Gene
KWMTBOMO09944
Pre Gene Modal
BGIBMGA013241
Annotation
cytochrome_P450_CYP6AE7_[Bombyx_mori]
Location in the cell
PlasmaMembrane Reliability : 2.819
Sequence
CDS
ATGATATTAACTATAATATTCATTCTGAGTTTATGTGTTTACATTCTCTATAGGATATCTACAAGGAAATTTGATTATTGGCAAAAAAAGAATGTGAGCTACGTAGAGCCGGCTCCTTTCCTCGGAAACTACTCTGGGTACATACTTCTAAAGGAAAACTTATTGGACGTGGTGCATAATTTGAGTAAATTGTTTCCAAATGACCCCTACGTTGGTGCATTCTTTGGCACAAAACCCACTTTAATCGTTAAAGACCCGGAATTCATTAAACTGGTACTAATTAAAGATTTCTACTATTTCCACGGGAGAGAAGGCTCCAAATACACCCACAATGAAGTCATCACGCAGGGCATATTTTTCACGTACGGCGACCGCTGGAAGGTCATTCGACAAAACCTGACGCCGCTTTTCTCTTCTTCAAAAATTAGAAACATGTTTCGTACTATCGAAAAATGTTCTGGTGTTTTCGAGAATTTGCTGGATGAAGAATCTTTAGCACCCGAAGTTGAAATGAGGTCTGTGATGTCGCGGTTCACAATGGACTGCATAGGCGGTTGTGCGTTCGGAGTAGACACTAATGCGATGCAGGAACCCAAGGATAACATCTTCAAAACTATGGGATACCTTTTCTTCGAGAGCACCACGCACCGAGGGATAAAGAACGTTTTCAAAGCTATCTGGCCGGAAATATTCTATGGACTAGGTTTCAAAGTATTTCCAACGGACTTAAATGAGTTTTTTAGTAAACTTCTAGTTGGGATTTTCGAAGCACGGGACTACAAGCCTTCCTCACAGAATGTTTTCATCAACCTGTTGTTAAATTTGAAGAAGAACAGACATATCGTCGGCGACCGTTTGCTAAAGATTAAAACAGGAAACGTAAGAGCAGAGAGTAAAATAAAACTAGAAGTGGATGATGAGCTGTTAGTTTCGCAATGCGTTGCCTTCTTTATCGCAGGCTTTGAAACTTCATCCACTATTTCAAGCTTTACTTTGTACGAGCTGGCAAAGAATCCAGATGTTCAGAAAAGAGCGCAGAAAGAAGTGGACGAATACATTAAGAAGCACAACAATAAACTAGATTATGATTGCGTGAAGGAGCTCCCGTTTGTGGAGGCGTGTATAGATGAAGCCCTACGTTTGTATCCCGTTCTTGGAGTATTGACAAGAGAAGTAATGGAACAATACACATTCCCGACCGGCCTGACTCTCGACAAAGGAGATCGCGTTCATATTCCCGTGTATCATCTGCAACGCGACCCGGAATACTTTCCGGAGCCAGAGCTCTTTAAGCCGGAGAGGTTCTATGGGGAAGAGAAAAGGAATATCAGGCCCTTTACGTATTTACCATTTGGAGCTGGGCCCCGAACTTGTATTGGTCAAAGATTTGCGAAGATGCAGATGTTGGCCGGCCTGGTGACCATACTGAAAAGGTACACGGTGCGTCTGGCCGAGGGCATGCCGGAGACGATCAACTTCGAACAGAGAGCAATTGTCACGCAGCCCAACATTGGAATACGACTGAATCTCTTGCCTAGAAACAATTAA
Protein
MILTIIFILSLCVYILYRISTRKFDYWQKKNVSYVEPAPFLGNYSGYILLKENLLDVVHNLSKLFPNDPYVGAFFGTKPTLIVKDPEFIKLVLIKDFYYFHGREGSKYTHNEVITQGIFFTYGDRWKVIRQNLTPLFSSSKIRNMFRTIEKCSGVFENLLDEESLAPEVEMRSVMSRFTMDCIGGCAFGVDTNAMQEPKDNIFKTMGYLFFESTTHRGIKNVFKAIWPEIFYGLGFKVFPTDLNEFFSKLLVGIFEARDYKPSSQNVFINLLLNLKKNRHIVGDRLLKIKTGNVRAESKIKLEVDDELLVSQCVAFFIAGFETSSTISSFTLYELAKNPDVQKRAQKEVDEYIKKHNNKLDYDCVKELPFVEACIDEALRLYPVLGVLTREVMEQYTFPTGLTLDKGDRVHIPVYHLQRDPEYFPEPELFKPERFYGEEKRNIRPFTYLPFGAGPRTCIGQRFAKMQMLAGLVTILKRYTVRLAEGMPETINFEQRAIVTQPNIGIRLNLLPRNN
Summary
Cofactor
heme
Similarity
Belongs to the cytochrome P450 family.
Uniprot
H9JUM8
A4GUB8
H9JUM6
H9JUM5
D5L0M6
Q2XSW1
+ More
A0A068EVK6 A0A2A4J2E3 A0A172MF86 A0A068ETU8 A0A0L7LJH7 A0A068F0X2 A0A248QEX7 A0A248QHF4 A0A2A4JAK3 A0A068F1Z7 A0A248QEF8 A0A2H1V0E7 J7FJF2 A0A0L7KVL1 H2CZF1 A0A2H1WID4 A0A068ETU4 A0A068EWQ5 A0A2A4J7V4 H2CZE9 A0A2A4JAM9 B3VT94 C1LZ54 A0A2A4J698 A0A068EU65 F4YCW0 A0A0K0YD30 Q069I9 A0A068EU72 A0A0K8TUL3 A0A0C5C1I6 D6MLU9 A0A3S2P9Y7 A0A068EVL0 L0N7C5 A0A3S2LFF7 L0N766 A0A1V0D9D6 A0A0L7KJY7 A0A068EVD2 B6VFR9 A0A0A7DH57 A9YXV8 L0N6H5 A0A2A4J918 A3RIC0 H9JUM4 A9QW13 A0A068EWQ0 A0A1V0D9D9 Q7YZS2 A0A437B5H6 A5JM34 A5HG34 A4GUB9 A0A0L7LRY0 A9QW15 A5HG33 J7FKN1 A5HKM1 L0N6M3 A0A2H1V048 A0A0K8TUI1 A0A0A7DGX3 A0A0C5CGV6 A0A1V0D9E0 J7FKN2 A0A222NX20 A0A286QUG7 W5VY13 A0A411E5Q0 A0A0A7DH65 X5D9B5 A0A0N1ID06 A0A0N1IP03 A0A0K8TV92 C7U1L4 A0A3S2LW02 A0A0A7DH10 A0A248QG18 A0A2A4JWC6 A0A2W1BGY9 A0A2H1VU17 D2JLK6 A0A068F0X7 J7FJF3 A0A248QEH8 C1KJL7 A0A0A7DH14 D5L0M5 A0A2H1VSH4 A0A0A7DG86 A0A411AG09 A0A2H1VSG7
A0A068EVK6 A0A2A4J2E3 A0A172MF86 A0A068ETU8 A0A0L7LJH7 A0A068F0X2 A0A248QEX7 A0A248QHF4 A0A2A4JAK3 A0A068F1Z7 A0A248QEF8 A0A2H1V0E7 J7FJF2 A0A0L7KVL1 H2CZF1 A0A2H1WID4 A0A068ETU4 A0A068EWQ5 A0A2A4J7V4 H2CZE9 A0A2A4JAM9 B3VT94 C1LZ54 A0A2A4J698 A0A068EU65 F4YCW0 A0A0K0YD30 Q069I9 A0A068EU72 A0A0K8TUL3 A0A0C5C1I6 D6MLU9 A0A3S2P9Y7 A0A068EVL0 L0N7C5 A0A3S2LFF7 L0N766 A0A1V0D9D6 A0A0L7KJY7 A0A068EVD2 B6VFR9 A0A0A7DH57 A9YXV8 L0N6H5 A0A2A4J918 A3RIC0 H9JUM4 A9QW13 A0A068EWQ0 A0A1V0D9D9 Q7YZS2 A0A437B5H6 A5JM34 A5HG34 A4GUB9 A0A0L7LRY0 A9QW15 A5HG33 J7FKN1 A5HKM1 L0N6M3 A0A2H1V048 A0A0K8TUI1 A0A0A7DGX3 A0A0C5CGV6 A0A1V0D9E0 J7FKN2 A0A222NX20 A0A286QUG7 W5VY13 A0A411E5Q0 A0A0A7DH65 X5D9B5 A0A0N1ID06 A0A0N1IP03 A0A0K8TV92 C7U1L4 A0A3S2LW02 A0A0A7DH10 A0A248QG18 A0A2A4JWC6 A0A2W1BGY9 A0A2H1VU17 D2JLK6 A0A068F0X7 J7FJF3 A0A248QEH8 C1KJL7 A0A0A7DH14 D5L0M5 A0A2H1VSH4 A0A0A7DG86 A0A411AG09 A0A2H1VSG7
Pubmed
EMBL
BABH01022705
EF432656
ABO21682.1
BABH01022698
BABH01022694
GU731531
+ More
ADE05581.1 DQ256407 ABB69054.1 KM016736 AID54888.1 NWSH01003425 PCG66327.1 KU145393 ANC96676.1 KM016740 AID54892.1 JTDY01000942 KOB75351.1 KM016738 AID54890.1 KX443435 ASO98010.1 KX443432 ASO98007.1 NWSH01002369 PCG68452.1 KJ671576 AID55428.1 KX443430 ASO98005.1 ODYU01000082 SOQ34236.1 JX310078 AFP20589.1 JTDY01005369 KOB67074.1 JN375491 AEY75585.1 ODYU01008863 SOQ52829.1 KM016735 AID54887.1 KM016744 AID54896.1 NWSH01002736 PCG67604.1 JN375489 AEY75583.1 PCG68453.1 EU807990 ACF17813.2 FN356971 CAX94849.1 PCG67605.1 KM016737 AID54889.1 GU937509 ADW23116.1 KR095600 AKS48888.1 DQ986461 JN176574 ABI84381.1 AET11931.1 KM016742 AID54894.1 GCVX01000058 JAI18172.1 KP001126 AJN91171.1 GQ241737 ADD70250.1 RSAL01000158 RVE45548.1 KM016741 AID54893.1 AK289291 AK343204 BAM73818.1 BAM73890.1 RVE45547.1 AK343205 BAM73891.1 KY212045 ARA91609.1 JTDY01009617 KOB63214.1 KJ671575 AID55427.1 FJ387169 ACJ10074.1 KJ645906 AIJ00763.1 EU309477 ABY40426.1 AK289290 BAM73817.1 PCG68451.1 EF415296 ABN71367.1 EU273875 ABX64438.1 KM016739 AID54891.1 KY212043 ARA91607.1 AY295774 AAP83689.1 RVE45549.1 EF547545 ABQ42339.1 EF528486 ABP99020.1 EF432657 ABO21683.1 JTDY01000272 KOB77971.1 EU273877 ABX64440.1 EF528485 ABP99019.1 JX310077 AFP20588.1 EF535809 ABQ08711.1 AK343206 BAM73892.1 ODYU01000083 SOQ34238.1 GCVX01000081 JAI18149.1 KJ645907 AIJ00764.1 KP001127 AJN91172.1 KY212044 ARA91608.1 JX310082 AFP20593.1 KX844829 ASQ44127.1 KX443431 ASO98006.1 KF853191 AHH92929.1 MH236461 QBA57466.1 KJ645905 AIJ00762.1 KF701137 AHW57307.1 KQ459388 KPJ01135.1 KQ460655 KPJ12961.1 GCVX01000049 JAI18181.1 FN544262 CBB07053.1 RVE45546.1 KJ645904 AIJ00761.1 KX443433 ASO98008.1 NWSH01000502 PCG75998.1 KZ150141 PZC72944.1 ODYU01004424 SOQ44278.1 GQ915322 ACZ97416.2 KM016743 AID54895.1 JX310083 AFP20594.1 KX443434 ASO98009.1 FJ843077 ACO55746.1 KJ645909 AIJ00766.1 GU731530 ADE05580.1 ODYU01004193 SOQ43795.1 KJ645908 AIJ00765.1 MH151918 QAX33066.1 ODYU01004192 SOQ43793.1
ADE05581.1 DQ256407 ABB69054.1 KM016736 AID54888.1 NWSH01003425 PCG66327.1 KU145393 ANC96676.1 KM016740 AID54892.1 JTDY01000942 KOB75351.1 KM016738 AID54890.1 KX443435 ASO98010.1 KX443432 ASO98007.1 NWSH01002369 PCG68452.1 KJ671576 AID55428.1 KX443430 ASO98005.1 ODYU01000082 SOQ34236.1 JX310078 AFP20589.1 JTDY01005369 KOB67074.1 JN375491 AEY75585.1 ODYU01008863 SOQ52829.1 KM016735 AID54887.1 KM016744 AID54896.1 NWSH01002736 PCG67604.1 JN375489 AEY75583.1 PCG68453.1 EU807990 ACF17813.2 FN356971 CAX94849.1 PCG67605.1 KM016737 AID54889.1 GU937509 ADW23116.1 KR095600 AKS48888.1 DQ986461 JN176574 ABI84381.1 AET11931.1 KM016742 AID54894.1 GCVX01000058 JAI18172.1 KP001126 AJN91171.1 GQ241737 ADD70250.1 RSAL01000158 RVE45548.1 KM016741 AID54893.1 AK289291 AK343204 BAM73818.1 BAM73890.1 RVE45547.1 AK343205 BAM73891.1 KY212045 ARA91609.1 JTDY01009617 KOB63214.1 KJ671575 AID55427.1 FJ387169 ACJ10074.1 KJ645906 AIJ00763.1 EU309477 ABY40426.1 AK289290 BAM73817.1 PCG68451.1 EF415296 ABN71367.1 EU273875 ABX64438.1 KM016739 AID54891.1 KY212043 ARA91607.1 AY295774 AAP83689.1 RVE45549.1 EF547545 ABQ42339.1 EF528486 ABP99020.1 EF432657 ABO21683.1 JTDY01000272 KOB77971.1 EU273877 ABX64440.1 EF528485 ABP99019.1 JX310077 AFP20588.1 EF535809 ABQ08711.1 AK343206 BAM73892.1 ODYU01000083 SOQ34238.1 GCVX01000081 JAI18149.1 KJ645907 AIJ00764.1 KP001127 AJN91172.1 KY212044 ARA91608.1 JX310082 AFP20593.1 KX844829 ASQ44127.1 KX443431 ASO98006.1 KF853191 AHH92929.1 MH236461 QBA57466.1 KJ645905 AIJ00762.1 KF701137 AHW57307.1 KQ459388 KPJ01135.1 KQ460655 KPJ12961.1 GCVX01000049 JAI18181.1 FN544262 CBB07053.1 RVE45546.1 KJ645904 AIJ00761.1 KX443433 ASO98008.1 NWSH01000502 PCG75998.1 KZ150141 PZC72944.1 ODYU01004424 SOQ44278.1 GQ915322 ACZ97416.2 KM016743 AID54895.1 JX310083 AFP20594.1 KX443434 ASO98009.1 FJ843077 ACO55746.1 KJ645909 AIJ00766.1 GU731530 ADE05580.1 ODYU01004193 SOQ43795.1 KJ645908 AIJ00765.1 MH151918 QAX33066.1 ODYU01004192 SOQ43793.1
Proteomes
PRIDE
Interpro
SUPFAM
SSF48264
SSF48264
Gene 3D
ProteinModelPortal
H9JUM8
A4GUB8
H9JUM6
H9JUM5
D5L0M6
Q2XSW1
+ More
A0A068EVK6 A0A2A4J2E3 A0A172MF86 A0A068ETU8 A0A0L7LJH7 A0A068F0X2 A0A248QEX7 A0A248QHF4 A0A2A4JAK3 A0A068F1Z7 A0A248QEF8 A0A2H1V0E7 J7FJF2 A0A0L7KVL1 H2CZF1 A0A2H1WID4 A0A068ETU4 A0A068EWQ5 A0A2A4J7V4 H2CZE9 A0A2A4JAM9 B3VT94 C1LZ54 A0A2A4J698 A0A068EU65 F4YCW0 A0A0K0YD30 Q069I9 A0A068EU72 A0A0K8TUL3 A0A0C5C1I6 D6MLU9 A0A3S2P9Y7 A0A068EVL0 L0N7C5 A0A3S2LFF7 L0N766 A0A1V0D9D6 A0A0L7KJY7 A0A068EVD2 B6VFR9 A0A0A7DH57 A9YXV8 L0N6H5 A0A2A4J918 A3RIC0 H9JUM4 A9QW13 A0A068EWQ0 A0A1V0D9D9 Q7YZS2 A0A437B5H6 A5JM34 A5HG34 A4GUB9 A0A0L7LRY0 A9QW15 A5HG33 J7FKN1 A5HKM1 L0N6M3 A0A2H1V048 A0A0K8TUI1 A0A0A7DGX3 A0A0C5CGV6 A0A1V0D9E0 J7FKN2 A0A222NX20 A0A286QUG7 W5VY13 A0A411E5Q0 A0A0A7DH65 X5D9B5 A0A0N1ID06 A0A0N1IP03 A0A0K8TV92 C7U1L4 A0A3S2LW02 A0A0A7DH10 A0A248QG18 A0A2A4JWC6 A0A2W1BGY9 A0A2H1VU17 D2JLK6 A0A068F0X7 J7FJF3 A0A248QEH8 C1KJL7 A0A0A7DH14 D5L0M5 A0A2H1VSH4 A0A0A7DG86 A0A411AG09 A0A2H1VSG7
A0A068EVK6 A0A2A4J2E3 A0A172MF86 A0A068ETU8 A0A0L7LJH7 A0A068F0X2 A0A248QEX7 A0A248QHF4 A0A2A4JAK3 A0A068F1Z7 A0A248QEF8 A0A2H1V0E7 J7FJF2 A0A0L7KVL1 H2CZF1 A0A2H1WID4 A0A068ETU4 A0A068EWQ5 A0A2A4J7V4 H2CZE9 A0A2A4JAM9 B3VT94 C1LZ54 A0A2A4J698 A0A068EU65 F4YCW0 A0A0K0YD30 Q069I9 A0A068EU72 A0A0K8TUL3 A0A0C5C1I6 D6MLU9 A0A3S2P9Y7 A0A068EVL0 L0N7C5 A0A3S2LFF7 L0N766 A0A1V0D9D6 A0A0L7KJY7 A0A068EVD2 B6VFR9 A0A0A7DH57 A9YXV8 L0N6H5 A0A2A4J918 A3RIC0 H9JUM4 A9QW13 A0A068EWQ0 A0A1V0D9D9 Q7YZS2 A0A437B5H6 A5JM34 A5HG34 A4GUB9 A0A0L7LRY0 A9QW15 A5HG33 J7FKN1 A5HKM1 L0N6M3 A0A2H1V048 A0A0K8TUI1 A0A0A7DGX3 A0A0C5CGV6 A0A1V0D9E0 J7FKN2 A0A222NX20 A0A286QUG7 W5VY13 A0A411E5Q0 A0A0A7DH65 X5D9B5 A0A0N1ID06 A0A0N1IP03 A0A0K8TV92 C7U1L4 A0A3S2LW02 A0A0A7DH10 A0A248QG18 A0A2A4JWC6 A0A2W1BGY9 A0A2H1VU17 D2JLK6 A0A068F0X7 J7FJF3 A0A248QEH8 C1KJL7 A0A0A7DH14 D5L0M5 A0A2H1VSH4 A0A0A7DG86 A0A411AG09 A0A2H1VSG7
PDB
6MJM
E-value=5.86535e-53,
Score=526
Ontologies
KEGG
GO
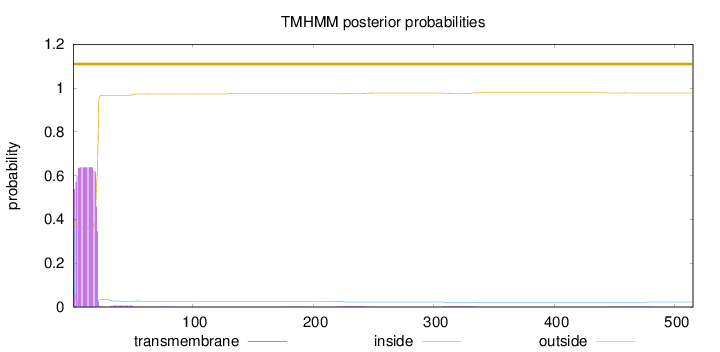
Topology
Length:
515
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
12.34126
Exp number, first 60 AAs:
12.17863
Total prob of N-in:
0.60497
POSSIBLE N-term signal
sequence
outside
1 - 515
Population Genetic Test Statistics
Pi
260.270549
Theta
186.291388
Tajima's D
1.039487
CLR
0
CSRT
0.669566521673916
Interpretation
Uncertain
