Gene
KWMTBOMO08486
Annotation
PREDICTED:_piggyBac_transposable_element-derived_protein_4-like_isoform_X4_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 1.576 PlasmaMembrane Reliability : 1.06
Sequence
CDS
ATGGAAAGATTTAACCTTATTGCAATTGAGGATTTGCCCGCTGGGGAACCGATAGAGCAGGCATTGACGGAGCGAAGTACGTTGAATGTGGCGGCAGTGGTACGAGCACGCCATCTTCTAGATATAATATACAGATTTCTGTACATGGCAGCCGTAGTCCTTCGCCGTCACATAAACGTGTGCGTCGATCGCTTTGCCCAGACCAAGAACATGATACTACGAAGTGACGACGATGACAGCGATGATGAGAGCCAGAATGATAACGACGAAATCGTTCCCATGATTATCCCCCGGGCAGCTGGAATACTGATGGACACTCCGGACGACGAGGAGATTGAGATTGACACGGCGGCGGCAACTAAACCTCTTGTGACATCTTCCGCGCCTTCCATCATTTCAGCACCTACATCACCAGCGCGTGATACCGGTATGAAGCCTAGTCATTTTTTTGAATTTGACTGGGGAACTTTCCCTGACTCTCCTATACCGCCAACAGAAAGGCGTGAATCTTTTAAGGAACACTCTGGTCCCACTGTTGCTGTAAGTGACCCTTACGAGATTTTTAGACTTATTTGGGACCAGGAGTTCATGGAATTTATTGTGCAGGAAACAAATAGATATGCCCAACAATTAGCTGCTGAAATGTTGGACGGCGGGGAATTACAGGCCTCCAGTAGGATTACAGAATGGAAAGAAACCAATGTAGACGAGTTATTGGTGTTCTTCGGAATATTACTGGCGATGGGAATCGTCATAAAGAACCGGGTTGAAGAGTACTGGAATACGGAACAAAACATCTTCTCCACTCCAGGTTTCAAAGTATATATGTCGCTTAGGAGGTTCCAGTTACTCAGCAGGTGCTTACATTTTAATAATTCCGAGAACTTGAGAAACCTCAACCTGGATCCCTCACAGGCAAAGCTTTTTAAAGTGGAGCCTGTGATTAGCCATTTGAATTCCAAGTTCACGGAATTATATATAATGAAGCAGAACATTGCTTTAGATGAATCGCTGCTGCAGTGGAAAGGTTGGTTGAACATCAACCAATTTATTCCAAACAAAGCTGCAGCAGTGGGCATAAAAACTTACGAAATATGTGAGTCGCAGACGGGGTATTTATGGCGATTTAAAATCCATGCCCATAAGGCTTCGCCTACAGTCTCAGAGGCTGATCCATTCACAGCGAGTACACCGGCGTTGGTTCTTAACCTTATCAAAGGTTTAGAACACAAAGGATACACGTTGTGGATGGATAACTTCTACAATTCCCCAGCACTTGCCCGAAAACTCAAATCGATTGGGTTTGATTGTGTCGGTACGTTGCGGACAAACCGCAAGTATGTCCCGACGGAACTGACCAATCTGAAAAAATCTCAGATGAAGCCCGGGCAAGTAGTAGGCTATACCAGTGGAGATGTCGATTGCATAATATGGAGAGATCAGAACCGCGTAGCGACGATCTCAACGTACCATGGCAATGCCGTCTCCACTAAAAACGGAGTAACAAAACTTATTTTGATACGTGACTACAACATCTGCATGGGTGGCGTGGATAAAAAAGATCAGATGTTGGCTGCATTCCCAATTGAACGCAAGAGGACACAAATTTGGTACAAGAAATTGTTTAAAAGGAGAACACAAACTGGAAAATGCAGTGCAGGACGCCCTCGGGCCCGGTGGAGTGATGATCTGTGCAGGGTGGCTGGCAGGAACTGGATGAGTGAAACCGAGGATCGTGCTCAGTGGCGAACAATTGGAGAGGCTCATGTCCAGCAGTGGACTGCTATAGACTGA
Protein
MERFNLIAIEDLPAGEPIEQALTERSTLNVAAVVRARHLLDIIYRFLYMAAVVLRRHINVCVDRFAQTKNMILRSDDDDSDDESQNDNDEIVPMIIPRAAGILMDTPDDEEIEIDTAAATKPLVTSSAPSIISAPTSPARDTGMKPSHFFEFDWGTFPDSPIPPTERRESFKEHSGPTVAVSDPYEIFRLIWDQEFMEFIVQETNRYAQQLAAEMLDGGELQASSRITEWKETNVDELLVFFGILLAMGIVIKNRVEEYWNTEQNIFSTPGFKVYMSLRRFQLLSRCLHFNNSENLRNLNLDPSQAKLFKVEPVISHLNSKFTELYIMKQNIALDESLLQWKGWLNINQFIPNKAAAVGIKTYEICESQTGYLWRFKIHAHKASPTVSEADPFTASTPALVLNLIKGLEHKGYTLWMDNFYNSPALARKLKSIGFDCVGTLRTNRKYVPTELTNLKKSQMKPGQVVGYTSGDVDCIIWRDQNRVATISTYHGNAVSTKNGVTKLILIRDYNICMGGVDKKDQMLAAFPIERKRTQIWYKKLFKRRTQTGKCSAGRPRARWSDDLCRVAGRNWMSETEDRAQWRTIGEAHVQQWTAID
Summary
Uniprot
EMBL
Proteomes
Pfam
PF13843 DDE_Tnp_1_7
Interpro
IPR029526
PGBD
ProteinModelPortal
Ontologies
GO
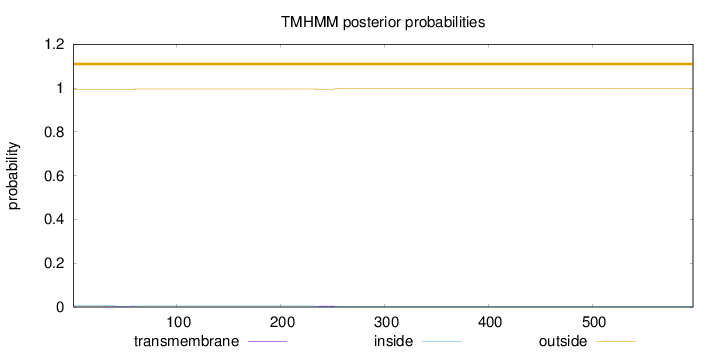
Topology
Length:
597
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.15071
Exp number, first 60 AAs:
0.05759
Total prob of N-in:
0.00646
outside
1 - 597
Population Genetic Test Statistics
Pi
183.726399
Theta
142.387758
Tajima's D
-0.64459
CLR
2.311419
CSRT
0.207739613019349
Interpretation
Uncertain
