Gene
KWMTBOMO07898
Annotation
PREDICTED:_uncharacterized_protein_LOC106720343_[Papilio_machaon]
Location in the cell
Mitochondrial Reliability : 1.577 Nuclear Reliability : 1.843
Sequence
CDS
ATGACTGTGACACAAAAGGCAGGAAGTGTTGTAGCGTCCGTGTACGACCGTGAGAGTGATGATAGGTATTTGCATTTACGGCCTATTTTCAAAATTCAAGTTCTCGGTAAACAGGGTACTGCACTTATAGATACAGCAGCTAAGCATTGTATTGCTGGTCACACGCTGTATGCTCTCCTTCTGCATAAGGGTCACCCTCTTACTCCCTCCACTAGGCGAGTCAAACTTGCTGATGGGCATGTTCGGAATATGGATGTATTGACAACCATATTACCAGTGAGGTTAGAACACATAGTGATTAATATTCCATTTATAATCTTCCCTGATTCTAATGATAATGAAACTTTACTGGGTATTGATTTTATTAATGCTGCCAAAGTTATTATTGATTTTCATCGTCATGAATGGTATTTTAGTACTGATGCTAAGGTGCATTATAAATTACTTTTTGAACCTATGTCTCGTGGTGTGTGCATAGCTTCTACTAACATGCTTCGGGATGATGAAGGCATGCACTTGCAATCAGCTGAGCGCCAAGCATTGTCTGAGTTCCTTATTCGGCATGAAAGTATTTTTACAGCAGGGGGAGGTCCGACACCATTCATAGAACATTGCATAGACACCGGAGATCACCCCCCGATAGCTGTGCCGCCCTATCGCCTTAACCCTTCCAAGAAGGAGACGATGAAGAATGGAATAGAGAAGATGCTGGCAGATGACATCATTGAAGAGTGCGAATCCGCCTGGTGCTCTCCTGCATTGATGATACCGAAGTCCAATGGTAATGTAAGATTCTGCGTGGATTACAGGCGCTTAAATGAAGTAACCAAATCGGATACTTACCCACTCCCACGCATTGATGATCTTCTCCAGAGTACGAAGAAGAATTGTTACATGACCACAATAGACCTTCGTTCTTCATATTGGCAAGTTATGGTCCGAGAAGCAGACAGAGACAAAACAGCTTTTGTCTGCCCCCTAGGCACTTACCGTTTTAAAAGAATGCCGTTCGGCTTGAAGAATGCGCCGGCGACTTTCCAAAGGCTTATTGATCGTCTACGCTCCTGTGCTGCTTTGAAGGATGTTACAGTACTTGCTTATCTAGACGATTTGTTAATTATTTCAGAAGGATTCCAGCAACACCTACAAGATCTGGAAGCTGTTTTTAGTCGTTTGTCTGAATTTAAACTTCACGTAAACAGAGAGAAGTGCACTTTTGCCAAAGAAAGGGTCCGTTATCTGGGCCACGTCATCACTCCAGACGGTGTGTCTCCTGACCCTGAGAAGGTCAGTGCTGTCCTTAATATGCTGGAACCATCTAACCTAAAACATTTAAAAACTTTTCTCCAAACATGCTCTTGGTTCAGGAAGTTTATTCCAAACTTTTCAAAGATAGCAGAACCGCTTACTCGACTGACCAAGAAGAATTACACATGGAGCTGGGGACCTGAACAAACACAGGCATTCAAAGAATTGAAGCGTCTGCTGACCACGGCGCCCGTACTTGTTCAAGCGAATTTTAAACAACCCTTAGTACTGCGTACCGACGCGAGCAACTATGCACTTGGCGCAGTTTTAGCTCAAGGGGAAGGCAAGGACGAAAGGCCTATCGAGTATGCTAGCCGCCTACTCACCGCAGCGGAGCGCAACTACTCTACTACAGAACGGGAAGCGCTTGCTGTAGTCTGGGCCGTGGAGCGTTTCAGGCCATACCTCGACGGCCAACCAATTATTATAGGCAGTGATCACCAACCCTTGCGTTGGTTGCTTTCTTTAAAGTCTCCTGCTGGTAGGTTGGTCCGTTGGGCGCTTAAACTCCAAGAATTTGACATCAGATTCGAGTACACCCCAGGAAAAGCTAACGTTGTCGCTGACACGTTAAGTAGACCGATCTGTTCCAACGAAACACAAAATAACTGCGGCATCTGTTCTGTCATTTGCGACCTGCCCACCAAGTCACCTACCCAGTTACGTCAAGACCAGTTGACTGATCCGGAAATACAGAAGATCGTCACAGAACTGGAAGGAGTGGATGAACTTGCTGCTAAAAGATGGTCAGAAAGAGGATTTCTAATGGAACAAGGTGTTTTATATCGACTTAATCCGGATTCTGATGCCGAGGCCCCACAATTAGTCATTCCTGCCCATCAAGTGACCGACGTTCTCAAAGAGCTTCACGATGCCCCTACCGCTGGACATGCAGGAATTGACCGGACTTACCAACAAGTCTCACGTCTGTTCTACTTCACAGGAATAAGAAGAATCATAACCGACTATGTAAAGGCATGTATCCATTGTCAACGCTATAAGGCTGCTAATACCAAGCCTCCAGGTCTATTGCAGACGCCGGTGATGAACCAGCGCAACGAAGTCTTGGCAGTTGATCTTTTCGGCCCGTTGCCACCTGGAAAACAGGGGGAGCGTTGGATCTTGCTTATAGAAGATACTGCTACGAGGTGGACAGAACTTTTCCCATTAAAGGAAGCTACTGCTGAGGCGTGTGCACATGTCCTCATAGAGGAGTATTTTATGAGGTTTGGTCTTCCACGCCGACTAGTTTCTGATAATGGCGTACAATTTATTTCGGCAGTCATGCGTCAATGCATGTCCATCCTGGGGATAAAGCAAAACCTTATCCCTCTCTATCACCCAGAAGCAAACCCCGCCGAACGTAAAAACAGAGATCTTAAGACTTTATTAGCACAACTCGTGGAATGTGACCACACTTCCTGGCCAAACATGTTACCAGTGATTCGATTCGCTCTTAATAACGCTAAATGTCGCACCACAGGTATGTCACCTGCTTACTTGTCATTTGGCAGAGAGATGAGGTCCCCCACAGAAGTCACCCATGACCTTCGTGCTGTTCTAGACAAAGATAACTTTGTTCCCCAAATAACTCCCTATCTCAGGAAATTTGTTAACTCCTTTTCAGCGGTGCGCGAACGTGTAGAGACGCTTCAAGATAAAGCTAAAGGGTATGCTGATCGTTCGAGACGGCCCATTGAAACATTCAACGAAGGTGACATGGTTCTCATCAAATCACATGTCCTCAGTAAGAGCGCTAATGGCCTTGCTTCTAAGTTTGTTCCAAAACGCGATGGTCCGTATCGCATCGTTAAAAAGGTCAGCCCTACTACTTACCACGTCGCGCATGTTGACAAACCTGATGAAGTTTTAGGGAAATATCATGTGAACGATCTCACTCTATACCGTGAGAATCAGAGTGATGCTCTTCCGCGGCCGGTGATGCCTAAAAAGAAACGTGGTCGTCCCCCAAATAATGTTAAAACAGATTCTCGAGTTCCAGGGCATATGACGACACGGCATGCTGAACCATGTGCTGAAGCTCCACGTCTTTCTCCCGCAGCTGGTGGTATGGTTGGTCTTGCGCCTGTCCCTCAGCCCAGTAGTAATGAGATGCTAGTCCAGGAGCGAGGGCGTCCTCCAGGACTAGAGGGGGAGTATGTAGCAAACCGACCGGTACGTCACTCTCGCGGCCGTATGCCCGCTCGCTATACATAG
Protein
MTVTQKAGSVVASVYDRESDDRYLHLRPIFKIQVLGKQGTALIDTAAKHCIAGHTLYALLLHKGHPLTPSTRRVKLADGHVRNMDVLTTILPVRLEHIVINIPFIIFPDSNDNETLLGIDFINAAKVIIDFHRHEWYFSTDAKVHYKLLFEPMSRGVCIASTNMLRDDEGMHLQSAERQALSEFLIRHESIFTAGGGPTPFIEHCIDTGDHPPIAVPPYRLNPSKKETMKNGIEKMLADDIIEECESAWCSPALMIPKSNGNVRFCVDYRRLNEVTKSDTYPLPRIDDLLQSTKKNCYMTTIDLRSSYWQVMVREADRDKTAFVCPLGTYRFKRMPFGLKNAPATFQRLIDRLRSCAALKDVTVLAYLDDLLIISEGFQQHLQDLEAVFSRLSEFKLHVNREKCTFAKERVRYLGHVITPDGVSPDPEKVSAVLNMLEPSNLKHLKTFLQTCSWFRKFIPNFSKIAEPLTRLTKKNYTWSWGPEQTQAFKELKRLLTTAPVLVQANFKQPLVLRTDASNYALGAVLAQGEGKDERPIEYASRLLTAAERNYSTTEREALAVVWAVERFRPYLDGQPIIIGSDHQPLRWLLSLKSPAGRLVRWALKLQEFDIRFEYTPGKANVVADTLSRPICSNETQNNCGICSVICDLPTKSPTQLRQDQLTDPEIQKIVTELEGVDELAAKRWSERGFLMEQGVLYRLNPDSDAEAPQLVIPAHQVTDVLKELHDAPTAGHAGIDRTYQQVSRLFYFTGIRRIITDYVKACIHCQRYKAANTKPPGLLQTPVMNQRNEVLAVDLFGPLPPGKQGERWILLIEDTATRWTELFPLKEATAEACAHVLIEEYFMRFGLPRRLVSDNGVQFISAVMRQCMSILGIKQNLIPLYHPEANPAERKNRDLKTLLAQLVECDHTSWPNMLPVIRFALNNAKCRTTGMSPAYLSFGREMRSPTEVTHDLRAVLDKDNFVPQITPYLRKFVNSFSAVRERVETLQDKAKGYADRSRRPIETFNEGDMVLIKSHVLSKSANGLASKFVPKRDGPYRIVKKVSPTTYHVAHVDKPDEVLGKYHVNDLTLYRENQSDALPRPVMPKKKRGRPPNNVKTDSRVPGHMTTRHAEPCAEAPRLSPAAGGMVGLAPVPQPSSNEMLVQERGRPPGLEGEYVANRPVRHSRGRMPARYT
Summary
Uniprot
A0A224XHC7
A0A2L2Y6Q3
A0A1Y1NDK0
D7EM19
A0A3B3IK80
A0A224X632
+ More
A0A3B5R366 A0A0V0G672 L7LYQ9 L7LYP4 L7LV95 V5I839 L7ML53 A0A1E1XKD7 L7LWL7 A0A3B1JJM2 A0A3P9MP57 L7MAW1 A0A0J7K8R1 L7LW11 A0A139W930 A0A3B3BX27 A0A3B5QFT0 A0A147BPL7 A0A224XHI1 A0A023EY78 A0A224XHE2 A0A096XIB4 A0A1E1X293 A0A1B6IL28 A0A147BUF5 A0A2R5L8F2 A0A3B3SLA7 A0A147BLY7 A0A0J7KPJ7 A0A2R5LEZ4 A0A147BJI2 A0A1Y1MZ56 A0A131XQ77 A0A0J7K4Z4 L7MEQ5 A0A146TD60 L7MFA7 A0A1Z5L5G8 A0A1Y1LZS5 A0A147BST7 A0A1Y1JVH7 L7MKE1 A0A2R5LP11 L7LY95 L7LV55 L7MMB3 A0A0J7KA76 L7MCS8 A0A131XRN7 L7MA76 A0A1E1XL81 A0A3P8Q5A9 L7MAT3 A0A131XL91 A0A1Y1MED9 W8B595 A0A0A9XMW3 L7LW12 A0A0J7KA70 A0A090XDG5 A0A023EY81 A0A087UDP9 A0A0J7K9T8 A0A1Y1NGJ5 H3A4B4 A0A147BIK4 L7LVQ9 A0A164KLJ0 A0A087SUZ9 A0A224XGU7 A0A0A9Z8Q1 A0A087TFE2 A0A0J7KF66 A0A087TFA8 A0A087U8V5 A0A087TH40 A0A164P0I2 A0A087TUL9 A0A087U9J6 A0A1Y1JW20 A0A087TRB6 A0A087U713
A0A3B5R366 A0A0V0G672 L7LYQ9 L7LYP4 L7LV95 V5I839 L7ML53 A0A1E1XKD7 L7LWL7 A0A3B1JJM2 A0A3P9MP57 L7MAW1 A0A0J7K8R1 L7LW11 A0A139W930 A0A3B3BX27 A0A3B5QFT0 A0A147BPL7 A0A224XHI1 A0A023EY78 A0A224XHE2 A0A096XIB4 A0A1E1X293 A0A1B6IL28 A0A147BUF5 A0A2R5L8F2 A0A3B3SLA7 A0A147BLY7 A0A0J7KPJ7 A0A2R5LEZ4 A0A147BJI2 A0A1Y1MZ56 A0A131XQ77 A0A0J7K4Z4 L7MEQ5 A0A146TD60 L7MFA7 A0A1Z5L5G8 A0A1Y1LZS5 A0A147BST7 A0A1Y1JVH7 L7MKE1 A0A2R5LP11 L7LY95 L7LV55 L7MMB3 A0A0J7KA76 L7MCS8 A0A131XRN7 L7MA76 A0A1E1XL81 A0A3P8Q5A9 L7MAT3 A0A131XL91 A0A1Y1MED9 W8B595 A0A0A9XMW3 L7LW12 A0A0J7KA70 A0A090XDG5 A0A023EY81 A0A087UDP9 A0A0J7K9T8 A0A1Y1NGJ5 H3A4B4 A0A147BIK4 L7LVQ9 A0A164KLJ0 A0A087SUZ9 A0A224XGU7 A0A0A9Z8Q1 A0A087TFE2 A0A0J7KF66 A0A087TFA8 A0A087U8V5 A0A087TH40 A0A164P0I2 A0A087TUL9 A0A087U9J6 A0A1Y1JW20 A0A087TRB6 A0A087U713
Pubmed
EMBL
GFTR01008566
JAW07860.1
IAAA01017336
LAA03833.1
GEZM01007731
JAV94970.1
+ More
KQ972234 EFA12514.2 GFTR01008489 JAW07937.1 GECL01003261 JAP02863.1 GACK01009040 JAA55994.1 GACK01009041 JAA55993.1 GACK01009043 JAA55991.1 GALX01005355 JAB63111.1 GACK01000334 JAA64700.1 GFAA01003652 JAT99782.1 GACK01009042 JAA55992.1 GACK01004027 JAA61007.1 LBMM01011454 KMQ86828.1 GACK01009039 JAA55995.1 KQ972733 KXZ75794.1 GEGO01002704 JAR92700.1 GFTR01008516 JAW07910.1 GBBI01004690 JAC14022.1 GFTR01008424 JAW08002.1 KF925840 AHN53418.1 GFAC01005811 JAT93377.1 GECU01030157 GECU01020076 JAS77549.1 JAS87630.1 GEGO01001242 JAR94162.1 GGLE01001609 MBY05735.1 GEGO01003620 JAR91784.1 LBMM01004568 KMQ92283.1 GGLE01003869 MBY07995.1 GEGO01004488 JAR90916.1 GEZM01020516 GEZM01020515 JAV89256.1 GEFM01006999 JAP68797.1 LBMM01013676 KMQ85528.1 GACK01002722 JAA62312.1 GCES01095459 JAQ90863.1 GACK01003086 JAA61948.1 GFJQ02004522 JAW02448.1 GEZM01046053 JAV77555.1 GEGO01001548 JAR93856.1 GEZM01099479 JAV53309.1 GACK01000363 JAA64671.1 GGLE01007069 MBY11195.1 GACK01009195 JAA55839.1 GACK01009547 JAA55487.1 GACK01000610 JAA64424.1 LBMM01010645 KMQ87338.1 GACK01004001 JAA61033.1 GEFM01006971 JAP68825.1 GACK01004074 JAA60960.1 GFAA01003693 JAT99741.1 GACK01003909 JAA61125.1 GEFH01002290 JAP66291.1 GEZM01035170 GEZM01035168 JAV83378.1 GAMC01021623 GAMC01021621 JAB84934.1 GBHO01041895 GBHO01025146 GBHO01025145 GBHO01025144 GBHO01025143 GBHO01014600 GBHO01014569 JAG01709.1 JAG18458.1 JAG18459.1 JAG18460.1 JAG18461.1 JAG29004.1 JAG29035.1 GACK01009237 JAA55797.1 LBMM01010896 KMQ87169.1 GBIH01000760 JAC93950.1 GBBI01004634 JAC14078.1 KK119366 KFM75488.1 LBMM01010852 KMQ87188.1 GEZM01005662 GEZM01005661 JAV96000.1 AFYH01233110 GEGO01004785 JAR90619.1 GACK01009357 JAA55677.1 LRGB01003288 KZS03371.1 KK112080 KFM56688.1 GFTR01008624 JAW07802.1 GBHO01002002 GBHO01001990 JAG41602.1 JAG41614.1 KK114971 KFM63831.1 LBMM01008414 KMQ88902.1 KK114965 KFM63797.1 KK118759 KFM73794.1 KK115210 KFM64429.1 LRGB01002748 KZS06406.1 KK116803 KFM68808.1 KK118857 KFM74035.1 GEZM01099475 JAV53313.1 KK116393 KFM67655.1 KK118517 KFM73152.1
KQ972234 EFA12514.2 GFTR01008489 JAW07937.1 GECL01003261 JAP02863.1 GACK01009040 JAA55994.1 GACK01009041 JAA55993.1 GACK01009043 JAA55991.1 GALX01005355 JAB63111.1 GACK01000334 JAA64700.1 GFAA01003652 JAT99782.1 GACK01009042 JAA55992.1 GACK01004027 JAA61007.1 LBMM01011454 KMQ86828.1 GACK01009039 JAA55995.1 KQ972733 KXZ75794.1 GEGO01002704 JAR92700.1 GFTR01008516 JAW07910.1 GBBI01004690 JAC14022.1 GFTR01008424 JAW08002.1 KF925840 AHN53418.1 GFAC01005811 JAT93377.1 GECU01030157 GECU01020076 JAS77549.1 JAS87630.1 GEGO01001242 JAR94162.1 GGLE01001609 MBY05735.1 GEGO01003620 JAR91784.1 LBMM01004568 KMQ92283.1 GGLE01003869 MBY07995.1 GEGO01004488 JAR90916.1 GEZM01020516 GEZM01020515 JAV89256.1 GEFM01006999 JAP68797.1 LBMM01013676 KMQ85528.1 GACK01002722 JAA62312.1 GCES01095459 JAQ90863.1 GACK01003086 JAA61948.1 GFJQ02004522 JAW02448.1 GEZM01046053 JAV77555.1 GEGO01001548 JAR93856.1 GEZM01099479 JAV53309.1 GACK01000363 JAA64671.1 GGLE01007069 MBY11195.1 GACK01009195 JAA55839.1 GACK01009547 JAA55487.1 GACK01000610 JAA64424.1 LBMM01010645 KMQ87338.1 GACK01004001 JAA61033.1 GEFM01006971 JAP68825.1 GACK01004074 JAA60960.1 GFAA01003693 JAT99741.1 GACK01003909 JAA61125.1 GEFH01002290 JAP66291.1 GEZM01035170 GEZM01035168 JAV83378.1 GAMC01021623 GAMC01021621 JAB84934.1 GBHO01041895 GBHO01025146 GBHO01025145 GBHO01025144 GBHO01025143 GBHO01014600 GBHO01014569 JAG01709.1 JAG18458.1 JAG18459.1 JAG18460.1 JAG18461.1 JAG29004.1 JAG29035.1 GACK01009237 JAA55797.1 LBMM01010896 KMQ87169.1 GBIH01000760 JAC93950.1 GBBI01004634 JAC14078.1 KK119366 KFM75488.1 LBMM01010852 KMQ87188.1 GEZM01005662 GEZM01005661 JAV96000.1 AFYH01233110 GEGO01004785 JAR90619.1 GACK01009357 JAA55677.1 LRGB01003288 KZS03371.1 KK112080 KFM56688.1 GFTR01008624 JAW07802.1 GBHO01002002 GBHO01001990 JAG41602.1 JAG41614.1 KK114971 KFM63831.1 LBMM01008414 KMQ88902.1 KK114965 KFM63797.1 KK118759 KFM73794.1 KK115210 KFM64429.1 LRGB01002748 KZS06406.1 KK116803 KFM68808.1 KK118857 KFM74035.1 GEZM01099475 JAV53313.1 KK116393 KFM67655.1 KK118517 KFM73152.1
Proteomes
PRIDE
Pfam
Interpro
IPR012337
RNaseH-like_sf
+ More
IPR001584 Integrase_cat-core
IPR041588 Integrase_H2C2
IPR036397 RNaseH_sf
IPR041373 RT_RNaseH
IPR000477 RT_dom
IPR021109 Peptidase_aspartic_dom_sf
IPR001878 Znf_CCHC
IPR041577 RT_RNaseH_2
IPR005162 Retrotrans_gag_dom
IPR001969 Aspartic_peptidase_AS
IPR003034 SAP_dom
IPR036875 Znf_CCHC_sf
IPR001995 Peptidase_A2_cat
IPR019103 Peptidase_aspartic_DDI1-type
IPR018061 Retropepsins
IPR001584 Integrase_cat-core
IPR041588 Integrase_H2C2
IPR036397 RNaseH_sf
IPR041373 RT_RNaseH
IPR000477 RT_dom
IPR021109 Peptidase_aspartic_dom_sf
IPR001878 Znf_CCHC
IPR041577 RT_RNaseH_2
IPR005162 Retrotrans_gag_dom
IPR001969 Aspartic_peptidase_AS
IPR003034 SAP_dom
IPR036875 Znf_CCHC_sf
IPR001995 Peptidase_A2_cat
IPR019103 Peptidase_aspartic_DDI1-type
IPR018061 Retropepsins
Gene 3D
ProteinModelPortal
A0A224XHC7
A0A2L2Y6Q3
A0A1Y1NDK0
D7EM19
A0A3B3IK80
A0A224X632
+ More
A0A3B5R366 A0A0V0G672 L7LYQ9 L7LYP4 L7LV95 V5I839 L7ML53 A0A1E1XKD7 L7LWL7 A0A3B1JJM2 A0A3P9MP57 L7MAW1 A0A0J7K8R1 L7LW11 A0A139W930 A0A3B3BX27 A0A3B5QFT0 A0A147BPL7 A0A224XHI1 A0A023EY78 A0A224XHE2 A0A096XIB4 A0A1E1X293 A0A1B6IL28 A0A147BUF5 A0A2R5L8F2 A0A3B3SLA7 A0A147BLY7 A0A0J7KPJ7 A0A2R5LEZ4 A0A147BJI2 A0A1Y1MZ56 A0A131XQ77 A0A0J7K4Z4 L7MEQ5 A0A146TD60 L7MFA7 A0A1Z5L5G8 A0A1Y1LZS5 A0A147BST7 A0A1Y1JVH7 L7MKE1 A0A2R5LP11 L7LY95 L7LV55 L7MMB3 A0A0J7KA76 L7MCS8 A0A131XRN7 L7MA76 A0A1E1XL81 A0A3P8Q5A9 L7MAT3 A0A131XL91 A0A1Y1MED9 W8B595 A0A0A9XMW3 L7LW12 A0A0J7KA70 A0A090XDG5 A0A023EY81 A0A087UDP9 A0A0J7K9T8 A0A1Y1NGJ5 H3A4B4 A0A147BIK4 L7LVQ9 A0A164KLJ0 A0A087SUZ9 A0A224XGU7 A0A0A9Z8Q1 A0A087TFE2 A0A0J7KF66 A0A087TFA8 A0A087U8V5 A0A087TH40 A0A164P0I2 A0A087TUL9 A0A087U9J6 A0A1Y1JW20 A0A087TRB6 A0A087U713
A0A3B5R366 A0A0V0G672 L7LYQ9 L7LYP4 L7LV95 V5I839 L7ML53 A0A1E1XKD7 L7LWL7 A0A3B1JJM2 A0A3P9MP57 L7MAW1 A0A0J7K8R1 L7LW11 A0A139W930 A0A3B3BX27 A0A3B5QFT0 A0A147BPL7 A0A224XHI1 A0A023EY78 A0A224XHE2 A0A096XIB4 A0A1E1X293 A0A1B6IL28 A0A147BUF5 A0A2R5L8F2 A0A3B3SLA7 A0A147BLY7 A0A0J7KPJ7 A0A2R5LEZ4 A0A147BJI2 A0A1Y1MZ56 A0A131XQ77 A0A0J7K4Z4 L7MEQ5 A0A146TD60 L7MFA7 A0A1Z5L5G8 A0A1Y1LZS5 A0A147BST7 A0A1Y1JVH7 L7MKE1 A0A2R5LP11 L7LY95 L7LV55 L7MMB3 A0A0J7KA76 L7MCS8 A0A131XRN7 L7MA76 A0A1E1XL81 A0A3P8Q5A9 L7MAT3 A0A131XL91 A0A1Y1MED9 W8B595 A0A0A9XMW3 L7LW12 A0A0J7KA70 A0A090XDG5 A0A023EY81 A0A087UDP9 A0A0J7K9T8 A0A1Y1NGJ5 H3A4B4 A0A147BIK4 L7LVQ9 A0A164KLJ0 A0A087SUZ9 A0A224XGU7 A0A0A9Z8Q1 A0A087TFE2 A0A0J7KF66 A0A087TFA8 A0A087U8V5 A0A087TH40 A0A164P0I2 A0A087TUL9 A0A087U9J6 A0A1Y1JW20 A0A087TRB6 A0A087U713
PDB
4OL8
E-value=3.0254e-72,
Score=696
Ontologies
GO
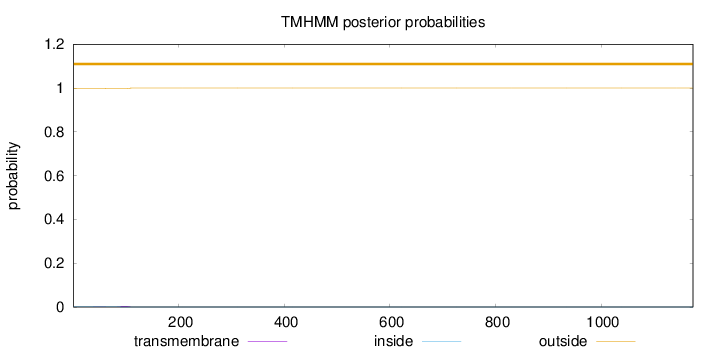
Topology
Length:
1174
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.0650199999999997
Exp number, first 60 AAs:
0.01371
Total prob of N-in:
0.00320
outside
1 - 1174
Population Genetic Test Statistics
Pi
169.673688
Theta
149.960327
Tajima's D
-0.775061
CLR
0
CSRT
0.181390930453477
Interpretation
Uncertain
