Gene
KWMTBOMO07916
Annotation
reverse_transcriptase_[Danaus_plexippus]
Location in the cell
Mitochondrial Reliability : 1.622 PlasmaMembrane Reliability : 1.103
Sequence
CDS
ATGGACGAGGGGTGGAACGTGGTGGCCCGGCGGGGTAACAAGAAGGCCCCATTGACAGCTCCACAAGTGGGTGGAGTGTCGGCGGCCACGGCGGCTCAGGCTGCTCCCCAACAGGGTCGAGCGCCTGTAGCCGCCAAAAATAAAAATAAAAACAAAAAAACAAAGAAGCCACAGCGCGCGCCGCGATCGGCGGCAGTGGTCCTAACGCTGCTGCCGGCTGCCAAAGCAAAGGGCATGACATACGAGATGCAGACAGCCAATGGGGCCAAGCTGCTAGAATGTCCCGTTGCCGACAGTCCCGGCATCGCGGAGCGTCTGGCGGCAAGGCTCCGTATGGTTCTGCCTGAAGAGGAGGTGCGGGTGCTTAGGCCGGTCAAGATGGCTGAATTGAGAGTGACCGGCCTGGACGACTGCACCACCAGGGGTGAGGTGGCGGCGGCCGTCGCTGCCCAGGGCAACTGTGCCCCGCAGAATATAACCGTAGGCGAGCTGCGCTCTGGTTATACCGGGGCCGGCTCCGTGTGGGTACGTTGCCCGGTGGAGGCGGCCGTCGCCCTGGCTGCTCCTCCGCCTGGGAGACCGTCGGGGGATCCGGGCAGGTTGCGCGTGGGCTGGATAGCAGCCCACGTGCGCCTGTTAGAACCACGTGCCTGGCGGTGCCTAAGGTGCTTCAGCACTGGGCACTGCCTCGCCAGGTGCGTGAGTGCGGTTGACCGCAGTGGACTGTGTTTCCGCTGTGGTCAGCCTGGACACAAGGCGGCCGTGTGTGTGGCTGCGCCGCATTGTTCATTATGCGCGGCGGCCGGACGCAAGGCTGATCACCGGACCGGGGGCAAGGCATGCCCCCCGGCCGGCAAAAATGCAATTCGAAATTGGCGCCGCCAAGCTAAGCGGCGCCAAAAGAAGCGGTCGGCCGGGGGAGCACCGGCTGACACTTTGGCCCCCGCTGAGGCGTTCGAGCCCGATACCGGGGACAATAGAGTAGGGGAGGGGGCGATGGATGACATGCTGCTCCAGACTATGGCGGAGTGGTCAATTGACGTGGCTGTGGTCGCCGAGCCGTACTTCGTTCCCCCAGCGGACGACTGCTGGTTTGGGGACGTTGATGGCTTGGCGGCCATACATATAAGAAGATCCGCGATGATTCCTCCCCTGGCGATGGTGACCAGGGGCCCCGGGATCGTCGCGGTACAACTAGAAAATACCGTCGTGATCGGGGTGTACTTCTCTCCAAATCGGCCTACCGTCGAGTTCGAGCGGTTTCTGGACGGGCTAGAGGTGATCGCTCGTCACCTCTCTCCCCGTTCGGTGATACTGGCGGGGGACTTCAATGCGAAGTCTGTCGCTTGGGGTTCCCCATCCACGGACGCTCGTGGTAGGCTGTTGGAGGAGTGGGCGGTCGCGGCCGATCTCTGTATCGTTAACAGGGGCTCGGTCGTGACATGCGTGCCTTGGATGGGAGAGTCAATCATGGACTTGACGTTCGCGAGCTCGTCCATCGCACAGCGCATCCTGGGCTGGGGCGTGGTGGTGGGGGCGGAATCGTTGTCGGATCACCGGTATATCCGGTTCGATCTTTCCGCTCCCGCGCCGACGCCGGCCGCAGGCGCTCGCGGGGTTAACCCCCCGCGGAGTGCGTCCCGATCATTCCCTAGGTGGGCATTGAAGCTCCTGAAGAAGGCGCTCCTGGTGGAGGCCTCTATAGTGGCGGCGTGGACGCCCATGCGTCCACACCCTTTCGACGTGGAGGCTGAGGCCGAATGGTTCCGCGGCGCGATGCACCGCATATGCGATGCCTCGATGCCCCGGGTCGGCCCTCGGCGTTGTGGACGGCGGCAGTCGCCCTGGTGGTCGCCCGAAATTGCGAGACTGCGTGCAGTCTCCATTCGGGCGAGCCGCCGGTGCGCTAGACACCGCCGTCGCCGCCACTTGCGGCGCGACGGCCCCGTCGCGTTCGCGGAGGAGGAGACCCGGCTGCACGGCGAATATCGTGCTGCGAAGGTCGCCCTGCGGCTGGCCATCAGGCGGGCCAAGGAGCAGAACATGGAGAGACTCTTGGAGGCGCTCGAAGCGGATCCATGGGGGAGCCCGTATAGGCTGGTGCGCGGCAAACTGCGCCCGTGGCCCCCCCCTTTGACTGAGCGACTTCAGCCTCAACAGCTGCGGGACATCGTTTCGGCGTTGTTCCCGCAGGAGCGGGAGGGTTTTGTTCCCCCTGCTATGGGTGAGCCGTCGGACGCCATCGACGGCGAAGCCCCTGCTGAGGTGCCCCGTATCACTGGGGCAGAGCTCCGTGTGGCCGTTGTCAGGATGACTGCAAAGGACACGGCCCCCGACCCGGACGGAGTCCATGGCCGGGTGCTGGCCTTGGCCCTCGATGCCCTGGGGGACCGGCTCCTGGAGCTATTTAATGGTTGCCTGGAGTCGGGACGGTTTCCGTTGCTCTGGCGGACGGGCAGACTTGTGTTGTTGAGAAAGGAGGGGCACCCGGTGGATACAGCCGCCGGGTATCGTCTTATCGTACTGCTGGACGAGGCGGGAAAATTGCTGGAGCGTATTCTGGCTGCCCGCATCGTTCAGCACCTGGTCGGGGTTGGACCTGACCTGTCGGCGGAGCAGTTCGGCTTCCGGGAGGGCCGTTCGACCGTGGATGCGATACTTCGCGTGCGGGCCCTCTCGGACGAGGCTGTCTCACGGGGTGGGGTGGCGCTGGCGGTGTCTCTTGACATCGCCAACGCATTTAACACCCTGCCCTGGTCTGTGATAGGGGGGGCACTAGAGAGGCATGGAGTGCCCCTCTACCTCCGCCGGCTAGTTGGCTCCTATTTGGAGGCCAGGTCGGTCGAATGTACCGGGTACGGTGGGACCCTTCACCGTTTTCCGGTCGTGCGTGGTGTTCCACAGGGGTCGGTTCTCGGACCCCTCTTGTGGAACATCGGGTACGACTGGGTGCTGAGGGGTGCCCTCCTCCCGGGCCTCCGTGTTATTTGTTACGCGGACGACACTTTGGTCGTGGCCCGGGGGATGATTATAGAGAGTCTGCCCGTCTCGCCACAGCGGGAGTGGCCCTCGTCGTCGGGAGGATAA
Protein
MDEGWNVVARRGNKKAPLTAPQVGGVSAATAAQAAPQQGRAPVAAKNKNKNKKTKKPQRAPRSAAVVLTLLPAAKAKGMTYEMQTANGAKLLECPVADSPGIAERLAARLRMVLPEEEVRVLRPVKMAELRVTGLDDCTTRGEVAAAVAAQGNCAPQNITVGELRSGYTGAGSVWVRCPVEAAVALAAPPPGRPSGDPGRLRVGWIAAHVRLLEPRAWRCLRCFSTGHCLARCVSAVDRSGLCFRCGQPGHKAAVCVAAPHCSLCAAAGRKADHRTGGKACPPAGKNAIRNWRRQAKRRQKKRSAGGAPADTLAPAEAFEPDTGDNRVGEGAMDDMLLQTMAEWSIDVAVVAEPYFVPPADDCWFGDVDGLAAIHIRRSAMIPPLAMVTRGPGIVAVQLENTVVIGVYFSPNRPTVEFERFLDGLEVIARHLSPRSVILAGDFNAKSVAWGSPSTDARGRLLEEWAVAADLCIVNRGSVVTCVPWMGESIMDLTFASSSIAQRILGWGVVVGAESLSDHRYIRFDLSAPAPTPAAGARGVNPPRSASRSFPRWALKLLKKALLVEASIVAAWTPMRPHPFDVEAEAEWFRGAMHRICDASMPRVGPRRCGRRQSPWWSPEIARLRAVSIRASRRCARHRRRRHLRRDGPVAFAEEETRLHGEYRAAKVALRLAIRRAKEQNMERLLEALEADPWGSPYRLVRGKLRPWPPPLTERLQPQQLRDIVSALFPQEREGFVPPAMGEPSDAIDGEAPAEVPRITGAELRVAVVRMTAKDTAPDPDGVHGRVLALALDALGDRLLELFNGCLESGRFPLLWRTGRLVLLRKEGHPVDTAAGYRLIVLLDEAGKLLERILAARIVQHLVGVGPDLSAEQFGFREGRSTVDAILRVRALSDEAVSRGGVALAVSLDIANAFNTLPWSVIGGALERHGVPLYLRRLVGSYLEARSVECTGYGGTLHRFPVVRGVPQGSVLGPLLWNIGYDWVLRGALLPGLRVICYADDTLVVARGMIIESLPVSPQREWPSSSGG
Summary
Uniprot
EMBL
AGBW02011529
OWR46521.1
LBMM01004272
KMQ92618.1
LBMM01012412
KMQ86235.1
+ More
LBMM01006996 KMQ90130.1 LBMM01005474 KMQ91485.1 LBMM01008278 KMQ89009.1 RSAL01003504 RVE40273.1 LBMM01008445 KMQ88871.1 LBMM01008226 KMQ89040.1 LBMM01003988 KMQ92922.1 LBMM01008902 KMQ88543.1 ABLF02011183 ABLF02041884 ABLF02018942 ABLF02041885 ABLF02013358 ABLF02013361 ABLF02054869 ABLF02011238 GGMR01019493 MBY32112.1 ABLF02030663 ABLF02041312 ABLF02041886 ABLF02018098 ABLF02019402 ABLF02059310 ABLF02027248 KQ971310 KYB29499.1 KYB29500.1 ABLF02018808 KQ972544 KXZ75869.1 ABLF02018758 ABLF02018759 ABLF02011927
LBMM01006996 KMQ90130.1 LBMM01005474 KMQ91485.1 LBMM01008278 KMQ89009.1 RSAL01003504 RVE40273.1 LBMM01008445 KMQ88871.1 LBMM01008226 KMQ89040.1 LBMM01003988 KMQ92922.1 LBMM01008902 KMQ88543.1 ABLF02011183 ABLF02041884 ABLF02018942 ABLF02041885 ABLF02013358 ABLF02013361 ABLF02054869 ABLF02011238 GGMR01019493 MBY32112.1 ABLF02030663 ABLF02041312 ABLF02041886 ABLF02018098 ABLF02019402 ABLF02059310 ABLF02027248 KQ971310 KYB29499.1 KYB29500.1 ABLF02018808 KQ972544 KXZ75869.1 ABLF02018758 ABLF02018759 ABLF02011927
Proteomes
Pfam
Interpro
Gene 3D
ProteinModelPortal
PDB
1WDU
E-value=3.08588e-09,
Score=152
Ontologies
GO
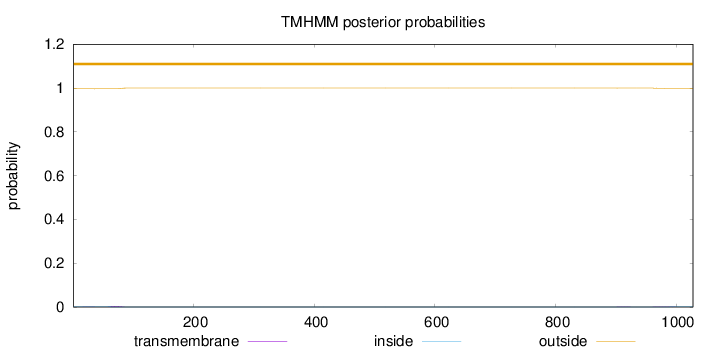
Topology
Length:
1028
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.10198
Exp number, first 60 AAs:
0.00781
Total prob of N-in:
0.00370
outside
1 - 1028
Population Genetic Test Statistics
Pi
0
Theta
0
Tajima's D
0
CLR
0.024691
CSRT
0
Interpretation
Uncertain
