Gene
KWMTBOMO09854
Pre Gene Modal
BGIBMGA013151
Annotation
PREDICTED:_fatty_acid_synthase-like_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 2.255
Sequence
CDS
ATGGCCCCTACACCTCAAGACAACAAGGTACTAGAAGATGTACCGAAGCAACCTGTCATCGGCGAAAGGGTTGTAATTTCCGGAATGTCTGGCCTGTACCCTGACTGTAAATCTGTTAAGGATTTGGCAGATATTTTGTATAATAAGAAAGATCCTATAACATCCGAGCGTCAACGTTGGAGTTATTCCCACCCTGAAGTGTCACAATGCACTGGAAAGCTCCTTGATCTTGACCGTTTCGATGCTCAATTCTTCAAGGTGCATTTTCGTCTTGGAAACAGCATGGATTCAATGGGAAGAAAAGTGCTGGAGCAAACATATCAAGCAATATATGATGCCGGTGTTAATCCACAACATCTTGCTGGAAAAAAAGTAGGAGTGTACATTGGGACATGTTTCTCTGAAACTGAAAAGACTAGCTTTTATGTAGCTAACTCTAGGACAGGTTTTGGCATTGCAGGGTGCAGCAAAACTATGTTTGCAAATCGTATTTCATACTGGTTAAATGTCAAAGGACCGTCGATGGCAATAGACGAAGCCTGTTGTAGCTCTACTGCAGCTATTGAACAAGCCTACCTCGCCATGACTCGAGGGGAATGCGAAGCGGCTATCGTCGGAGGAACTAACATATGCCTTCATCCACATTCCTTGATTCATCACGGAAGGCTCTTGACTATTTGCATGGATGCGAAAACGAAATCCTTCGATCAAAACGCAGATGGTTGTGCCAAATCGGAAGCTATAAACGTTTTATTTCTTCAGAAGGCCGAAAATGCGCTCAGAGTTTACGCAGAAATTGTTTTTGTGAAAACAGAATTTACGGCTTTACCAAATGCCGTAACTGGACCAAAGTTTGGATTTTGTCGTGATCCAGATGTCATAGCGAATTTCCTAAAGAAGTTGTATGAAAATGCTGGAGTGTCACCTGATGCTGTTGAATACGTAGAAGCTTTTGGGTCAGGTGAATAA
Protein
MAPTPQDNKVLEDVPKQPVIGERVVISGMSGLYPDCKSVKDLADILYNKKDPITSERQRWSYSHPEVSQCTGKLLDLDRFDAQFFKVHFRLGNSMDSMGRKVLEQTYQAIYDAGVNPQHLAGKKVGVYIGTCFSETEKTSFYVANSRTGFGIAGCSKTMFANRISYWLNVKGPSMAIDEACCSSTAAIEQAYLAMTRGECEAAIVGGTNICLHPHSLIHHGRLLTICMDAKTKSFDQNADGCAKSEAINVLFLQKAENALRVYAEIVFVKTEFTALPNAVTGPKFGFCRDPDVIANFLKKLYENAGVSPDAVEYVEAFGSGE
Summary
Similarity
Belongs to the thiolase-like superfamily. Beta-ketoacyl-ACP synthases family.
Uniprot
H9JUD8
A0A2H1WMZ3
A0A0L7L3I7
A0A2H1WNC5
A0A212EWU6
A0A0N1I4Z6
+ More
A0A2A4K3Y5 A0A2H1V636 A0A194PKD7 A0A2A4K4K9 A0A0L7L2S3 A0A2A4K3Y0 A0A3S2P698 A0A0L7L2W8 A0A0L7KRD1 A0A437B1L3 A0A3S2NJM7 O01677 A0A0L7K4N3 A0A0L7LI88 A0A2W1BEL1 A0A2A4JTZ4 A0A1S4EB00 A0A1B6CGM1 A0A0C9Q3R4 E9IT55 A0A1B6GA85 A0A212FMU5 A0A3S2LA71 A0A195CP13 A0A195BMY6 A0A158NZW8 E0VU95 A0A2H1VGF0 A0A195F360 A0A067RHL3 A0A1B6IHH2 A0A1B6JP24 O01678 A0A2A4J2W6 A0A2A4J169 E2B583 A0A232EXC2 A0A0N1IP09 A0A2R7W720 A0A2H8TWM7 K7IZC8 F4WIF1 A0A2S2RA89 A0A151JRB8 J9K3D3 A0A0G3YL52 A0A194Q6Q7 A0A026WNV9 E2ADN8 A0A2H1VF66 A0A2J7PQX9 A0A232F4D3 A0A2A4K5B4 K7IZC9 A0A154PMH1 A0A1A9XHP8 A0A1B0G1J3 A0A2H1VFB6 A0A2P8XKN7 A0A1W4XV87 A0A1A9XHQ3 A0A1B0AA77 A0A1B6LEY2 A0A088ALX8 A0A182GNP7 A0A2A3E4Z2 H9JGD2 A0A1S4FIT0 Q16ZI9 A0A1B0B9K0 A0A1A9W891 A0A0L0BQ63 E1ZXI7 E2AUN4 A0A310SRI2 B0WMC9 A0A1B0G6Y6 A0A2J7PQX7 A0A0L7LNK4 Q7PLB8 T1P9U1 A0A1I8N1J7 A0A1A9W883 B5DVP9 A0A0K8UG84 A0A0M8ZRC2 A0A0R1E948 B3N4Q4 A0A0L7R328 W5J299 A0A1Q3FS82
A0A2A4K3Y5 A0A2H1V636 A0A194PKD7 A0A2A4K4K9 A0A0L7L2S3 A0A2A4K3Y0 A0A3S2P698 A0A0L7L2W8 A0A0L7KRD1 A0A437B1L3 A0A3S2NJM7 O01677 A0A0L7K4N3 A0A0L7LI88 A0A2W1BEL1 A0A2A4JTZ4 A0A1S4EB00 A0A1B6CGM1 A0A0C9Q3R4 E9IT55 A0A1B6GA85 A0A212FMU5 A0A3S2LA71 A0A195CP13 A0A195BMY6 A0A158NZW8 E0VU95 A0A2H1VGF0 A0A195F360 A0A067RHL3 A0A1B6IHH2 A0A1B6JP24 O01678 A0A2A4J2W6 A0A2A4J169 E2B583 A0A232EXC2 A0A0N1IP09 A0A2R7W720 A0A2H8TWM7 K7IZC8 F4WIF1 A0A2S2RA89 A0A151JRB8 J9K3D3 A0A0G3YL52 A0A194Q6Q7 A0A026WNV9 E2ADN8 A0A2H1VF66 A0A2J7PQX9 A0A232F4D3 A0A2A4K5B4 K7IZC9 A0A154PMH1 A0A1A9XHP8 A0A1B0G1J3 A0A2H1VFB6 A0A2P8XKN7 A0A1W4XV87 A0A1A9XHQ3 A0A1B0AA77 A0A1B6LEY2 A0A088ALX8 A0A182GNP7 A0A2A3E4Z2 H9JGD2 A0A1S4FIT0 Q16ZI9 A0A1B0B9K0 A0A1A9W891 A0A0L0BQ63 E1ZXI7 E2AUN4 A0A310SRI2 B0WMC9 A0A1B0G6Y6 A0A2J7PQX7 A0A0L7LNK4 Q7PLB8 T1P9U1 A0A1I8N1J7 A0A1A9W883 B5DVP9 A0A0K8UG84 A0A0M8ZRC2 A0A0R1E948 B3N4Q4 A0A0L7R328 W5J299 A0A1Q3FS82
Pubmed
EMBL
BABH01022903
ODYU01009786
SOQ54441.1
JTDY01003263
KOB69849.1
ODYU01009787
+ More
SOQ54442.1 AGBW02011888 OWR45966.1 KQ461178 KPJ07769.1 NWSH01000156 PCG78967.1 ODYU01000872 SOQ36288.1 KQ459602 KPI93194.1 PCG78966.1 JTDY01003390 KOB69631.1 PCG78965.1 RSAL01000013 RVE53442.1 KOB69848.1 JTDY01006902 KOB65606.1 RSAL01000223 RVE44005.1 RSAL01000068 RVE49259.1 U67866 AAB53257.1 JTDY01011546 KOB55720.1 JTDY01001031 KOB75079.1 KZ150116 PZC73268.1 NWSH01000591 PCG75471.1 GEDC01024813 JAS12485.1 GBYB01008708 JAG78475.1 GL765434 EFZ16301.1 GECZ01010430 JAS59339.1 AGBW02007651 OWR55047.1 RSAL01000071 RVE49124.1 KQ977574 KYN01849.1 KQ976440 KYM86527.1 ADTU01005072 ADTU01005073 ADTU01005074 ADTU01005075 DS235784 EEB16951.1 ODYU01002246 SOQ39472.1 KQ981856 KYN34617.1 KK852680 KDR18643.1 GECU01021338 JAS86368.1 GECU01006695 JAT01012.1 U67867 AAB53258.1 NWSH01003873 PCG65733.1 PCG65731.1 GL445739 EFN89136.1 NNAY01001760 OXU23005.1 KQ460651 KPJ13003.1 KK854269 PTY14005.1 GFXV01006890 MBW18695.1 GL888175 EGI66082.1 GGMS01017661 MBY86864.1 KQ978607 KYN29892.1 ABLF02036897 KM507100 AKM28424.1 KQ459582 KPI99075.1 KK107139 EZA57700.1 GL438820 EFN68400.1 ODYU01002244 SOQ39469.1 NEVH01022635 PNF18739.1 NNAY01001038 OXU25389.1 PCG78963.1 KQ434960 KZC12654.1 CCAG010017947 ODYU01002245 SOQ39471.1 PYGN01001841 PSN32561.1 GEBQ01017893 JAT22084.1 JXUM01076840 KQ562972 KXJ74732.1 KZ288370 PBC26760.1 BABH01033622 BABH01033623 BABH01033624 CH477489 EAT40085.1 JXJN01010482 JXJN01010483 JRES01001625 KNC21349.1 GL435066 EFN74070.1 GL442834 EFN62855.1 KQ761246 OAD57941.1 DS231997 EDS30986.1 CCAG010013878 PNF18741.1 JTDY01000529 KOB76786.1 AE014296 EAA46042.3 KA644718 AFP59347.1 CM000070 EDY68457.3 GDHF01026736 JAI25578.1 KQ435885 KOX69550.1 CH893166 KRK05689.1 CH954177 EDV60008.2 KQ414663 KOC65238.1 ADMH02002134 ETN58382.1 GFDL01004709 JAV30336.1
SOQ54442.1 AGBW02011888 OWR45966.1 KQ461178 KPJ07769.1 NWSH01000156 PCG78967.1 ODYU01000872 SOQ36288.1 KQ459602 KPI93194.1 PCG78966.1 JTDY01003390 KOB69631.1 PCG78965.1 RSAL01000013 RVE53442.1 KOB69848.1 JTDY01006902 KOB65606.1 RSAL01000223 RVE44005.1 RSAL01000068 RVE49259.1 U67866 AAB53257.1 JTDY01011546 KOB55720.1 JTDY01001031 KOB75079.1 KZ150116 PZC73268.1 NWSH01000591 PCG75471.1 GEDC01024813 JAS12485.1 GBYB01008708 JAG78475.1 GL765434 EFZ16301.1 GECZ01010430 JAS59339.1 AGBW02007651 OWR55047.1 RSAL01000071 RVE49124.1 KQ977574 KYN01849.1 KQ976440 KYM86527.1 ADTU01005072 ADTU01005073 ADTU01005074 ADTU01005075 DS235784 EEB16951.1 ODYU01002246 SOQ39472.1 KQ981856 KYN34617.1 KK852680 KDR18643.1 GECU01021338 JAS86368.1 GECU01006695 JAT01012.1 U67867 AAB53258.1 NWSH01003873 PCG65733.1 PCG65731.1 GL445739 EFN89136.1 NNAY01001760 OXU23005.1 KQ460651 KPJ13003.1 KK854269 PTY14005.1 GFXV01006890 MBW18695.1 GL888175 EGI66082.1 GGMS01017661 MBY86864.1 KQ978607 KYN29892.1 ABLF02036897 KM507100 AKM28424.1 KQ459582 KPI99075.1 KK107139 EZA57700.1 GL438820 EFN68400.1 ODYU01002244 SOQ39469.1 NEVH01022635 PNF18739.1 NNAY01001038 OXU25389.1 PCG78963.1 KQ434960 KZC12654.1 CCAG010017947 ODYU01002245 SOQ39471.1 PYGN01001841 PSN32561.1 GEBQ01017893 JAT22084.1 JXUM01076840 KQ562972 KXJ74732.1 KZ288370 PBC26760.1 BABH01033622 BABH01033623 BABH01033624 CH477489 EAT40085.1 JXJN01010482 JXJN01010483 JRES01001625 KNC21349.1 GL435066 EFN74070.1 GL442834 EFN62855.1 KQ761246 OAD57941.1 DS231997 EDS30986.1 CCAG010013878 PNF18741.1 JTDY01000529 KOB76786.1 AE014296 EAA46042.3 KA644718 AFP59347.1 CM000070 EDY68457.3 GDHF01026736 JAI25578.1 KQ435885 KOX69550.1 CH893166 KRK05689.1 CH954177 EDV60008.2 KQ414663 KOC65238.1 ADMH02002134 ETN58382.1 GFDL01004709 JAV30336.1
Proteomes
UP000005204
UP000037510
UP000007151
UP000053240
UP000218220
UP000053268
+ More
UP000283053 UP000079169 UP000078542 UP000078540 UP000005205 UP000009046 UP000078541 UP000027135 UP000008237 UP000215335 UP000002358 UP000007755 UP000078492 UP000007819 UP000053097 UP000000311 UP000235965 UP000076502 UP000092443 UP000092444 UP000245037 UP000192223 UP000092445 UP000005203 UP000069940 UP000249989 UP000242457 UP000008820 UP000092460 UP000091820 UP000037069 UP000002320 UP000000803 UP000095301 UP000001819 UP000053105 UP000002282 UP000008711 UP000053825 UP000000673
UP000283053 UP000079169 UP000078542 UP000078540 UP000005205 UP000009046 UP000078541 UP000027135 UP000008237 UP000215335 UP000002358 UP000007755 UP000078492 UP000007819 UP000053097 UP000000311 UP000235965 UP000076502 UP000092443 UP000092444 UP000245037 UP000192223 UP000092445 UP000005203 UP000069940 UP000249989 UP000242457 UP000008820 UP000092460 UP000091820 UP000037069 UP000002320 UP000000803 UP000095301 UP000001819 UP000053105 UP000002282 UP000008711 UP000053825 UP000000673
Pfam
Interpro
IPR020801
PKS_acyl_transferase
+ More
IPR014030 Ketoacyl_synth_N
IPR016039 Thiolase-like
IPR001227 Ac_transferase_dom_sf
IPR020841 PKS_Beta-ketoAc_synthase_dom
IPR016035 Acyl_Trfase/lysoPLipase
IPR014031 Ketoacyl_synth_C
IPR016036 Malonyl_transacylase_ACP-bd
IPR014043 Acyl_transferase
IPR042104 PKS_dehydratase_sf
IPR032821 KAsynt_C_assoc
IPR020843 PKS_ER
IPR036291 NAD(P)-bd_dom_sf
IPR011032 GroES-like_sf
IPR013149 ADH_C
IPR023102 Fatty_acid_synthase_dom_2
IPR029058 AB_hydrolase
IPR020807 PKS_dehydratase
IPR013968 PKS_KR
IPR001031 Thioesterase
IPR018201 Ketoacyl_synth_AS
IPR036736 ACP-like_sf
IPR009081 PP-bd_ACP
IPR020806 PKS_PP-bd
IPR020846 MFS_dom
IPR036259 MFS_trans_sf
IPR011701 MFS
IPR006162 Ppantetheine_attach_site
IPR014030 Ketoacyl_synth_N
IPR016039 Thiolase-like
IPR001227 Ac_transferase_dom_sf
IPR020841 PKS_Beta-ketoAc_synthase_dom
IPR016035 Acyl_Trfase/lysoPLipase
IPR014031 Ketoacyl_synth_C
IPR016036 Malonyl_transacylase_ACP-bd
IPR014043 Acyl_transferase
IPR042104 PKS_dehydratase_sf
IPR032821 KAsynt_C_assoc
IPR020843 PKS_ER
IPR036291 NAD(P)-bd_dom_sf
IPR011032 GroES-like_sf
IPR013149 ADH_C
IPR023102 Fatty_acid_synthase_dom_2
IPR029058 AB_hydrolase
IPR020807 PKS_dehydratase
IPR013968 PKS_KR
IPR001031 Thioesterase
IPR018201 Ketoacyl_synth_AS
IPR036736 ACP-like_sf
IPR009081 PP-bd_ACP
IPR020806 PKS_PP-bd
IPR020846 MFS_dom
IPR036259 MFS_trans_sf
IPR011701 MFS
IPR006162 Ppantetheine_attach_site
SUPFAM
CDD
ProteinModelPortal
H9JUD8
A0A2H1WMZ3
A0A0L7L3I7
A0A2H1WNC5
A0A212EWU6
A0A0N1I4Z6
+ More
A0A2A4K3Y5 A0A2H1V636 A0A194PKD7 A0A2A4K4K9 A0A0L7L2S3 A0A2A4K3Y0 A0A3S2P698 A0A0L7L2W8 A0A0L7KRD1 A0A437B1L3 A0A3S2NJM7 O01677 A0A0L7K4N3 A0A0L7LI88 A0A2W1BEL1 A0A2A4JTZ4 A0A1S4EB00 A0A1B6CGM1 A0A0C9Q3R4 E9IT55 A0A1B6GA85 A0A212FMU5 A0A3S2LA71 A0A195CP13 A0A195BMY6 A0A158NZW8 E0VU95 A0A2H1VGF0 A0A195F360 A0A067RHL3 A0A1B6IHH2 A0A1B6JP24 O01678 A0A2A4J2W6 A0A2A4J169 E2B583 A0A232EXC2 A0A0N1IP09 A0A2R7W720 A0A2H8TWM7 K7IZC8 F4WIF1 A0A2S2RA89 A0A151JRB8 J9K3D3 A0A0G3YL52 A0A194Q6Q7 A0A026WNV9 E2ADN8 A0A2H1VF66 A0A2J7PQX9 A0A232F4D3 A0A2A4K5B4 K7IZC9 A0A154PMH1 A0A1A9XHP8 A0A1B0G1J3 A0A2H1VFB6 A0A2P8XKN7 A0A1W4XV87 A0A1A9XHQ3 A0A1B0AA77 A0A1B6LEY2 A0A088ALX8 A0A182GNP7 A0A2A3E4Z2 H9JGD2 A0A1S4FIT0 Q16ZI9 A0A1B0B9K0 A0A1A9W891 A0A0L0BQ63 E1ZXI7 E2AUN4 A0A310SRI2 B0WMC9 A0A1B0G6Y6 A0A2J7PQX7 A0A0L7LNK4 Q7PLB8 T1P9U1 A0A1I8N1J7 A0A1A9W883 B5DVP9 A0A0K8UG84 A0A0M8ZRC2 A0A0R1E948 B3N4Q4 A0A0L7R328 W5J299 A0A1Q3FS82
A0A2A4K3Y5 A0A2H1V636 A0A194PKD7 A0A2A4K4K9 A0A0L7L2S3 A0A2A4K3Y0 A0A3S2P698 A0A0L7L2W8 A0A0L7KRD1 A0A437B1L3 A0A3S2NJM7 O01677 A0A0L7K4N3 A0A0L7LI88 A0A2W1BEL1 A0A2A4JTZ4 A0A1S4EB00 A0A1B6CGM1 A0A0C9Q3R4 E9IT55 A0A1B6GA85 A0A212FMU5 A0A3S2LA71 A0A195CP13 A0A195BMY6 A0A158NZW8 E0VU95 A0A2H1VGF0 A0A195F360 A0A067RHL3 A0A1B6IHH2 A0A1B6JP24 O01678 A0A2A4J2W6 A0A2A4J169 E2B583 A0A232EXC2 A0A0N1IP09 A0A2R7W720 A0A2H8TWM7 K7IZC8 F4WIF1 A0A2S2RA89 A0A151JRB8 J9K3D3 A0A0G3YL52 A0A194Q6Q7 A0A026WNV9 E2ADN8 A0A2H1VF66 A0A2J7PQX9 A0A232F4D3 A0A2A4K5B4 K7IZC9 A0A154PMH1 A0A1A9XHP8 A0A1B0G1J3 A0A2H1VFB6 A0A2P8XKN7 A0A1W4XV87 A0A1A9XHQ3 A0A1B0AA77 A0A1B6LEY2 A0A088ALX8 A0A182GNP7 A0A2A3E4Z2 H9JGD2 A0A1S4FIT0 Q16ZI9 A0A1B0B9K0 A0A1A9W891 A0A0L0BQ63 E1ZXI7 E2AUN4 A0A310SRI2 B0WMC9 A0A1B0G6Y6 A0A2J7PQX7 A0A0L7LNK4 Q7PLB8 T1P9U1 A0A1I8N1J7 A0A1A9W883 B5DVP9 A0A0K8UG84 A0A0M8ZRC2 A0A0R1E948 B3N4Q4 A0A0L7R328 W5J299 A0A1Q3FS82
PDB
3HHD
E-value=1.80201e-57,
Score=563
Ontologies
GO
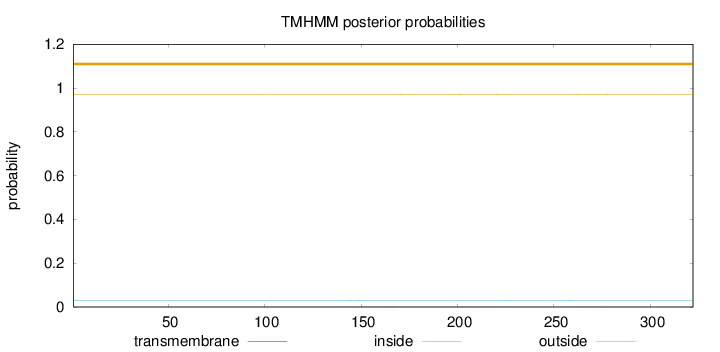
Topology
Length:
322
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.01605
Exp number, first 60 AAs:
0.00015
Total prob of N-in:
0.03057
outside
1 - 322
Population Genetic Test Statistics
Pi
189.296887
Theta
149.933981
Tajima's D
0.246215
CLR
0.212398
CSRT
0.437778111094445
Interpretation
Possibly Positive selection
