Gene
KWMTBOMO08822
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 1.308 Mitochondrial Reliability : 1.731 Nuclear Reliability : 1.609
Sequence
CDS
ATGATTGCACACCGCAGAACCAGGCCCCTAGTCCCCGGCGGCCCCGCTAGTCCTGGTTACGGTAGCGGCCATGGTAACGGCGGGGCAAGGGGTGCTAAGAATCCCCGGCAGAGATCCGACTACCACCCCCCTTCACAACGACAACTGGCCCTGGTGACATACAACGTGCGCACTTTGAGGTTGGACGAGAAGCTAATAGAGCTGGAAGAAGAAATGAACAAGTTGCGCTGGGACGTTATTGGTTTATCGGAGATCCGAAGAGAGGGGGAGGATACGAAAACCTTACAGTCCGGCCACTTGTTCTACTACCGGGAGGGAGAACAGAAGTCCCAAGGTGGTGTTGGGTTTATCGTCCACAAATCTCTCATAAAGAACATTATGGCCATCGAAAGTGTGTCGAGTATACCTTGTCCTCAAAATCTCAAAGCGGTATTCGCTGAAGATTGTACAGGCGATGATGAGTTGAAAGTCGGGGCCATTTGGATTTGGGCAGCGGAATCCCAGAGGACAGATGCTGGCGGACTTTATGGAGAAGGAAGGACTCTTTATGATGAATTCTTTCTTCAAGAAGCCGCCATAACGGAAATGGACCTGCGTATCAAAGAGACGATTGAGCGAAACCAAGGCTCTAAAGTGTTCGCAAGAGACGCTTCTATTGGGCAAAGCCAGCTAACGAAGCTGAAGACCGAAGATGGACGTGTAACGTCGAATAGGTCAGAGGTCCTGCGGGAGATAGAGACGTTCTACGGGCAGCTGTTTACCTCGGTCTCAGCTGAACCGCAAGGTTTAGCTGCAGACCCTAGAGCCAAGCTAACCCGACACTACACCGAAGACATCCCGGACATCAGCCTGTACGAGATTAGGATGGCTCTCAAACAGCTTAAGAATAACGAGGCACCGGGCGAGGATGGAATTACTACGGAGCTTCTGAAAGCGGGCGGGACGCCGGTACTGAAAGTCCTTCAGAAGCTCTTCAATTCCTCTTTGCTCGATGGAAAACCGCCACAGGCATGGAATAGGGGTGTTGTGATCCTGTTCTTCAAAAAAGGTGACAAAACCTTATTGAAGAACTACAGGCCCATAGCTCTGCTGAGCCATGTCTACAAGCTGTTCTCGAGAGTCATCACGAAGCGTCTCGAACGCAGGTTTGATGACTTCCAGCCTACCGAACAAGCCGGTTTCCGAAAAGGCTACAGTACCATAGACCACATACATACGCTGCGGCAGATTGTACAGAAGACCGAAGAGTACAATCGGCCATTATGCTTAGCGTTTGTGGACTATGAGAAAGCCTTTCATTCCGTCGAAACATGGGCGGTCCTTAGATCTCTCCAGCGGTGCCGTATTGACCACCGGCCGCAACTACTTCGCAGCCAGGTGTAA
Protein
MIAHRRTRPLVPGGPASPGYGSGHGNGGARGAKNPRQRSDYHPPSQRQLALVTYNVRTLRLDEKLIELEEEMNKLRWDVIGLSEIRREGEDTKTLQSGHLFYYREGEQKSQGGVGFIVHKSLIKNIMAIESVSSIPCPQNLKAVFAEDCTGDDELKVGAIWIWAAESQRTDAGGLYGEGRTLYDEFFLQEAAITEMDLRIKETIERNQGSKVFARDASIGQSQLTKLKTEDGRVTSNRSEVLREIETFYGQLFTSVSAEPQGLAADPRAKLTRHYTEDIPDISLYEIRMALKQLKNNEAPGEDGITTELLKAGGTPVLKVLQKLFNSSLLDGKPPQAWNRGVVILFFKKGDKTLLKNYRPIALLSHVYKLFSRVITKRLERRFDDFQPTEQAGFRKGYSTIDHIHTLRQIVQKTEEYNRPLCLAFVDYEKAFHSVETWAVLRSLQRCRIDHRPQLLRSQV
Summary
Uniprot
EMBL
Proteomes
Pfam
PF00078 RVT_1
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
Ontologies
GO
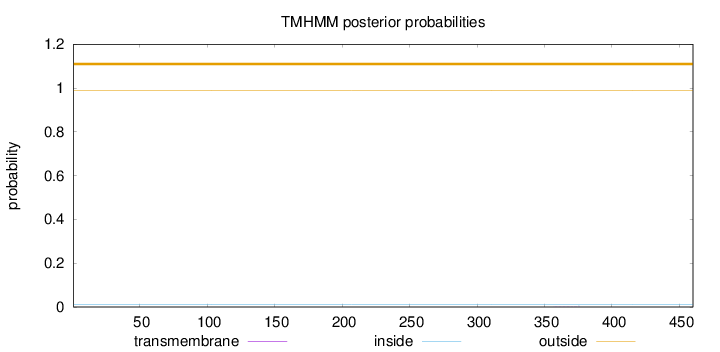
Topology
Length:
460
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00798
Exp number, first 60 AAs:
0.00034
Total prob of N-in:
0.00988
outside
1 - 460
Population Genetic Test Statistics
Pi
272.870512
Theta
209.410799
Tajima's D
0.929065
CLR
0
CSRT
0.63686815659217
Interpretation
Uncertain
