Gene
KWMTBOMO09557
Pre Gene Modal
BGIBMGA013019
Annotation
PREDICTED:_uncharacterized_protein_LOC105842057_[Bombyx_mori]
Transcription factor
Location in the cell
Nuclear Reliability : 3.727
Sequence
CDS
ATGCCGAAGGTTACGGATGGAAAAAAATATACAAAAGCATATACAGAAGAAGATATAGAAAAAGCTTTGCAAGCCATAAAAAATGGAATGGGTAAGAAAACTGCTGCAAATAAATTTAATATCCCAAGAGCAACTTTGCAGTTCCGCCTATCCAACCAATTTGTCAAAAGTCGACCTGGTCCAGATACAGTTTTATCTAGTGCAGAAGAACAAGAGATAGTTAAATGGATAATTGCAGCTTGTAGAAAAGGATTTCCACGGCGTAAAGAGGATGTCATAAAGAGCGTCAAACTGTATTTGGATGAACATCCCAGACCAAACAGTTTTATAGATAATACACCAGGCGAGGGATGGTATAAATTATTTTTGAAGAGGCATCCTGAGCTTTCTATACGAACATCAGAGGCTGTAACTTCAGCAAGCTCAAAGTTGTCAGAGAAGCAAATTAGAAACTGGTTTACACAAGTAGAAGAATATCTGAAAGAAAATGATCTCTTTCATATTTTGAGTGATCCATCTAGAGTTTACAATGGGGATGAAACGAATTTTATTTTATGCCCAAAGAACACCAAAGTTATCGCTTTAAAGGGAACCAAACATGTGTATGAGGTAGATCACGCTCAGGCTAAATCATGTGTTACTGTAATGTTCACATTTTCGGCGGATGGTGTAACAACACCACCTATGATTATTTACCCTTTAAAGAGAATGCGACCTGAGATTCAAAAGTCTGTGCCTAGCGAGTGGGGTATTGGCTTGTCAGATAATGGATGGATGAAGGCGGAGCTTTTCATCCAGTACATACAGAATGTTCTGCATCCAGCGCTAATGAAGATGAATATCACTTTTCCAGTAATTCTTTTTTTAGATGGACATAAAAGCCACACAACACTTCAGCTTACTCAACTTTGTCAAAATCTTGGTATAATACTTATAGCTCTATACCCTAACGCGACACGGATTTTGCAACCAGCAGACGTAGCGGCATTTCGTCCAATAAAAGTAATGTGGCGTAAAGCAGTCCTAGAATGGCGAACAGATAACTTAACAAAATCTTTACTAAAAACAGATGTTGCTCCTATACTGGAGAAGCTTTTACCCAAATTAAATAAAACGACTTTGAAGAACGGCTTTGAAGCTTGTGGTTTATGCCCATTTAATTCAGATCAAGTGGACTACACAAAATGTTTAATGCCCTATAAACCAGTCCAAGAAATTTTGGATGACAACTGTTCTACAAGTCTTAACTATCAGAAATTCGAAGAAATAGTTGGACCACAGTTAATCAGAAAGATGAAAAATCAGGTGGTAGGTAAATCAAGAGAAGCTAAAAAGCTTCACGATTTGTATAAATATCTCGTACCAAAAATGTCAACACATGAGTCATCGTCACAAATCATAGAAACAACTTCTAAGGAACTAATGCAATACAAGGACAGGAGCAATTTAAATGAACAAAACAACGATGGTTCAGAAGATTTCACCACAATGGCAGCCCTCAATGAATTAACGACAATGCCAACAGACGTAGAACTTTTGCTAAATGAACCAGAACTGTCAACGCATGAACCTCAATCCAAAGACACTGGCACTTCTAAAACAAACACGATGATTGATGATTATTCAGGAACCTCAGGACACCTAAAACCTCTGAACGTTGAACTTCAGCATCAGGGGGTTGAGGAATTATCTTTGGCATCCTGTATAAATCTTACATCGACAAACGATCCAATACCAGAAGCTGTACAAATGGTTGAGGTTATTTCTGAGTGTGAAAAAGAAAACCAAACACCAATATACGTAATACCTATTCCATCCACTAGTCGTCAAAGTTTAGAGCTGATTGGAGAAAAATGGGTACTAGTGGATGACATGCCAACCGAAGTCCCTTCAAAGAGTCCAATAAAGAAACAAAATATAAAAGATTTTTTAAAGTTACCAAAGACTCCAATTAAAACACAAAAAGCTGGTATAGAACGTCGCTCTTTTGTTTTAACGAGTAAAGATTGGGAAGCTGTAGAACTAGAAAAACAACAAAAAAAGATGAAAGAGATACAAGAAAAGGAGAACAAACGAATAGCAAGACTAGAAAAAGCCAAAGAGAAACGCTTACTCAAGGAAACAAAACAAGCTGAAACTATAAATAAAAAGAACAAAGCTAAAAATTTAAATTTAATTAAGAAAAGTAAGCCAAATTCTGATAGTATCAAATCAAAAATTGTTATTCAATCAAATGTTTTAATTAAGCCTGCAAATAGAATCTCAAGTGATATTAAAGAAAAAAGAGAATATACAAAAGAAATTAAAAAAGGACCTATGAACGAAGTTAAACCTGCAGATATTACTGCCCTGAAGGAAATAAAAAACAGGCCAATTTTAGATGCAGTTGAAGCTGCAAATGAAAGACTCTTTAAAGAATCGGAACAAGTACAGATGGTAAAAAAAGAATTAAAGAAAGGACCAACTTTAGATGAACTTAAACCTCCGAACATAACTTTCCGAGATTCAAGGGAAACCGGAAAAGATAAGAAGATAAAAATTAAAGTAAATAGATTGAGACCAATGACAGATGAAGAATTGTTAAAAATGTTGGAAGATGATTAG
Protein
MPKVTDGKKYTKAYTEEDIEKALQAIKNGMGKKTAANKFNIPRATLQFRLSNQFVKSRPGPDTVLSSAEEQEIVKWIIAACRKGFPRRKEDVIKSVKLYLDEHPRPNSFIDNTPGEGWYKLFLKRHPELSIRTSEAVTSASSKLSEKQIRNWFTQVEEYLKENDLFHILSDPSRVYNGDETNFILCPKNTKVIALKGTKHVYEVDHAQAKSCVTVMFTFSADGVTTPPMIIYPLKRMRPEIQKSVPSEWGIGLSDNGWMKAELFIQYIQNVLHPALMKMNITFPVILFLDGHKSHTTLQLTQLCQNLGIILIALYPNATRILQPADVAAFRPIKVMWRKAVLEWRTDNLTKSLLKTDVAPILEKLLPKLNKTTLKNGFEACGLCPFNSDQVDYTKCLMPYKPVQEILDDNCSTSLNYQKFEEIVGPQLIRKMKNQVVGKSREAKKLHDLYKYLVPKMSTHESSSQIIETTSKELMQYKDRSNLNEQNNDGSEDFTTMAALNELTTMPTDVELLLNEPELSTHEPQSKDTGTSKTNTMIDDYSGTSGHLKPLNVELQHQGVEELSLASCINLTSTNDPIPEAVQMVEVISECEKENQTPIYVIPIPSTSRQSLELIGEKWVLVDDMPTEVPSKSPIKKQNIKDFLKLPKTPIKTQKAGIERRSFVLTSKDWEAVELEKQQKKMKEIQEKENKRIARLEKAKEKRLLKETKQAETINKKNKAKNLNLIKKSKPNSDSIKSKIVIQSNVLIKPANRISSDIKEKREYTKEIKKGPMNEVKPADITALKEIKNRPILDAVEAANERLFKESEQVQMVKKELKKGPTLDELKPPNITFRDSRETGKDKKIKIKVNRLRPMTDEELLKMLEDD
Summary
Uniprot
Pubmed
EMBL
Proteomes
PRIDE
Interpro
SUPFAM
SSF46689
SSF46689
ProteinModelPortal
Ontologies
GO
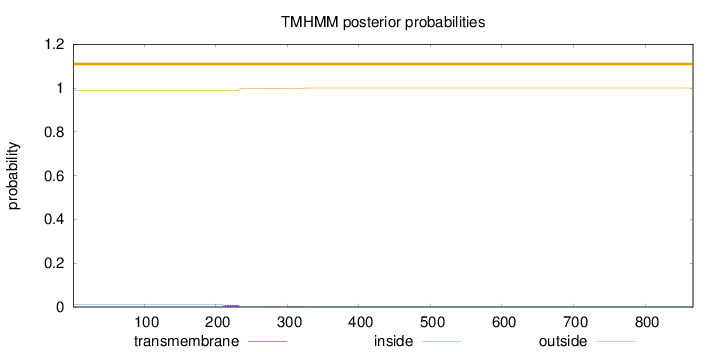
Topology
Length:
867
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.24782
Exp number, first 60 AAs:
0
Total prob of N-in:
0.01071
outside
1 - 867
Population Genetic Test Statistics
Pi
261.164593
Theta
201.482251
Tajima's D
1.090883
CLR
100.991823
CSRT
0.684665766711664
Interpretation
Uncertain
