Gene
KWMTBOMO09350
Annotation
PREDICTED:_RNA-directed_DNA_polymerase_from_mobile_element_jockey-like?_partial_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 2.022 Nuclear Reliability : 1.558
Sequence
CDS
ATGTATGGCAGCGAAACATGGGCGGCAACCAAAAGGAACGTACATACCATTCAAGTTGCTGAAATGAAAATGCTGAGGTGGATGTGCGGCATGACTAGGCTCGATAAAATCCGCAACGAGTACGTGCGTGGAAGTCTAGGTGTACGGGACATCGCCGACAAGATGCAGGAGAGTAGGCTGAGATGGTATGGTCACGTAAAAAGGAAGCCACCTGACTACGTCGGAAATTTGGCGCTACTGCTCGACCTCCCCGGTCGGAGACCTAGAGGCAGACCCAAGACAAGGTGGAGGGACGTAGTGCTGAAGGACCAGAGAGAGAATGCAATGTTGCTGACGAGGATGTCGAAGACAGAGCGAAGTGGTGCAGTAAAGTCAGGAAAGCTGACCCCACCACCATGTGGGATAAATAGCTGGGAAGAGAGAGAGAGAGAGAGCGTACAGACGACTATTTCATGTACCTAA
Protein
MYGSETWAATKRNVHTIQVAEMKMLRWMCGMTRLDKIRNEYVRGSLGVRDIADKMQESRLRWYGHVKRKPPDYVGNLALLLDLPGRRPRGRPKTRWRDVVLKDQRENAMLLTRMSKTERSGAVKSGKLTPPPCGINSWEERERESVQTTISCT
Summary
Uniprot
EMBL
Pfam
PF00078 RVT_1
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
Ontologies
KEGG
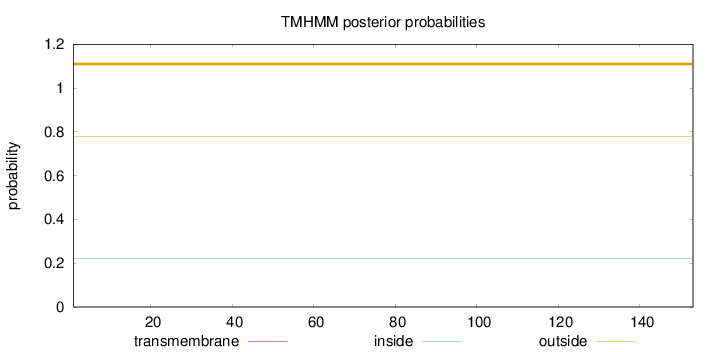
Topology
Length:
153
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00188
Exp number, first 60 AAs:
0.00162
Total prob of N-in:
0.22165
outside
1 - 153
Population Genetic Test Statistics
Pi
217.788257
Theta
138.600448
Tajima's D
1.868835
CLR
8.234085
CSRT
0.865706714664267
Interpretation
Uncertain
