Gene
KWMTBOMO10241 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA012651
Annotation
PREDICTED:_uncharacterized_protein_LOC106139693_isoform_X1_[Amyelois_transitella]
Location in the cell
Mitochondrial Reliability : 2.088
Sequence
CDS
ATGTCTTTGAAGAATAGGTTAATTGCATGTCTGCAAAAACTGAAGGAATACGATTACACTGACGTAGACAAGGCAAGAGCTAGTGACGCGAAGGCCGAGAATGCCCTCAACTTAATATACCAGACTGCTTTAAAGAAAACAACTTTTACACCAGAAGAGAAGAAAATTGTAGGTTCTCTTTTATTTGATGCCATCACACCGATACAAAGCAACATAGAAGGAACGCTCTGCGAATACAGAACTAGATTAGATACCTCTGTGCTTATGAAAAGATCTGCAATACAATTCTTGATTGACGAATACGGGAAGTTTCCAGTTGAGAATTCTATCTTGGCTACAAAGTTAGAGGAAGCCGGCATAGCAGAGAGCGTTGAAATACTCAATGAAATTATAAATAGGACATCGGCGGTCCTAGCAGGAGGAGCAGTTGGATGCCTCGCAGGACCGGCAGGAGCCGCATTGGGAGGAGTATACGGAGGCATGATTACAGACGGCTTGGTTTCTGCGGCTAAGAACGAGCCTGTAGGTATTTTTGAGTCAAGTGCAAATATAGGTAGAGACATTGCTTCAGGCAAAAATCCAATGGCTTCTGTGGGTTCCGCAAGTTTCAAAATCGCCATGGACGGTATAGGTGGCTTTGGCATGGGAGGAGTTTCTAGCGTTGCCAGTAAAGTCCCGGCGAAGGCCGTCGCAAAAACCATAGAAGGCACTGTAAAAAAAGCTGCTGCAACTATTGGGGAGGAGATGATTGAGAAGTCAGTGCAATTAACTGCAGAAGAAATGACAAAAATTGTGGCAAAGAAAGTGTCAGAAAAAGCCGTGAAAAAAGCTGTCGAAACTGCAGCAATACAAACACAAATGCTTCTGGTGTCTGCTCACGTCGGAGTAAATGCTAGTGAAAAAACCAAAGCTGAAACGCGTGCTGAAGAAGAACGTAAACAACAGAAACAAAAAGAAGAAGAAAAAAAGGCGCGTGAAAATCGCGATAAAAAAAATACTGGATCGTGGCAACCGCCACGCGAGAACAGAAACAATAAAGAGTCCAATGTTGATAATGAAGAGTCCGACGAGGAGTTGTATTTTATGACCAAAACCAAAGGTAAACCGACTCGTCAATCAGATGCCGACCGAAATGAGCGTCGAAGGAACGCCATCAGAGAACTTCAAAACAAAAGGGATATAATTCAGAAAATTGCTGATCTATTCAGACATTTACAAGCGGAAATCATAGCTGAAACGAGCGCGCCTAATATAATTTCCCCCTATTTTCGTGATATATTCTCAGCTGTGTATCATTATTACAAACACAGACGCGTGCCCGTGCCAGGCCATGAGAAGCTGACAGTCAAACAATATTTCGACTTGATAAGGATGGTAATTATAAAAATGAAAGAATCACCGGAATACAAAAAAATGCAGAAGGCCAGAAATGGAACGGCCCTCACTTGTGAAATGTACGTGGTCTTGTTCGGTAGGATGTTCAGAATTATTATTACTGTTCGAGATGGTGAAATTGTATTGAGTACCGCCTATGTAGTCAAGCGTATGAACTTTAATGCCAAAGAAGGCATTCTTTACATAAAATTCGGTGATGGAGATGTACAGAAGTAA
Protein
MSLKNRLIACLQKLKEYDYTDVDKARASDAKAENALNLIYQTALKKTTFTPEEKKIVGSLLFDAITPIQSNIEGTLCEYRTRLDTSVLMKRSAIQFLIDEYGKFPVENSILATKLEEAGIAESVEILNEIINRTSAVLAGGAVGCLAGPAGAALGGVYGGMITDGLVSAAKNEPVGIFESSANIGRDIASGKNPMASVGSASFKIAMDGIGGFGMGGVSSVASKVPAKAVAKTIEGTVKKAAATIGEEMIEKSVQLTAEEMTKIVAKKVSEKAVKKAVETAAIQTQMLLVSAHVGVNASEKTKAETRAEEERKQQKQKEEEKKARENRDKKNTGSWQPPRENRNNKESNVDNEESDEELYFMTKTKGKPTRQSDADRNERRRNAIRELQNKRDIIQKIADLFRHLQAEIIAETSAPNIISPYFRDIFSAVYHYYKHRRVPVPGHEKLTVKQYFDLIRMVIIKMKESPEYKKMQKARNGTALTCEMYVVLFGRMFRIIITVRDGEIVLSTAYVVKRMNFNAKEGILYIKFGDGDVQK
Summary
Uniprot
ProteinModelPortal
Ontologies
GO
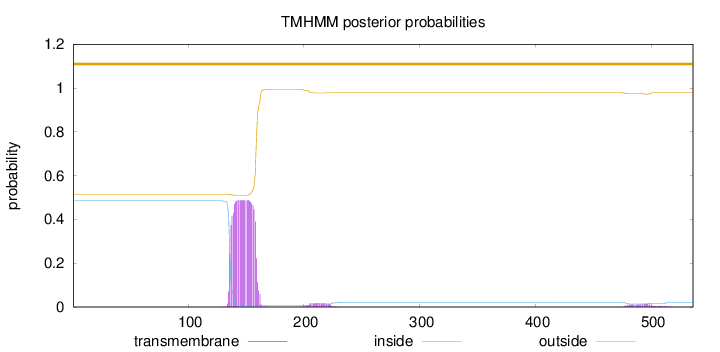
Topology
Length:
536
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
11.77963
Exp number, first 60 AAs:
0.00025
Total prob of N-in:
0.48524
outside
1 - 536
Population Genetic Test Statistics
Pi
332.32171
Theta
182.541204
Tajima's D
2.691911
CLR
0.512259
CSRT
0.95965201739913
Interpretation
Uncertain
