Gene
KWMTBOMO09264
Annotation
PREDICTED:_LOW_QUALITY_PROTEIN:_uncharacterized_protein_LOC105841899?_partial_[Bombyx_mori]
Location in the cell
PlasmaMembrane Reliability : 1.041
Sequence
CDS
ATGGAATTGGATAGGACAACAGCTATCGGGGACCGCAGCAACCTGTGCTTCCGCTGCGGCCAACCGGGGCACAAGGCAGCCTCCTGCACCGCAGCCGCGCCGCACTGCGTCTTGTGCGACGCGGCTAAAAGGAAGGCTGACCACCGGGCCGGGGGCCCGGCGTGCAAGTCCGCCCCCTCCTCTACAAAAACGAGGAGGGGCGGAAAGAAAAAGAAGCCGGCTGCAACAAGGGCAGCCGGCGAGTCAGTTCCCGCCGCGGCTGTTGTCGAGCCGCAGGGCGGAACAGACGAGGGAGCGATGGATGATCTCCTCGTCCACACCATGGCGGAGTGGTCAATCGACGTCGCTGTCGCTGCGGAGCCGTACTTCGTCCCCACCGACCAGGACTGCTGGATGGGGGACGTCGACGGCTCCGTGGCCATTGTGTTGAGACAGTCCGCGGCGTTACCCCCCTTGGAAATGGTGGCCAGGGGACCCGGGTACGTCGCGGCTAGGATCGACGGGACCGTCGTGATCGGGGCGTACTTTTCTCCGAACAGGAGCCTCGCCGAGTTCGAGCGGTTTCTGGGCGGGCTCGAGGCGCTTTTACACCGCCTCGGGTCCCGCCCTGTCATCGTCGCGGGGGACTTCAATGCCAAGTGCACGGCTTGGGGTTCCCCCCGCACGGATGCGCGAGGCGAGGCCCTTTCGGAGTGGGCGATCGCGACCGGCCTCTGTCTCCTAAACAGAGGCACTGTCGCGACTTGCGTGCGGTGGAACGGGGAATCTCACGTGGACGTGACGTTCGCGTCCCCTTCCGCCGTGCGCCGCACCCGCGGCTGGCGCGTACTCGAGGGGGCGGAGACGCTCTCGGACCACAGATACGTCCGATTCGAGCTCTCCGCCTCCTCCTTGTCGTCGTCGTCAGCCGCGGGCGCCCGCGGCGAGGAGGAGCTCCCGCAAGGTGCGCCCCGTTCGTTCCCCAGGTGGGCCTTGAAACGGATGAACAAGGAGTTGCTGATGGAGGCGGCCGCTGTTGCTGCGTGGGCGCCAAAACCCGCGCGGCTCGCGGACGTGGATGCGGAGGTCGACTGGTTCCGGGGCACCATGGCAAACATTTGCGATGCCGCCATGCCCCGGGTCGGCCCCCGAGCACCACGTGGGGGTGCGTTCTGGTGGTCGCCCGAGATCGCAAGACTCCGAGAGGAGTGCGTGAGGGCGCGCCGCCGGAGCGCACGCCACCGCCGCCGCCGTCGTCGCGACGCTGCGTTCGCGGAGACGGCGGCCCAGCTGCACGCTGACTGTCGTCAAAAGCAGACGGCGCTGCAGCTGGCCATCAGCGAGGCCAAGAAGCAGAGCATGAAGACTCTCCTGGAGTCGCTCGACGAAGATCCCTGGGGCGCCCATACAAGATGGGGACCGGCTCGGGGGCTTTTCGAAGCGTGCCTCGAGTCGGGACGGTTCCCGTCGAAATGGAAGACGGGCAGACTGGTCCTTCTGCGGAAGGACGGGCGACCCGCGGACTCACCCGCGGGCTATCGCCCCATCGTGTTGCTGGACGAAGCGGGCAAGATGCTCGAGAGGATCGTCGCGGCCCGCATCGTCCGGCACCTGACCGAGACGGGTCCCGATCTTTCGGCGGAGCAATACGGCTTCAGAGAGGGCCGCTCGACTATCGATGCAATTCTGCGCGTGCGGGCCCTCTCCGATGAAGCCGTATCCCGGGGCGGGGTGGCGATGGCGTTATCCTTAGATGTAAGGAACGCCTTCGACACCCTGCCCTGGTCGGTGATCGCGGGGGCGCTGGAGTATCACGGCGTCCCCGCATACCTCCGTCGACTGATCGGCTCCTACCTCGAGGGCAGGTCGATCCAGTGCATCGGACACGGTGGGGCGATGTACCGCTTCCCCGTCGAGCGCGGTGTTCCGCAGGGGTCCGTCCTGGGCCCCCTGCTGTGGAACATCGGCTACGACTGGGTCCTGCGGGGCGTCATTCGGGGACCCCTCCCCGGGCTCAGCGTAATTTGTTACGCTGACGACACGTTGGTCGTAGCTCCGGGGAGGGACTACCGGGAGTCTGCCCGTCTGGCGTGCGCAGGCGTGGCACACGTCGTCACCAGGATCCGACGGTTGGGGCTCGAGGTGGCGCTCGACAAAACCCAGGCCCTGCTCTTTCACGGGCCGGGACGAGCGCCGCCTGTGGGTGCCCACCTCGTGATCGGAGGCGTCCGCGTCGGGGTCGGGGTGACCGGTCTTCGGTACCTCGGCCTCGAACTGGACGGTCGGTGGAACTTCCGCGCTCACTTTGAGAAGTTGGGCCCTCGGCTGATGGCAACGGCCGGCTCTTTGAGCCGGCTCCTTCCAAACGTCGGGGGTCCCGATCAGGTGGCGCGCCGCCTCTACATGGGGGTGGTGCGGTCGATGGCACTATACGGCGCGCCCGTATGGTGCCACGCCCTGACCCGCCAGAACGTCGCCGCGCTGCGACGTCCGCAGCGCGCGATCGCGGTTAGGGCGATCCGAGGATACCGCACCGTCTCCTTCGAGGCGGCGTGCTTGCTCGCCGGGGCGCCACCCTGGGACCTGGAGGCGGAGGCGCTCGCTGCCGATTACCGGTGGCGTAGCGACCTTCGCTCTAGGGGGGAAGGGCGCCCCGGCGAAGGAGTAGTTCGAGCGCGGAGGCTCCACTCTCGGCGGTCCGTGCTGGAAGCGTGGTCTCGCCGCTTGGCGGACCCGTCGGCCGGCCTCCGTACGGCCTCCGTACGGCGCGTTGCTCTGGTTTCGACGAGCAACGCGCCGCCCTCGTCGCGGTCATAG
Protein
MELDRTTAIGDRSNLCFRCGQPGHKAASCTAAAPHCVLCDAAKRKADHRAGGPACKSAPSSTKTRRGGKKKKPAATRAAGESVPAAAVVEPQGGTDEGAMDDLLVHTMAEWSIDVAVAAEPYFVPTDQDCWMGDVDGSVAIVLRQSAALPPLEMVARGPGYVAARIDGTVVIGAYFSPNRSLAEFERFLGGLEALLHRLGSRPVIVAGDFNAKCTAWGSPRTDARGEALSEWAIATGLCLLNRGTVATCVRWNGESHVDVTFASPSAVRRTRGWRVLEGAETLSDHRYVRFELSASSLSSSSAAGARGEEELPQGAPRSFPRWALKRMNKELLMEAAAVAAWAPKPARLADVDAEVDWFRGTMANICDAAMPRVGPRAPRGGAFWWSPEIARLREECVRARRRSARHRRRRRRDAAFAETAAQLHADCRQKQTALQLAISEAKKQSMKTLLESLDEDPWGAHTRWGPARGLFEACLESGRFPSKWKTGRLVLLRKDGRPADSPAGYRPIVLLDEAGKMLERIVAARIVRHLTETGPDLSAEQYGFREGRSTIDAILRVRALSDEAVSRGGVAMALSLDVRNAFDTLPWSVIAGALEYHGVPAYLRRLIGSYLEGRSIQCIGHGGAMYRFPVERGVPQGSVLGPLLWNIGYDWVLRGVIRGPLPGLSVICYADDTLVVAPGRDYRESARLACAGVAHVVTRIRRLGLEVALDKTQALLFHGPGRAPPVGAHLVIGGVRVGVGVTGLRYLGLELDGRWNFRAHFEKLGPRLMATAGSLSRLLPNVGGPDQVARRLYMGVVRSMALYGAPVWCHALTRQNVAALRRPQRAIAVRAIRGYRTVSFEAACLLAGAPPWDLEAEALAADYRWRSDLRSRGEGRPGEGVVRARRLHSRRSVLEAWSRRLADPSAGLRTASVRRVALVSTSNAPPSSRS
Summary
Uniprot
EMBL
Proteomes
Interpro
Gene 3D
ProteinModelPortal
PDB
1WDU
E-value=6.67678e-10,
Score=157
Ontologies
GO
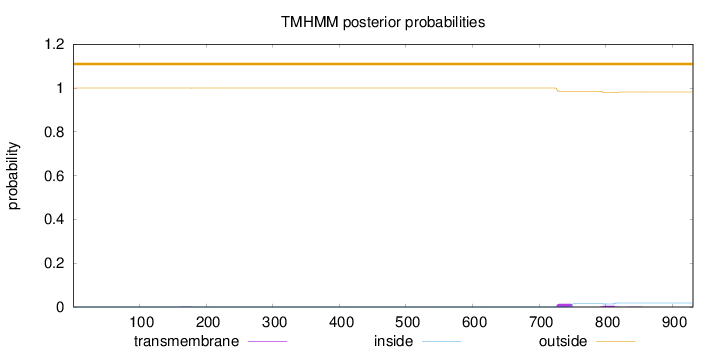
Topology
Length:
931
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.46028
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00033
outside
1 - 931
Population Genetic Test Statistics
Pi
22.990987
Theta
18.141642
Tajima's D
0
CLR
110.529918
CSRT
0.373631318434078
Interpretation
Uncertain
