Gene
KWMTBOMO08018
Pre Gene Modal
BGIBMGA010315
Annotation
Gag-like_protein_[Operophtera_brumata]
Location in the cell
Nuclear Reliability : 2.546
Sequence
CDS
ATGAGAAGCAATCACTTCAAAAGTATTGATGGAGGGCCGCGTGGACCCTCAGCTGTTAGAAAACGCTCACGCGACAATGCTCCAGTGGGAGATGGGTCAGTGGACCTGTCCTCAGATGAGGAAGGCCTGCTGCCCCCGAAGAAATTGCCGACCACGAGAAGAGTTCGGCAGGTTTCGAGAGGGATGAGTGCTTCACGGCCGCGGAGGAACGCACAGGGTAGGTTCGTGTGCTTCAATACTCCGGCGGCCAGTAAAGCAGATAGTGACATGGGGGTCACTACCGAAGACCCTGGTGGCGAGAGCTGTGCCTCGCTAGGGAGCTCAAAGGCGGAGATCAACGCCGCCAGAAGAGAGCAGCGCAAAGCCGTCGCGGCCGACGAACTTGCCTTGGACGGGGTGGACTTAGTTCTGAAGGTTGCCACCAAATCCGGCAGCCTGAAGGGTACGTTCACCCGCGGTCTGAAGGAAGCTGCCGCAGACATTAAGGAGGCGATTGGCATCCTCCTCAACAGGACGGCCTCTGACGAGGTTGCGAAGTTGCAGGAGGAAAACAGCCGTCTCCGAAATGATATGGAGGACCTCCGGCGACGAGTCACTGCGTTGAGCGAGCAGCAGCAGCGACGTACGTCCACTGATGCCGCATCGGTAGTGGCCCCGGCTCTCGCACCGAGACCGGTAAGCACTCATACCGACGATGAGGTCGAGCGCATTGTCCGGCTCTGCATGCTCCAGTGCGGGAGCATGGTCAATGCTCGTATGGAGGCAATTTCTCGGCGCCTCCCTGCGGAAATCCTCCGTCCGCCACTAGCGGCCGATACACGGCGGAGGGTTGAGGAGCCGCCGAGACCTAAGCCTAGGGAGGGGAAACCCGCGGAGGGTGTTAAAAAGCCCGTCGAGGGAGCCCCATCGAGCGATCAGCCCACGACTGCTGGGTCTAAGGGTGAGACCTGGGTTACAGTCATGGGCCGCAAGAAGGCACGCAAGGCCGCCAAGGCGGCCTCAGCAGCAACGCATGCCCCCGGCCAAACCGCAAAGGCGGTTGCGCAGCCTGCGCGGCGGACTGCCAAAGGCGGTCGCAAAGGACTGGTGATACGTGCTCCACGATCCGAAGCTGTGACGCTCACGCTACAGCCCGGAGCTGCGGAGCGCGGCGTAACGTACCAGTCGGTCATCGCCGAAGCCAAGGCCAAGATCAAATTATCAGATCTTGGTCTTCAGGCCGCCACCCGCAGGCAGGCTGCCACGGGTGCACGGCTGTTCGAGGTGGCTGGTACTGCGAGTGGCAGTGCCGAAAAGGCGGACGCTCTGGCCGCCAAGATGAGGGAGGTCCTCAGCCCCGAGGACGTCCGGGTCTCCAGGCCAATGAAAACCGCAGAGGTGCGGATTGCTGGCCTGGATGACTCCGTGACCTCTGAGGAGGTGGTAGCGGCCGTTGCCCGAAGTGGAGAGTGTCCATCGGACAAGGTGCGGGCCGGTGATATACGCACCGACGCCACCAGACTCGGCGTGGTCTGGGTTCGGTGCCCTGTAGCGTCGGCGAAAAAGATCGCCTTAAGCGGCAGATTGCTGGTTTTGCAGTCCGGGTCGTTTCGTAATGGCGATGTGCCATAA
Protein
MRSNHFKSIDGGPRGPSAVRKRSRDNAPVGDGSVDLSSDEEGLLPPKKLPTTRRVRQVSRGMSASRPRRNAQGRFVCFNTPAASKADSDMGVTTEDPGGESCASLGSSKAEINAARREQRKAVAADELALDGVDLVLKVATKSGSLKGTFTRGLKEAAADIKEAIGILLNRTASDEVAKLQEENSRLRNDMEDLRRRVTALSEQQQRRTSTDAASVVAPALAPRPVSTHTDDEVERIVRLCMLQCGSMVNARMEAISRRLPAEILRPPLAADTRRRVEEPPRPKPREGKPAEGVKKPVEGAPSSDQPTTAGSKGETWVTVMGRKKARKAAKAASAATHAPGQTAKAVAQPARRTAKGGRKGLVIRAPRSEAVTLTLQPGAAERGVTYQSVIAEAKAKIKLSDLGLQAATRRQAATGARLFEVAGTASGSAEKADALAAKMREVLSPEDVRVSRPMKTAEVRIAGLDDSVTSEEVVAAVARSGECPSDKVRAGDIRTDATRLGVVWVRCPVASAKKIALSGRLLVLQSGSFRNGDVP
Summary
Uniprot
EMBL
JTDY01000314
KOB77738.1
NWSH01001765
PCG70205.1
AGBW02010526
OWR48347.1
+ More
AB078928 BAC06449.1 JTDY01004969 KOB67533.1 D85594 BAA19775.1 JTDY01002012 KOB72329.1 NWSH01003254 PCG66687.1 NWSH01006437 PCG63434.1 NWSH01002890 PCG67363.1 NWSH01000260 PCG77924.1 JTDY01000669 KOB76277.1 NWSH01000688 PCG74775.1 ODYU01001274 SOQ37269.1
AB078928 BAC06449.1 JTDY01004969 KOB67533.1 D85594 BAA19775.1 JTDY01002012 KOB72329.1 NWSH01003254 PCG66687.1 NWSH01006437 PCG63434.1 NWSH01002890 PCG67363.1 NWSH01000260 PCG77924.1 JTDY01000669 KOB76277.1 NWSH01000688 PCG74775.1 ODYU01001274 SOQ37269.1
Proteomes
PRIDE
Pfam
PF00098 zf-CCHC
Gene 3D
ProteinModelPortal
Ontologies
KEGG
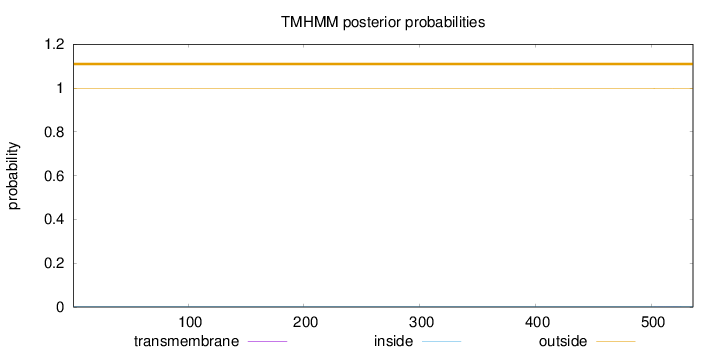
Topology
Length:
536
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00611
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00279
outside
1 - 536
Population Genetic Test Statistics
Pi
3.53614
Theta
15.39417
Tajima's D
-1.516278
CLR
0.294352
CSRT
0.0543972801359932
Interpretation
Uncertain
