Gene
KWMTBOMO10137
Pre Gene Modal
BGIBMGA005562
Annotation
PREDICTED:_uncharacterized_protein_KIAA0195?_partial_[Bombyx_mori]
Location in the cell
PlasmaMembrane Reliability : 4.388
Sequence
CDS
ATGCTCACACAGGGTACCGCAGATATAGTGTTAGACTCGTGCGTAGATTACTGGAACGGGCGGGACTTAACACCGATATCTGCTTCCGACCGAAAGAAGATCATAGATTTTTACCAAAGAAACTCACTGACGGCCTATTGTACCGCTTTCGCTTACAAGCCGGTGACGCGCGGCGTGTCGTCGCCCGTGGCGGACATGTACCTGGAGGTGCCGGCGGAGGGGCGCGGGGGGCTGGCGCCGCGCACGCAGCACTGGCCGGACGCGCCCGCACTGCACTTTCACTCAACTGATTCCCTACTGTTCAATGAAGTGACCGACGACGATGTTGTTGACGCCGATGGATATTTCGATCTGCAATGCAATCAAGTGTTCATAGGCATGGTGACGATGCAGTACCAGGCGCAGAGCGACATGGTGGAGCTGATAGAGCGGCTAGAGCGGGCCTGCATACGCTTCGTCCACTTCAGCAAGGAGAACGAGCTCCGGTCGAGGGTGTTCAGCGAAAAGATGGGCTTGGAGTCCGGCTGGAACTGTCACATATCGCTCATGAGCGACACCGAAGCCAGCAAAAGCGCATCGCCGCTGTCCTCGCAGAATATCGCGCAGCCAAAGGAACCTAACATCGAGAGGCGGCCCTCTAGGGGCTTCGAAACTTTTGACGCTCTCGGCTTTTGTCATAGCAATAGTATGGCAACATCGAAAGCGTTGTCTATGTCGGCGCCGGGAGCGATCAATCTAGATCACACTACCGTCAAATTCGACAACGAGGTCAAGAAAGCGAGGCACTCCATCACAACTGGAATCAAGATGCAGCGTGGCAGCATAGCGACGACGCATCCGTCGGCGGACTCGGCTCTGGACTCGCGCGAGGAGCCGGAGTCCCCGGTGGACGCGTGCCGCTCGCTGTCCTGCCTCACCGACTCCACCGACCACTCGGCGCAGCTCAACTTCGACATGAGCAATAGGGCTAAGTTGCCCCGCGGCATCGAGAACATCCGACCCCACATCGAGCAAGTGGACAACGTTCCGCTCCTCGTCTCGCTGTTCACGGACTGCACCGCGCGCTCCGTCCAGCTCATGATACAGATCATGCAGGATTACAGCGAGGTGGTTTGCGTGATGGGCTCCGCGGCCAACTGTCTCAACATGGAAATCTTCATGCAGGCCGACGCTAGCATCGCGGTTGAGCCTCTATATCCGGTACTGTGCCAAAAGATGCAACCGTACTTGATTCCCGACTCCTGCATCGGTCCCATCGACCTAGCTAGAGCACTCAATTCGGTCCCCTGTTCTTTGTCGATGACGCGGGAATCGGATCTGTCACTGTTCACATTAATAACGACGTCGCGTCACTTCATGACGTGCCTGTGGAACTGCACTCAGTTCTGGATATGCAGCGTTTGTTTCATAGCCCTGTTACAGACAGGTTCCCTGTGCGGGTTCTTGCCGCTGGCGCTGAGCGTGGGCGGCGCGGCGTGGTGCGGGCTCGTGTCGGTGCCGCACGTGGCGGGCGCGCTCGCCCTCGCCCCTCCGCACCCCGCGCTCTCCCAGCGCGCTGCCACCAGGCCAGACCATCAGCTGAACCTCAAGATGGCGGTGTTCGTGTTCTGGTGCTACGCGTGCAAGTTCGTGCCGGCCGCCGTCTCCATCCTCGTGCTGTACGGGCTGACGCTGCGCAGCTTCTGCGAGCAGATCGCGGCCGCCACCAACGCGACGTCGTGCTGGATCGTGTACCCGGTGAACGTGGGCGCGTTCCCGCCCAGCCACGACCTCACCGCGAGGAGCTGGCACGGCTGGGGCGACGACTTCTACGACGGATTCTATTTGACCCAACATATCGTGCTCGCTTTGATAACTTTGCATTTAATTGTGATCTCATTATCATTCGTCCATCGCTACTCGTCGATATGGCAACGTGGCTTCTGGACCAATCGAGCGTGGACTATCACTGTTGTCGTACTGCTGGTAGTCCAGTGCGCGTTCAGCGGCGTGATGCTGAGCGGCACGTACGGGTCGTGCGCGCGCTGGTCGGTGTGCGCGCTGCGGTACCGCGTGCCGTGGCCGCTGCTGCTCGCGTACGCCGCCTCGCTGCTGCTCGTGCTCGCGCTCAACGAGCTCATCAAGTGGCAGGAGATCAAGGCAGACAGACGGAATCAACGCCGCGCCCGCCTCGATTTCGGAACCAAACTGGGAATGAATTCTCCATTCTAA
Protein
MLTQGTADIVLDSCVDYWNGRDLTPISASDRKKIIDFYQRNSLTAYCTAFAYKPVTRGVSSPVADMYLEVPAEGRGGLAPRTQHWPDAPALHFHSTDSLLFNEVTDDDVVDADGYFDLQCNQVFIGMVTMQYQAQSDMVELIERLERACIRFVHFSKENELRSRVFSEKMGLESGWNCHISLMSDTEASKSASPLSSQNIAQPKEPNIERRPSRGFETFDALGFCHSNSMATSKALSMSAPGAINLDHTTVKFDNEVKKARHSITTGIKMQRGSIATTHPSADSALDSREEPESPVDACRSLSCLTDSTDHSAQLNFDMSNRAKLPRGIENIRPHIEQVDNVPLLVSLFTDCTARSVQLMIQIMQDYSEVVCVMGSAANCLNMEIFMQADASIAVEPLYPVLCQKMQPYLIPDSCIGPIDLARALNSVPCSLSMTRESDLSLFTLITTSRHFMTCLWNCTQFWICSVCFIALLQTGSLCGFLPLALSVGGAAWCGLVSVPHVAGALALAPPHPALSQRAATRPDHQLNLKMAVFVFWCYACKFVPAAVSILVLYGLTLRSFCEQIAAATNATSCWIVYPVNVGAFPPSHDLTARSWHGWGDDFYDGFYLTQHIVLALITLHLIVISLSFVHRYSSIWQRGFWTNRAWTITVVVLLVVQCAFSGVMLSGTYGSCARWSVCALRYRVPWPLLLAYAASLLLVLALNELIKWQEIKADRRNQRRARLDFGTKLGMNSPF
Summary
Uniprot
A0A2A4JEV8
H9J7S0
A0A3S2L8H5
A0A194QSL0
L7X3W0
D0AB86
+ More
A0A212EPB0 A0A067QXM3 A0A2J7PXG9 A0A2J7PXI5 A0A2J7PXH2 A0A2J7PXH6 A0A2A3EAG4 A0A232ER02 A0A310SMW2 A0A0L7QQV6 A0A087ZXX4 E2B474 A0A3L8DJS8 A0A195CWZ6 F4WHR9 A0A195B5D9 A0A151JXP4 A0A158NM24 A0A0C9PL89 A0A151XIG3 A0A026WG49 A0A0C9RMK5 A0A0C9QMF3 A0A1B6EF44 A0A1B6DN27 A0A1B6BXT8 A0A0N0U4H9 A0A2J7PXH8 A0A224XDD0 A0A1B6BYR1 A0A1B6CCC8 A0A023F1Q2 E1ZV00 A0A2P8XCV9 E0VQ73 A0A1Y1JWC3 A0A1Y1JTV9 A0A0T6BEX0 A0A0A9XA88 A0A146LYF4 A0A146LUX9 A0A146M4X0 T1HGD1 A0A2R7VPD4 D6WX73 J9JWY3 A0A2S2P6G4 A0A1B6JBX6 A0A1B6HLI4 A0A0P8XY64 Q16VL0 A0A1S4FXP6 A0A0R1DJ30 Q16KU5 A0A0J9QVI0 A0A0R3NT85 A0A0Q5WID4 A0A0Q9WNB0 T1IVZ1 T1PI09 A0A0P8ZI66 A0A0P8XHV7 A0A0P9BRS4 B3MU39 B4NYJ8 A0A0R1DJ53 A0A0N8ES84 N6UJ90 U4UKU9 A0A0J9QVJ3 A0A0J9QVM0 A0A0Q9XEQ8 A0A182QD36 B0W7G1 A0A1I8Q6P6 A0A1I8Q6Q4 W8AQ00 A0A1I8MUR8 A0A1Q3FCQ9 A0A1Q3FCX4 A0A0Q5WLE1 B3N3R6 A0A1B6KK07 A0A1L8D957 A0A0M4EBF0 B4I385 Q7QFN1 W8B4C2 D4G7D1 A0A182JHX8 A0A182Y7H4
A0A212EPB0 A0A067QXM3 A0A2J7PXG9 A0A2J7PXI5 A0A2J7PXH2 A0A2J7PXH6 A0A2A3EAG4 A0A232ER02 A0A310SMW2 A0A0L7QQV6 A0A087ZXX4 E2B474 A0A3L8DJS8 A0A195CWZ6 F4WHR9 A0A195B5D9 A0A151JXP4 A0A158NM24 A0A0C9PL89 A0A151XIG3 A0A026WG49 A0A0C9RMK5 A0A0C9QMF3 A0A1B6EF44 A0A1B6DN27 A0A1B6BXT8 A0A0N0U4H9 A0A2J7PXH8 A0A224XDD0 A0A1B6BYR1 A0A1B6CCC8 A0A023F1Q2 E1ZV00 A0A2P8XCV9 E0VQ73 A0A1Y1JWC3 A0A1Y1JTV9 A0A0T6BEX0 A0A0A9XA88 A0A146LYF4 A0A146LUX9 A0A146M4X0 T1HGD1 A0A2R7VPD4 D6WX73 J9JWY3 A0A2S2P6G4 A0A1B6JBX6 A0A1B6HLI4 A0A0P8XY64 Q16VL0 A0A1S4FXP6 A0A0R1DJ30 Q16KU5 A0A0J9QVI0 A0A0R3NT85 A0A0Q5WID4 A0A0Q9WNB0 T1IVZ1 T1PI09 A0A0P8ZI66 A0A0P8XHV7 A0A0P9BRS4 B3MU39 B4NYJ8 A0A0R1DJ53 A0A0N8ES84 N6UJ90 U4UKU9 A0A0J9QVJ3 A0A0J9QVM0 A0A0Q9XEQ8 A0A182QD36 B0W7G1 A0A1I8Q6P6 A0A1I8Q6Q4 W8AQ00 A0A1I8MUR8 A0A1Q3FCQ9 A0A1Q3FCX4 A0A0Q5WLE1 B3N3R6 A0A1B6KK07 A0A1L8D957 A0A0M4EBF0 B4I385 Q7QFN1 W8B4C2 D4G7D1 A0A182JHX8 A0A182Y7H4
Pubmed
EMBL
NWSH01001832
PCG69963.1
BABH01020203
BABH01020204
BABH01020205
BABH01020206
+ More
RSAL01000090 RVE48102.1 KQ461155 KPJ08304.1 KC469893 AGC92690.2 FP102340 CBH09281.1 AGBW02013517 OWR43291.1 KK853134 KDR10862.1 NEVH01020858 PNF21038.1 PNF21040.1 PNF21041.1 PNF21037.1 KZ288308 PBC28715.1 NNAY01002685 OXU20778.1 KQ759847 OAD62585.1 KQ414784 KOC61013.1 GL445515 EFN89491.1 QOIP01000007 RLU20670.1 KQ977171 KYN05181.1 GL888167 EGI66276.1 KQ976604 KYM79404.1 KQ981561 KYN39827.1 ADTU01020204 GBYB01001768 JAG71535.1 KQ982109 KYQ60000.1 KK107238 EZA54621.1 GBYB01009580 JAG79347.1 GBYB01001767 JAG71534.1 GEDC01000814 JAS36484.1 GEDC01023274 GEDC01010258 JAS14024.1 JAS27040.1 GEDC01031434 GEDC01026722 JAS05864.1 JAS10576.1 KQ435834 KOX71551.1 PNF21039.1 GFTR01008638 JAW07788.1 GEDC01030912 GEDC01010908 JAS06386.1 JAS26390.1 GEDC01026348 JAS10950.1 GBBI01003783 JAC14929.1 GL434335 EFN74998.1 PYGN01002942 PSN29814.1 DS235392 EEB15529.1 GEZM01101576 GEZM01101573 GEZM01101570 GEZM01101569 GEZM01101566 GEZM01101565 GEZM01101553 GEZM01101549 GEZM01101547 JAV52508.1 GEZM01101567 GEZM01101556 JAV52523.1 LJIG01001147 KRT85793.1 GBHO01045650 GBHO01045645 GBHO01026686 GBHO01026683 GBHO01026682 GBHO01026681 GBHO01026680 GBHO01026678 JAF97953.1 JAF97958.1 JAG16918.1 JAG16921.1 JAG16922.1 JAG16923.1 JAG16924.1 JAG16926.1 GDHC01006927 JAQ11702.1 GDHC01007108 JAQ11521.1 GDHC01003801 JAQ14828.1 ACPB03015962 ACPB03015963 KK854009 PTY09219.1 KQ971361 EFA08800.1 ABLF02027300 GGMR01012391 MBY25010.1 GECU01011041 JAS96665.1 GECU01032144 JAS75562.1 CH902624 KPU74432.1 CH477589 EAT38586.1 CM000157 KRJ97269.1 CH477937 EAT34941.1 CM002910 KMY88057.1 CH379060 KRT04430.1 CH954177 KQS70089.1 CH940649 KRF81941.1 JH431601 KA648401 AFP63030.1 KPU74431.1 KPU74433.1 KPU74430.1 EDV33368.1 EDW87631.1 KRJ97268.1 GDUN01000178 JAN95741.1 APGK01022082 KB740398 ENN80731.1 KB632255 ERL90630.1 KMY88058.1 KMY88056.1 CH933807 KRG03524.1 AXCN02001394 DS231854 EDS37996.1 GAMC01018438 JAB88117.1 GFDL01009689 JAV25356.1 GFDL01009707 JAV25338.1 KQS70090.1 EDV57725.1 GEBQ01028201 JAT11776.1 GFDF01011093 JAV02991.1 CP012523 ALC38747.1 CH480820 EDW54230.1 AAAB01008846 EAA06333.5 GAMC01018439 JAB88116.1 BT122083 ADE06689.1 AXCP01007583
RSAL01000090 RVE48102.1 KQ461155 KPJ08304.1 KC469893 AGC92690.2 FP102340 CBH09281.1 AGBW02013517 OWR43291.1 KK853134 KDR10862.1 NEVH01020858 PNF21038.1 PNF21040.1 PNF21041.1 PNF21037.1 KZ288308 PBC28715.1 NNAY01002685 OXU20778.1 KQ759847 OAD62585.1 KQ414784 KOC61013.1 GL445515 EFN89491.1 QOIP01000007 RLU20670.1 KQ977171 KYN05181.1 GL888167 EGI66276.1 KQ976604 KYM79404.1 KQ981561 KYN39827.1 ADTU01020204 GBYB01001768 JAG71535.1 KQ982109 KYQ60000.1 KK107238 EZA54621.1 GBYB01009580 JAG79347.1 GBYB01001767 JAG71534.1 GEDC01000814 JAS36484.1 GEDC01023274 GEDC01010258 JAS14024.1 JAS27040.1 GEDC01031434 GEDC01026722 JAS05864.1 JAS10576.1 KQ435834 KOX71551.1 PNF21039.1 GFTR01008638 JAW07788.1 GEDC01030912 GEDC01010908 JAS06386.1 JAS26390.1 GEDC01026348 JAS10950.1 GBBI01003783 JAC14929.1 GL434335 EFN74998.1 PYGN01002942 PSN29814.1 DS235392 EEB15529.1 GEZM01101576 GEZM01101573 GEZM01101570 GEZM01101569 GEZM01101566 GEZM01101565 GEZM01101553 GEZM01101549 GEZM01101547 JAV52508.1 GEZM01101567 GEZM01101556 JAV52523.1 LJIG01001147 KRT85793.1 GBHO01045650 GBHO01045645 GBHO01026686 GBHO01026683 GBHO01026682 GBHO01026681 GBHO01026680 GBHO01026678 JAF97953.1 JAF97958.1 JAG16918.1 JAG16921.1 JAG16922.1 JAG16923.1 JAG16924.1 JAG16926.1 GDHC01006927 JAQ11702.1 GDHC01007108 JAQ11521.1 GDHC01003801 JAQ14828.1 ACPB03015962 ACPB03015963 KK854009 PTY09219.1 KQ971361 EFA08800.1 ABLF02027300 GGMR01012391 MBY25010.1 GECU01011041 JAS96665.1 GECU01032144 JAS75562.1 CH902624 KPU74432.1 CH477589 EAT38586.1 CM000157 KRJ97269.1 CH477937 EAT34941.1 CM002910 KMY88057.1 CH379060 KRT04430.1 CH954177 KQS70089.1 CH940649 KRF81941.1 JH431601 KA648401 AFP63030.1 KPU74431.1 KPU74433.1 KPU74430.1 EDV33368.1 EDW87631.1 KRJ97268.1 GDUN01000178 JAN95741.1 APGK01022082 KB740398 ENN80731.1 KB632255 ERL90630.1 KMY88058.1 KMY88056.1 CH933807 KRG03524.1 AXCN02001394 DS231854 EDS37996.1 GAMC01018438 JAB88117.1 GFDL01009689 JAV25356.1 GFDL01009707 JAV25338.1 KQS70090.1 EDV57725.1 GEBQ01028201 JAT11776.1 GFDF01011093 JAV02991.1 CP012523 ALC38747.1 CH480820 EDW54230.1 AAAB01008846 EAA06333.5 GAMC01018439 JAB88116.1 BT122083 ADE06689.1 AXCP01007583
Proteomes
UP000218220
UP000005204
UP000283053
UP000053240
UP000007151
UP000027135
+ More
UP000235965 UP000242457 UP000215335 UP000053825 UP000005203 UP000008237 UP000279307 UP000078542 UP000007755 UP000078540 UP000078541 UP000005205 UP000075809 UP000053097 UP000053105 UP000000311 UP000245037 UP000009046 UP000015103 UP000007266 UP000007819 UP000007801 UP000008820 UP000002282 UP000001819 UP000008711 UP000008792 UP000019118 UP000030742 UP000009192 UP000075886 UP000002320 UP000095300 UP000095301 UP000092553 UP000001292 UP000007062 UP000075880 UP000076408
UP000235965 UP000242457 UP000215335 UP000053825 UP000005203 UP000008237 UP000279307 UP000078542 UP000007755 UP000078540 UP000078541 UP000005205 UP000075809 UP000053097 UP000053105 UP000000311 UP000245037 UP000009046 UP000015103 UP000007266 UP000007819 UP000007801 UP000008820 UP000002282 UP000001819 UP000008711 UP000008792 UP000019118 UP000030742 UP000009192 UP000075886 UP000002320 UP000095300 UP000095301 UP000092553 UP000001292 UP000007062 UP000075880 UP000076408
Interpro
Gene 3D
ProteinModelPortal
A0A2A4JEV8
H9J7S0
A0A3S2L8H5
A0A194QSL0
L7X3W0
D0AB86
+ More
A0A212EPB0 A0A067QXM3 A0A2J7PXG9 A0A2J7PXI5 A0A2J7PXH2 A0A2J7PXH6 A0A2A3EAG4 A0A232ER02 A0A310SMW2 A0A0L7QQV6 A0A087ZXX4 E2B474 A0A3L8DJS8 A0A195CWZ6 F4WHR9 A0A195B5D9 A0A151JXP4 A0A158NM24 A0A0C9PL89 A0A151XIG3 A0A026WG49 A0A0C9RMK5 A0A0C9QMF3 A0A1B6EF44 A0A1B6DN27 A0A1B6BXT8 A0A0N0U4H9 A0A2J7PXH8 A0A224XDD0 A0A1B6BYR1 A0A1B6CCC8 A0A023F1Q2 E1ZV00 A0A2P8XCV9 E0VQ73 A0A1Y1JWC3 A0A1Y1JTV9 A0A0T6BEX0 A0A0A9XA88 A0A146LYF4 A0A146LUX9 A0A146M4X0 T1HGD1 A0A2R7VPD4 D6WX73 J9JWY3 A0A2S2P6G4 A0A1B6JBX6 A0A1B6HLI4 A0A0P8XY64 Q16VL0 A0A1S4FXP6 A0A0R1DJ30 Q16KU5 A0A0J9QVI0 A0A0R3NT85 A0A0Q5WID4 A0A0Q9WNB0 T1IVZ1 T1PI09 A0A0P8ZI66 A0A0P8XHV7 A0A0P9BRS4 B3MU39 B4NYJ8 A0A0R1DJ53 A0A0N8ES84 N6UJ90 U4UKU9 A0A0J9QVJ3 A0A0J9QVM0 A0A0Q9XEQ8 A0A182QD36 B0W7G1 A0A1I8Q6P6 A0A1I8Q6Q4 W8AQ00 A0A1I8MUR8 A0A1Q3FCQ9 A0A1Q3FCX4 A0A0Q5WLE1 B3N3R6 A0A1B6KK07 A0A1L8D957 A0A0M4EBF0 B4I385 Q7QFN1 W8B4C2 D4G7D1 A0A182JHX8 A0A182Y7H4
A0A212EPB0 A0A067QXM3 A0A2J7PXG9 A0A2J7PXI5 A0A2J7PXH2 A0A2J7PXH6 A0A2A3EAG4 A0A232ER02 A0A310SMW2 A0A0L7QQV6 A0A087ZXX4 E2B474 A0A3L8DJS8 A0A195CWZ6 F4WHR9 A0A195B5D9 A0A151JXP4 A0A158NM24 A0A0C9PL89 A0A151XIG3 A0A026WG49 A0A0C9RMK5 A0A0C9QMF3 A0A1B6EF44 A0A1B6DN27 A0A1B6BXT8 A0A0N0U4H9 A0A2J7PXH8 A0A224XDD0 A0A1B6BYR1 A0A1B6CCC8 A0A023F1Q2 E1ZV00 A0A2P8XCV9 E0VQ73 A0A1Y1JWC3 A0A1Y1JTV9 A0A0T6BEX0 A0A0A9XA88 A0A146LYF4 A0A146LUX9 A0A146M4X0 T1HGD1 A0A2R7VPD4 D6WX73 J9JWY3 A0A2S2P6G4 A0A1B6JBX6 A0A1B6HLI4 A0A0P8XY64 Q16VL0 A0A1S4FXP6 A0A0R1DJ30 Q16KU5 A0A0J9QVI0 A0A0R3NT85 A0A0Q5WID4 A0A0Q9WNB0 T1IVZ1 T1PI09 A0A0P8ZI66 A0A0P8XHV7 A0A0P9BRS4 B3MU39 B4NYJ8 A0A0R1DJ53 A0A0N8ES84 N6UJ90 U4UKU9 A0A0J9QVJ3 A0A0J9QVM0 A0A0Q9XEQ8 A0A182QD36 B0W7G1 A0A1I8Q6P6 A0A1I8Q6Q4 W8AQ00 A0A1I8MUR8 A0A1Q3FCQ9 A0A1Q3FCX4 A0A0Q5WLE1 B3N3R6 A0A1B6KK07 A0A1L8D957 A0A0M4EBF0 B4I385 Q7QFN1 W8B4C2 D4G7D1 A0A182JHX8 A0A182Y7H4
Ontologies
PANTHER
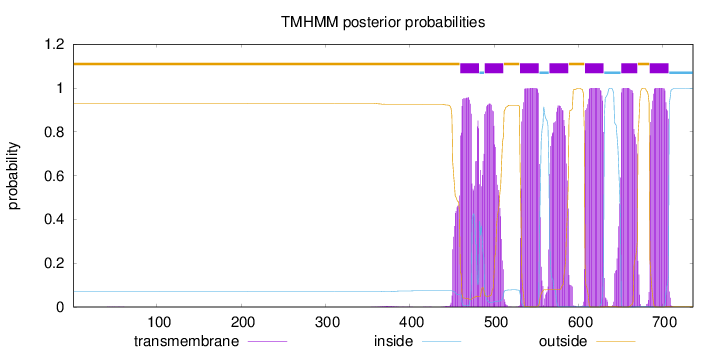
Topology
Length:
736
Number of predicted TMHs:
7
Exp number of AAs in TMHs:
151.93041
Exp number, first 60 AAs:
0.00697
Total prob of N-in:
0.07083
outside
1 - 459
TMhelix
460 - 482
inside
483 - 488
TMhelix
489 - 511
outside
512 - 530
TMhelix
531 - 553
inside
554 - 565
TMhelix
566 - 588
outside
589 - 607
TMhelix
608 - 630
inside
631 - 650
TMhelix
651 - 670
outside
671 - 684
TMhelix
685 - 707
inside
708 - 736
Population Genetic Test Statistics
Pi
277.131689
Theta
147.259594
Tajima's D
2.785931
CLR
0.105367
CSRT
0.967351632418379
Interpretation
Uncertain
