Gene
KWMTBOMO09584
Pre Gene Modal
BGIBMGA013001
Annotation
PREDICTED:_glucose_dehydrogenase_[FAD?_quinone]_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 1.445
Sequence
CDS
ATGGCGATTTCCACGGTTGCAGCTACAGCCATTAAATCAGCTCTTGCCATCGGCGGTCTGCATACACTTACTTTCATACCGATAATGCTAGCGGCCATGGCCTATTTCAACTATGAACTACTAGATCCCGAACAAAGACCGTTCAATCAGAAATATTTGAGAGAAACATATGATTTCATAGTGGTCGGAGGAGGATCTGCCGGATCTGTATTAGCTAATAGACTGTCTGAAGTTGAAGGTTGGAATGTCTTGCTTTTAGAAGCAGGAGGTCACGAAACCGACATCAGTGATGTTCCCTTACTGTCATTGTATTTGCATAAGAGCAAACTTGACTGGAAATACCGGACCCAACCGCAAGACACTGCTTGCCAAGCTATGATCGACAAACGATGCAGCTGGACTAAAGGCAAAGTCTTGGGAGGCTCTTCAGTTCTCAATACCATGTTGTATATAAGAGGGAACAAGCGTGATTTCGATCAGTGGGAGTCCTTTGGAAACCCGGGGTGGGGCTATGAAGATGTCTTGCCGTATTTCAAGAAATCTGAGGACCAACGCAATCCGTACTTGGCAAAGGATACCAAGTATCATCAAACTGGCGGGTATCTAACTGTCCAAGACTCCCCGTATAATACTCCGATCGGAGCTGCGCTACTCCAAGCTGGTGAGGAAATGGGATATGACTTTATCGACGTAAACGGAGCTCAACAAACTGGATATGCGTGGTACCAGTTTACTATTCGAAGAGGCACCAGATGTTCCACAGCAAAAGCTTTCCTGAGACCTGTAAGACTACGACAGAACCTTCATATCGCTCTATTCTCCCACGTCACTAAAGTTTTAATTGACAAGGATACGAAAAGAGCATACGGAGTCGAATTCCTTAGAGATGGCACTCAGCAAGTCGTTTATGCAAAACGAGAAGTTATATTGGCGGCTGGAGCAATAGCTTCGCCTCAATTACTTATGTTGTCTGGTGTCGGACCCGAGCAACACCTAAAAGAGGTGGGGATCGATGTGATTCATGATTCTCCTGGAGTCGGGAGAAACCTGCAAGATCACATCGCTGTCGGAGGCATTATCTTTCGAATCGATTATCCCGTAAGTTTGGTTATGAATCGTCTTGTGAACATTAACTCTGCTTTACGGTATGCCATCACCGAGGACGGTCCACTTACATCCAGCATCGGCCTCGAAGTTGTAGCTTTTATTAATACTAAATATGCAAATGCTACTGACGATTGGCCTGACATTGAATTTATGATGACATCATGTTCGACGCCTTCTGACGGAGGTACACAAGTGAAAAAAGCACACGGACTGACGGATGAGTTCTACAATGAAGTTTTTCAAGAAGTTAACAACAAAGACGTTTTCGGTATCTTTCCGATGATGTTACGTCCAAAAAGTAGAGGATTTATTAAACTACGCTCTACGAATCCCCTGGACTACCCTATAATGGTCCACAATTACTTGACGCATCCTGATGACGTTGGAGTACTGAGAGAAGGGGTGAAAGCCGCGGTAGCAGTTGGAGAAACAACAGCCATGAAACGTTTCGGATCAAGGTTTCATGCAAAACCTGTCCCGAATTGTAAGCACCTACCTCCTTATACTGACGAATATTGGGACTGTTTTATTCGTCAATACACCATGACCATTTATCACGTTTCGTGTACAGCGAAAATGGGTCCTTCTAGTGACCCAATGGCAGTTGTGGATCCAAGATTAAAAGTATATGGGATCGAAGGTCTTAGAGTGATAGACGCAAGTATTATGCCCACAATTACTAATGGAAATATTAATGCTCCAGTCATCATGATCGCTGAGAAAGGAGCTGATATGATCAAAGAAGATTGGTTACCAAAATCTAAAAAACGACGACGCAGAAGCATCAGATGCTCTAGATTAGAAAAAATTAGTTTAGGCCGTCATAATAGTAAGCTTTGCTCAATTAAAAGATGA
Protein
MAISTVAATAIKSALAIGGLHTLTFIPIMLAAMAYFNYELLDPEQRPFNQKYLRETYDFIVVGGGSAGSVLANRLSEVEGWNVLLLEAGGHETDISDVPLLSLYLHKSKLDWKYRTQPQDTACQAMIDKRCSWTKGKVLGGSSVLNTMLYIRGNKRDFDQWESFGNPGWGYEDVLPYFKKSEDQRNPYLAKDTKYHQTGGYLTVQDSPYNTPIGAALLQAGEEMGYDFIDVNGAQQTGYAWYQFTIRRGTRCSTAKAFLRPVRLRQNLHIALFSHVTKVLIDKDTKRAYGVEFLRDGTQQVVYAKREVILAAGAIASPQLLMLSGVGPEQHLKEVGIDVIHDSPGVGRNLQDHIAVGGIIFRIDYPVSLVMNRLVNINSALRYAITEDGPLTSSIGLEVVAFINTKYANATDDWPDIEFMMTSCSTPSDGGTQVKKAHGLTDEFYNEVFQEVNNKDVFGIFPMMLRPKSRGFIKLRSTNPLDYPIMVHNYLTHPDDVGVLREGVKAAVAVGETTAMKRFGSRFHAKPVPNCKHLPPYTDEYWDCFIRQYTMTIYHVSCTAKMGPSSDPMAVVDPRLKVYGIEGLRVIDASIMPTITNGNINAPVIMIAEKGADMIKEDWLPKSKKRRRRSIRCSRLEKISLGRHNSKLCSIKR
Summary
Cofactor
FAD
Similarity
Belongs to the GMC oxidoreductase family.
Uniprot
H9JTY9
A0A2H1V018
A0A2A4K2V8
A0A194RF86
A0A437BIK6
A0A0L7L287
+ More
A0A194PXZ8 A0A212FMP0 A0A151I752 E9IKG3 A0A0L7RJW5 K7J279 A0A158NUR0 A0A195BAJ7 A0A232ET33 A0A151K038 A0A1Y9H219 A0A151J2Y9 A0A182UPD3 Q7PS75 A0A182QYJ5 A0A182LAR5 E2BJK3 A0A088AJU9 A0A0L0C3Z5 A0A1I8NG66 A0A453YNK0 B4JKZ7 B4M8F4 B4L265 A0A0M8ZN69 A0A1W4VZR1 B0WJG5 A0A2M4BFU8 B4MTA1 A0A0A1WPB2 B4PW82 A0A182FZP6 B3MXK6 Q8MQN2 B4IJ75 Q9VY06 E2AQ26 B3NUX5 A0A310SVM0 A0A0K8U028 A0A2A3EJH4 A0A0M3QZ56 Q17DW1 A0A3B0J9R4 B4HAL5 Q29IS7 A0A151WNU3 A0A3L8D8C9 A0A2P8ZKR7 J9JSD3 A0A2H8TGH6 D2A3M7 A0A2S2PRC0 E2A0V6 W8CBS3 A0A2J7RBG0 A0A2S2R537 A0A026WCZ0 E0VUA9 A0A1J1IYP0 A0A2P8ZKR2 A0A023F5C8 A0A336LSM3 U4UF26 N6TYT9 A0A182JP93 A0A182VXW5 A0A182UF00 A0A1A9Y1T7 A0A1A9Z4U7 A0A1B0FB96 A0A1A9VSE8 A0A1B0CHA8 A0A084VW87 W5J4T9 A0A182JL50 A0A0C9RQK8 F4WRD0 A0A0R3P3C4 A0A182MI68 A0A182WV72 A0A182I8U7 A0A182GTK4 A0A0J7KTT7 T1I474 A0A2J7RBG4 E0VUA7 E2BJJ7 A0A1B6DYE3 A0A0L7RJU0 J9JXJ8 A0A1I8NVU5 K7J274 A0A2S2PIK7
A0A194PXZ8 A0A212FMP0 A0A151I752 E9IKG3 A0A0L7RJW5 K7J279 A0A158NUR0 A0A195BAJ7 A0A232ET33 A0A151K038 A0A1Y9H219 A0A151J2Y9 A0A182UPD3 Q7PS75 A0A182QYJ5 A0A182LAR5 E2BJK3 A0A088AJU9 A0A0L0C3Z5 A0A1I8NG66 A0A453YNK0 B4JKZ7 B4M8F4 B4L265 A0A0M8ZN69 A0A1W4VZR1 B0WJG5 A0A2M4BFU8 B4MTA1 A0A0A1WPB2 B4PW82 A0A182FZP6 B3MXK6 Q8MQN2 B4IJ75 Q9VY06 E2AQ26 B3NUX5 A0A310SVM0 A0A0K8U028 A0A2A3EJH4 A0A0M3QZ56 Q17DW1 A0A3B0J9R4 B4HAL5 Q29IS7 A0A151WNU3 A0A3L8D8C9 A0A2P8ZKR7 J9JSD3 A0A2H8TGH6 D2A3M7 A0A2S2PRC0 E2A0V6 W8CBS3 A0A2J7RBG0 A0A2S2R537 A0A026WCZ0 E0VUA9 A0A1J1IYP0 A0A2P8ZKR2 A0A023F5C8 A0A336LSM3 U4UF26 N6TYT9 A0A182JP93 A0A182VXW5 A0A182UF00 A0A1A9Y1T7 A0A1A9Z4U7 A0A1B0FB96 A0A1A9VSE8 A0A1B0CHA8 A0A084VW87 W5J4T9 A0A182JL50 A0A0C9RQK8 F4WRD0 A0A0R3P3C4 A0A182MI68 A0A182WV72 A0A182I8U7 A0A182GTK4 A0A0J7KTT7 T1I474 A0A2J7RBG4 E0VUA7 E2BJJ7 A0A1B6DYE3 A0A0L7RJU0 J9JXJ8 A0A1I8NVU5 K7J274 A0A2S2PIK7
Pubmed
19121390
26354079
26227816
22118469
21282665
20075255
+ More
21347285 28648823 12364791 14747013 17210077 20966253 20798317 26108605 25315136 17994087 25830018 17550304 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 17510324 15632085 30249741 29403074 18362917 19820115 24495485 24508170 20566863 25474469 23537049 24438588 20920257 23761445 21719571 23185243 26483478
21347285 28648823 12364791 14747013 17210077 20966253 20798317 26108605 25315136 17994087 25830018 17550304 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 17510324 15632085 30249741 29403074 18362917 19820115 24495485 24508170 20566863 25474469 23537049 24438588 20920257 23761445 21719571 23185243 26483478
EMBL
BABH01003801
ODYU01000059
SOQ34187.1
NWSH01000190
PCG78591.1
KQ460323
+ More
KPJ15965.1 RSAL01000051 RVE50266.1 JTDY01003428 KOB69572.1 KQ459595 KPI96025.1 AGBW02007651 OWR55006.1 KQ978473 KYM93712.1 GL763984 EFZ18946.1 KQ414579 KOC71129.1 AAZX01001588 ADTU01026684 KQ976532 KYM81563.1 NNAY01002337 OXU21519.1 KQ981296 KYN43078.1 KQ980314 KYN16690.1 AAAB01008844 EAA06050.6 AXCN02000040 GL448571 EFN84160.1 JRES01000943 KNC26962.1 CH916370 EDW00250.1 CH940653 EDW62430.1 CH933810 EDW06805.1 KQ435998 KOX67655.1 DS231960 EDS29161.1 GGFJ01002789 MBW51930.1 CH963851 EDW75340.1 GBXI01014029 JAD00263.1 CM000162 EDX01713.1 KRK06314.1 CH902630 EDV38471.1 AY128482 AAM75075.1 CH480847 EDW51054.1 AE014298 BT050468 AAF48400.1 ACJ13175.1 GL441651 EFN64469.1 CH954180 EDV47073.1 KQ759784 OAD62859.1 GDHF01032247 JAI20067.1 KZ288238 PBC31349.1 CP012528 ALC48843.1 CH477290 EAT44640.1 OUUW01000003 SPP78675.1 CH479242 EDW37626.1 CH379063 EAL32576.2 KQ982905 KYQ49513.1 QOIP01000012 RLU16178.1 PYGN01000027 PSN57089.1 ABLF02037931 GFXV01001389 MBW13194.1 KQ971348 EFA05528.1 GGMR01019401 MBY32020.1 GL435626 EFN73009.1 GAMC01000219 JAC06337.1 NEVH01005904 PNF38173.1 GGMS01015954 MBY85157.1 KK107261 EZA53940.1 DS235784 EEB16965.1 CVRI01000064 CRL05333.1 PSN57087.1 GBBI01002388 JAC16324.1 UFQT01000144 SSX20835.1 KB632330 ERL92579.1 APGK01055694 APGK01055695 APGK01055696 APGK01055697 APGK01055698 APGK01055699 APGK01055700 APGK01055701 APGK01055702 APGK01055703 KB741269 ENN71447.1 CCAG010007192 AJWK01012160 AJWK01012161 AJWK01012162 ATLV01017515 KE525172 KFB42231.1 ADMH02002107 ETN58956.1 GBYB01010765 JAG80532.1 GL888284 EGI63347.1 KRT06125.1 KRT06126.1 AXCM01003805 APCN01003121 JXUM01087326 KQ563625 KXJ73585.1 LBMM01003329 KMQ93654.1 ACPB03007739 ACPB03007740 ACPB03007741 PNF38176.1 EEB16963.1 EFN84154.1 GEDC01006623 JAS30675.1 KOC71135.1 ABLF02037922 AAZX01003729 GGMR01016673 MBY29292.1
KPJ15965.1 RSAL01000051 RVE50266.1 JTDY01003428 KOB69572.1 KQ459595 KPI96025.1 AGBW02007651 OWR55006.1 KQ978473 KYM93712.1 GL763984 EFZ18946.1 KQ414579 KOC71129.1 AAZX01001588 ADTU01026684 KQ976532 KYM81563.1 NNAY01002337 OXU21519.1 KQ981296 KYN43078.1 KQ980314 KYN16690.1 AAAB01008844 EAA06050.6 AXCN02000040 GL448571 EFN84160.1 JRES01000943 KNC26962.1 CH916370 EDW00250.1 CH940653 EDW62430.1 CH933810 EDW06805.1 KQ435998 KOX67655.1 DS231960 EDS29161.1 GGFJ01002789 MBW51930.1 CH963851 EDW75340.1 GBXI01014029 JAD00263.1 CM000162 EDX01713.1 KRK06314.1 CH902630 EDV38471.1 AY128482 AAM75075.1 CH480847 EDW51054.1 AE014298 BT050468 AAF48400.1 ACJ13175.1 GL441651 EFN64469.1 CH954180 EDV47073.1 KQ759784 OAD62859.1 GDHF01032247 JAI20067.1 KZ288238 PBC31349.1 CP012528 ALC48843.1 CH477290 EAT44640.1 OUUW01000003 SPP78675.1 CH479242 EDW37626.1 CH379063 EAL32576.2 KQ982905 KYQ49513.1 QOIP01000012 RLU16178.1 PYGN01000027 PSN57089.1 ABLF02037931 GFXV01001389 MBW13194.1 KQ971348 EFA05528.1 GGMR01019401 MBY32020.1 GL435626 EFN73009.1 GAMC01000219 JAC06337.1 NEVH01005904 PNF38173.1 GGMS01015954 MBY85157.1 KK107261 EZA53940.1 DS235784 EEB16965.1 CVRI01000064 CRL05333.1 PSN57087.1 GBBI01002388 JAC16324.1 UFQT01000144 SSX20835.1 KB632330 ERL92579.1 APGK01055694 APGK01055695 APGK01055696 APGK01055697 APGK01055698 APGK01055699 APGK01055700 APGK01055701 APGK01055702 APGK01055703 KB741269 ENN71447.1 CCAG010007192 AJWK01012160 AJWK01012161 AJWK01012162 ATLV01017515 KE525172 KFB42231.1 ADMH02002107 ETN58956.1 GBYB01010765 JAG80532.1 GL888284 EGI63347.1 KRT06125.1 KRT06126.1 AXCM01003805 APCN01003121 JXUM01087326 KQ563625 KXJ73585.1 LBMM01003329 KMQ93654.1 ACPB03007739 ACPB03007740 ACPB03007741 PNF38176.1 EEB16963.1 EFN84154.1 GEDC01006623 JAS30675.1 KOC71135.1 ABLF02037922 AAZX01003729 GGMR01016673 MBY29292.1
Proteomes
UP000005204
UP000218220
UP000053240
UP000283053
UP000037510
UP000053268
+ More
UP000007151 UP000078542 UP000053825 UP000002358 UP000005205 UP000078540 UP000215335 UP000078541 UP000075884 UP000078492 UP000075903 UP000007062 UP000075886 UP000075882 UP000008237 UP000005203 UP000037069 UP000095301 UP000075900 UP000001070 UP000008792 UP000009192 UP000053105 UP000192221 UP000002320 UP000007798 UP000002282 UP000069272 UP000007801 UP000001292 UP000000803 UP000000311 UP000008711 UP000242457 UP000092553 UP000008820 UP000268350 UP000008744 UP000001819 UP000075809 UP000279307 UP000245037 UP000007819 UP000007266 UP000235965 UP000053097 UP000009046 UP000183832 UP000030742 UP000019118 UP000075881 UP000075920 UP000075902 UP000092443 UP000092445 UP000092444 UP000078200 UP000092461 UP000030765 UP000000673 UP000075880 UP000007755 UP000075883 UP000076407 UP000075840 UP000069940 UP000249989 UP000036403 UP000015103 UP000095300
UP000007151 UP000078542 UP000053825 UP000002358 UP000005205 UP000078540 UP000215335 UP000078541 UP000075884 UP000078492 UP000075903 UP000007062 UP000075886 UP000075882 UP000008237 UP000005203 UP000037069 UP000095301 UP000075900 UP000001070 UP000008792 UP000009192 UP000053105 UP000192221 UP000002320 UP000007798 UP000002282 UP000069272 UP000007801 UP000001292 UP000000803 UP000000311 UP000008711 UP000242457 UP000092553 UP000008820 UP000268350 UP000008744 UP000001819 UP000075809 UP000279307 UP000245037 UP000007819 UP000007266 UP000235965 UP000053097 UP000009046 UP000183832 UP000030742 UP000019118 UP000075881 UP000075920 UP000075902 UP000092443 UP000092445 UP000092444 UP000078200 UP000092461 UP000030765 UP000000673 UP000075880 UP000007755 UP000075883 UP000076407 UP000075840 UP000069940 UP000249989 UP000036403 UP000015103 UP000095300
Interpro
SUPFAM
SSF51905
SSF51905
Gene 3D
ProteinModelPortal
H9JTY9
A0A2H1V018
A0A2A4K2V8
A0A194RF86
A0A437BIK6
A0A0L7L287
+ More
A0A194PXZ8 A0A212FMP0 A0A151I752 E9IKG3 A0A0L7RJW5 K7J279 A0A158NUR0 A0A195BAJ7 A0A232ET33 A0A151K038 A0A1Y9H219 A0A151J2Y9 A0A182UPD3 Q7PS75 A0A182QYJ5 A0A182LAR5 E2BJK3 A0A088AJU9 A0A0L0C3Z5 A0A1I8NG66 A0A453YNK0 B4JKZ7 B4M8F4 B4L265 A0A0M8ZN69 A0A1W4VZR1 B0WJG5 A0A2M4BFU8 B4MTA1 A0A0A1WPB2 B4PW82 A0A182FZP6 B3MXK6 Q8MQN2 B4IJ75 Q9VY06 E2AQ26 B3NUX5 A0A310SVM0 A0A0K8U028 A0A2A3EJH4 A0A0M3QZ56 Q17DW1 A0A3B0J9R4 B4HAL5 Q29IS7 A0A151WNU3 A0A3L8D8C9 A0A2P8ZKR7 J9JSD3 A0A2H8TGH6 D2A3M7 A0A2S2PRC0 E2A0V6 W8CBS3 A0A2J7RBG0 A0A2S2R537 A0A026WCZ0 E0VUA9 A0A1J1IYP0 A0A2P8ZKR2 A0A023F5C8 A0A336LSM3 U4UF26 N6TYT9 A0A182JP93 A0A182VXW5 A0A182UF00 A0A1A9Y1T7 A0A1A9Z4U7 A0A1B0FB96 A0A1A9VSE8 A0A1B0CHA8 A0A084VW87 W5J4T9 A0A182JL50 A0A0C9RQK8 F4WRD0 A0A0R3P3C4 A0A182MI68 A0A182WV72 A0A182I8U7 A0A182GTK4 A0A0J7KTT7 T1I474 A0A2J7RBG4 E0VUA7 E2BJJ7 A0A1B6DYE3 A0A0L7RJU0 J9JXJ8 A0A1I8NVU5 K7J274 A0A2S2PIK7
A0A194PXZ8 A0A212FMP0 A0A151I752 E9IKG3 A0A0L7RJW5 K7J279 A0A158NUR0 A0A195BAJ7 A0A232ET33 A0A151K038 A0A1Y9H219 A0A151J2Y9 A0A182UPD3 Q7PS75 A0A182QYJ5 A0A182LAR5 E2BJK3 A0A088AJU9 A0A0L0C3Z5 A0A1I8NG66 A0A453YNK0 B4JKZ7 B4M8F4 B4L265 A0A0M8ZN69 A0A1W4VZR1 B0WJG5 A0A2M4BFU8 B4MTA1 A0A0A1WPB2 B4PW82 A0A182FZP6 B3MXK6 Q8MQN2 B4IJ75 Q9VY06 E2AQ26 B3NUX5 A0A310SVM0 A0A0K8U028 A0A2A3EJH4 A0A0M3QZ56 Q17DW1 A0A3B0J9R4 B4HAL5 Q29IS7 A0A151WNU3 A0A3L8D8C9 A0A2P8ZKR7 J9JSD3 A0A2H8TGH6 D2A3M7 A0A2S2PRC0 E2A0V6 W8CBS3 A0A2J7RBG0 A0A2S2R537 A0A026WCZ0 E0VUA9 A0A1J1IYP0 A0A2P8ZKR2 A0A023F5C8 A0A336LSM3 U4UF26 N6TYT9 A0A182JP93 A0A182VXW5 A0A182UF00 A0A1A9Y1T7 A0A1A9Z4U7 A0A1B0FB96 A0A1A9VSE8 A0A1B0CHA8 A0A084VW87 W5J4T9 A0A182JL50 A0A0C9RQK8 F4WRD0 A0A0R3P3C4 A0A182MI68 A0A182WV72 A0A182I8U7 A0A182GTK4 A0A0J7KTT7 T1I474 A0A2J7RBG4 E0VUA7 E2BJJ7 A0A1B6DYE3 A0A0L7RJU0 J9JXJ8 A0A1I8NVU5 K7J274 A0A2S2PIK7
PDB
5NCC
E-value=4.99695e-64,
Score=622
Ontologies
GO
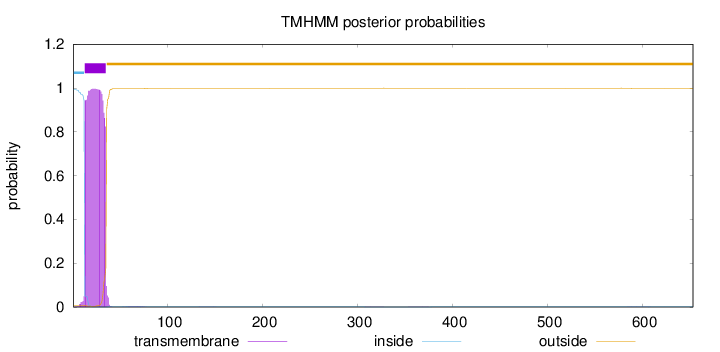
Topology
Length:
653
Number of predicted TMHs:
1
Exp number of AAs in TMHs:
22.64235
Exp number, first 60 AAs:
22.55924
Total prob of N-in:
0.99325
POSSIBLE N-term signal
sequence
inside
1 - 12
TMhelix
13 - 35
outside
36 - 653
Population Genetic Test Statistics
Pi
300.195572
Theta
206.969297
Tajima's D
1.322904
CLR
0.161108
CSRT
0.743612819359032
Interpretation
Uncertain
