATTGAAGAAAGATTTAATCGAATGGCTGGACGCTGAAATTACACGTAAAAATGAGCTAAATCCTAATAAGAAATCCTTAATCTGCAATCTGTGTAAAGGAGTTAGATACGCTTAA
Protein
MELTRENSRAMIYYDFRSGLTQKQCVDRMISAFGDEAPSKTTIYRWFAEFQRGRVKLSDDPRQGRLKTAATQENVDAVRKLIEEDRHVTYREIQAALDIGMSQIQIILQEQLGVKKLFSRWIPHSLCEEQKAARVTCTMDEHTSTDTVQESSNPTLHVEAPTDCDFNLTPVQIQQQYFENLGDCNWIRYEKCNRNHLLQNLAFGLFGPPITEGELIETEKMNAINLQGYNSRQKEKVDLVYAKICEQGKYSFEDEIYMSNLLVVCARPKPKNFLNIQPSKCWIDLHERSDSEIWLVPVFCIRKCIPIEMNAKPCKIYIDDNARVYHSWNAYLTENTLDKCVLIVPENGEYTGTLTEDGTDFTVNLTVAPSPALGLKSRVLSSVDTATTVVSLGAVGVVGVAALTPVAPVLLASAATVSVTAGIYGLIRSSLHLHDRRTHEQTINVRNSEARNSWLNVATSGVGLTAGAASNLLSKSAAAGTNLTTSGRFLTKSVDVLRHVNLLTGAASFTNGFVYLIIQYRKHGQKPSRIEIFQFAMSTLFFFNAVVSNQTAQEIIEQSQANKINEFRDTLRSNRHRKIFDKLSAEARRVRGTVQGNTEVIQGIKNIANKDEYFAQVLSINKDVNQHRLRISMTADGRVMLNNQHGYNPSELHSLGSEGRSQIFTEIGPAPPGPANVSTRLNSPVNRVDNYFGVDDEEGHLLVGIRPEEILQIGTYFVRISSSGTDEIVSMLQNLSEEVYTNLMTLSFNLLSKLVPSEIANLKLLDPERDLISQVVEFVFNYMKQRRPFGEARHDDNGIVEVVKDFFHDGRVNQDTILRLKKDLIEWLDAEITRKNELNPNKKSLICNLCKGVRYA
Summary
Uniprot
EMBL
Proteomes
Gene 3D
ProteinModelPortal
PDB
4U7B
E-value=1.00059e-07,
Score=138
Ontologies
GO
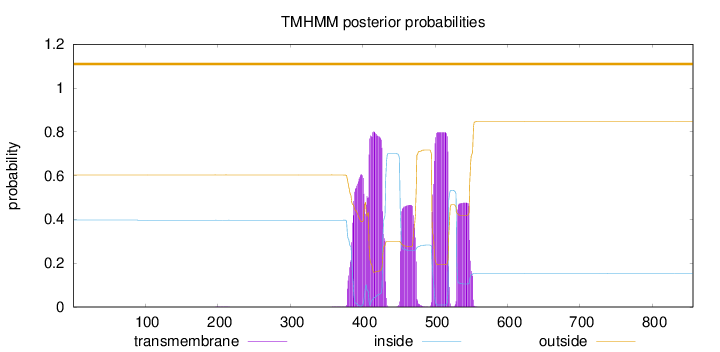
Topology
Length:
856
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
66.4274600000001
Exp number, first 60 AAs:
0.00019
Total prob of N-in:
0.39679
outside
1 - 856
Population Genetic Test Statistics
Pi
191.897272
Theta
163.846103
Tajima's D
0.253994
CLR
1.496439
CSRT
0.448577571121444
Interpretation
Uncertain
