Gene
KWMTBOMO06101
Pre Gene Modal
BGIBMGA001727
Annotation
PREDICTED:_lysosomal_alpha-mannosidase-like_[Bombyx_mori]
Full name
Alpha-mannosidase
Location in the cell
PlasmaMembrane Reliability : 1.7
Sequence
CDS
ATGGTATACACGCTGCTACGTCAAGGTAGACTCTTTATCGTGGGCGGGGCATGGGGGATGGTGGACGAGGCCACCACCAACTACCACGCCGTGATCGATCAGTTCACGTACTCGCTAAGGAAACTCAACGCTACGTTCCTAGAGTGCGGGCGTCCGCTGATGGCCTGGCAGGCAGATGTCTATGGGCACTCCCGCGAGATCGCCTCGCTGCTGGCCCAGATGGGCTTCGATGGGCTCTTCATAAACCCGATCAGCTTCGACGATGAGCTGGACAGAATGAGGAGGAAATCCATGGAGTTCTTGTGGAGGGGAAGCGATGATTTAGGCGCGGAGACTGATATCTACACCCACAAGCTGTTCGACGGGTACTGGTCTCCGCCAGGGTTCTGCTTCGGGAGCTACTGCGATGACCCCCTCCTCATCACTTCGGACTCAGTCTTCAGAAACATCGACGAGCGGGTCGACCTGTTCATAAAGTCGATCCTCCACCGCCAAGCTCCCTACTACTCCACCAGGAATGTCATGGTGATGATGGGTCAGAGGTTCGGCTACTATGACGCCTCTATATGGTTCGCCAACGTGGATAAATTGATCAACGAAGTGAACAAAAAGAGTTATGAGACCGGAGCCAATATCAATCTGTATTATTCAACGCCGGCGTGCTACCTGAAAGCGGTCTATGACTCCAACCCGACTTTGGACACGAAACAGGATGACTTCTTCCCTATGGCCTACGACAAGTTCTCTCACGCCACCGGCCTGTATACGTCGCGACCCACCATAAAGTACATGGCGAGAGAGGGACATGTATACCTGCAGATGGCCAAGCAGCTTCAGGTCCTCGCCAGACTGTGGAACAACGACCAGCTCTTTGAAGAGTTTCAGTGGGTCATGGGTAATGTTTAG
Protein
MVYTLLRQGRLFIVGGAWGMVDEATTNYHAVIDQFTYSLRKLNATFLECGRPLMAWQADVYGHSREIASLLAQMGFDGLFINPISFDDELDRMRRKSMEFLWRGSDDLGAETDIYTHKLFDGYWSPPGFCFGSYCDDPLLITSDSVFRNIDERVDLFIKSILHRQAPYYSTRNVMVMMGQRFGYYDASIWFANVDKLINEVNKKSYETGANINLYYSTPACYLKAVYDSNPTLDTKQDDFFPMAYDKFSHATGLYTSRPTIKYMAREGHVYLQMAKQLQVLARLWNNDQLFEEFQWVMGNV
Summary
Cofactor
Zn(2+)
Similarity
Belongs to the glycosyl hydrolase 38 family.
Feature
chain Alpha-mannosidase
Uniprot
H9IWU4
A0A2A4JTB7
A0A212F2R7
A0A2A4JU30
A0A2A4JTC5
A0A2W1BD84
+ More
E2B6B9 A0A2H1VPQ4 A0A154P4N7 A0A195DWW0 A0A195F0Z8 A0A087ZZ79 B4G755 B3N5F1 A0A151IBC4 A0A158NP11 V5GCU3 A0A3B0JHP2 A0A0J9R086 A0A1W4UUP0 E2ABS8 A0A3G5BIG9 A0A0P8XGR3 Q8IPB7 A0A0Q5WN53 A0A3B0K7V8 A0A3B0JH41 A0A182KEA5 A0A0L7QJS0 Q9VKV1 B3MJI9 A0A0R1DR73 A0A1I8M2X1 B4Q9B2 A0A0A1WZV2 A0A1I8M2X3 A0A336LVA7 A0A336LYZ7 U5EV98 A0A1W4UG04 A0A1A9W6T1 A0A2J7RN15 A0A0K8WDF9 W8APD9 A0A182TAD7 A0A0K8VT60 A0A0K8U3N4 A0A0K8UEK7 A0A0K8VTI9 A0A0A1XEF2 A0A1I8QCY1 A0A026VXZ2 A0A1J1I212 A0A034V6I7 B4LVB7 A0A2J7RN33 B4KHR0 A0A1A9ZH71 A0A1B0B0M4 A0A1Q3FMS6 A0A0Q9XIT5 E9IED8 A0A1Q3FMY3 A0A1I8QD07 A0A1A9XCE3 A0A2H1VKW2 A0A182IPC8 A0A0M4ER29 B4JQ04 A0A1J1I5Z3 A0A2P8ZEB4 B4MUA9 A0A1S4HD42 Q178W0 A0A0M9A0P7 A0A0Q9WJC0 A0A023EWA7 A0A1W4WS90 A0A2A4IXJ1 A0A1S4FBS2 A0A1L8DQQ1 A0A1B0GPJ0 A0A1L8DR39 A0A182MB09 A0A084VFG4 A0A182RK94 A0A182WFL0 A0A1S4GYN8 A0A182VJW0 A0A182GN05 A0A3G5BIM3 A0A182UGZ7 A0A182KMH3 Q5TS83 A0A023EX89 A0A067R124 N6TNX4 W5JQY8
E2B6B9 A0A2H1VPQ4 A0A154P4N7 A0A195DWW0 A0A195F0Z8 A0A087ZZ79 B4G755 B3N5F1 A0A151IBC4 A0A158NP11 V5GCU3 A0A3B0JHP2 A0A0J9R086 A0A1W4UUP0 E2ABS8 A0A3G5BIG9 A0A0P8XGR3 Q8IPB7 A0A0Q5WN53 A0A3B0K7V8 A0A3B0JH41 A0A182KEA5 A0A0L7QJS0 Q9VKV1 B3MJI9 A0A0R1DR73 A0A1I8M2X1 B4Q9B2 A0A0A1WZV2 A0A1I8M2X3 A0A336LVA7 A0A336LYZ7 U5EV98 A0A1W4UG04 A0A1A9W6T1 A0A2J7RN15 A0A0K8WDF9 W8APD9 A0A182TAD7 A0A0K8VT60 A0A0K8U3N4 A0A0K8UEK7 A0A0K8VTI9 A0A0A1XEF2 A0A1I8QCY1 A0A026VXZ2 A0A1J1I212 A0A034V6I7 B4LVB7 A0A2J7RN33 B4KHR0 A0A1A9ZH71 A0A1B0B0M4 A0A1Q3FMS6 A0A0Q9XIT5 E9IED8 A0A1Q3FMY3 A0A1I8QD07 A0A1A9XCE3 A0A2H1VKW2 A0A182IPC8 A0A0M4ER29 B4JQ04 A0A1J1I5Z3 A0A2P8ZEB4 B4MUA9 A0A1S4HD42 Q178W0 A0A0M9A0P7 A0A0Q9WJC0 A0A023EWA7 A0A1W4WS90 A0A2A4IXJ1 A0A1S4FBS2 A0A1L8DQQ1 A0A1B0GPJ0 A0A1L8DR39 A0A182MB09 A0A084VFG4 A0A182RK94 A0A182WFL0 A0A1S4GYN8 A0A182VJW0 A0A182GN05 A0A3G5BIM3 A0A182UGZ7 A0A182KMH3 Q5TS83 A0A023EX89 A0A067R124 N6TNX4 W5JQY8
EC Number
3.2.1.-
Pubmed
19121390
22118469
28756777
20798317
17994087
21347285
+ More
22936249 30400621 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 17550304 25315136 25830018 24495485 24508170 30249741 25348373 21282665 29403074 12364791 17510324 24945155 24438588 26483478 20966253 24845553 23537049 20920257 23761445
22936249 30400621 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 17550304 25315136 25830018 24495485 24508170 30249741 25348373 21282665 29403074 12364791 17510324 24945155 24438588 26483478 20966253 24845553 23537049 20920257 23761445
EMBL
BABH01020972
NWSH01000615
PCG75285.1
AGBW02010691
OWR48030.1
PCG75286.1
+ More
PCG75287.1 KZ150423 PZC70926.1 GL445930 EFN88812.1 ODYU01003694 SOQ42778.1 KQ434813 KZC06792.1 KQ980167 KYN17373.1 KQ981891 KYN33784.1 CH479180 EDW28309.1 CH954177 EDV59030.2 KQ978112 KYM96921.1 ADTU01022012 ADTU01022013 ADTU01022014 GALX01000516 JAB67950.1 OUUW01000006 SPP81715.1 CM002910 KMY89556.1 GL438356 EFN69105.1 MK075175 AYV99578.1 CH902620 KPU73915.1 AE014134 AAN10754.1 KQS70602.1 SPP81716.1 SPP81714.1 KQ415041 KOC58867.1 AF145601 AAD38576.1 AAF52958.2 EDV32357.1 CM000157 KRJ97497.1 CM000361 EDX04554.1 KMY89555.1 GBXI01009935 JAD04357.1 UFQS01000218 UFQT01000218 SSX01504.1 SSX21884.1 SSX01503.1 SSX21883.1 GANO01001972 JAB57899.1 NEVH01002545 PNF42230.1 GDHF01003182 JAI49132.1 GAMC01015917 GAMC01015916 GAMC01015915 GAMC01015914 JAB90640.1 GDHF01010271 GDHF01002021 JAI42043.1 JAI50293.1 GDHF01030990 JAI21324.1 GDHF01027225 GDHF01012263 JAI25089.1 JAI40051.1 GDHF01010454 JAI41860.1 GBXI01004996 JAD09296.1 KK107648 QOIP01000008 EZA48346.1 RLU19783.1 CVRI01000038 CRK93802.1 GAKP01021245 JAC37707.1 CH940649 EDW63296.2 PNF42229.1 CH933807 EDW13347.1 JXJN01006781 GFDL01006180 JAV28865.1 KRG03704.1 GL762576 EFZ21090.1 GFDL01006179 JAV28866.1 ODYU01002940 SOQ41072.1 CP012523 ALC39282.1 CH916372 EDV98984.1 CRK93801.1 PYGN01000082 PSN54844.1 CH963857 EDW76035.2 AAAB01008944 CH477358 EAT42739.1 KQ435775 KOX74960.1 KRF81022.1 GAPW01000098 JAC13500.1 NWSH01004815 PCG64677.1 GFDF01005288 JAV08796.1 AJVK01005894 AJVK01005895 GFDF01005289 JAV08795.1 AXCM01004170 ATLV01012389 KE524787 KFB36708.1 JXUM01075505 KQ562889 KXJ74905.1 MK075237 AYV99640.1 EAL40242.3 GAPW01000077 JAC13521.1 KK853110 KDR11229.1 APGK01026882 KB740648 ENN79713.1 ADMH02000752 ETN65164.1
PCG75287.1 KZ150423 PZC70926.1 GL445930 EFN88812.1 ODYU01003694 SOQ42778.1 KQ434813 KZC06792.1 KQ980167 KYN17373.1 KQ981891 KYN33784.1 CH479180 EDW28309.1 CH954177 EDV59030.2 KQ978112 KYM96921.1 ADTU01022012 ADTU01022013 ADTU01022014 GALX01000516 JAB67950.1 OUUW01000006 SPP81715.1 CM002910 KMY89556.1 GL438356 EFN69105.1 MK075175 AYV99578.1 CH902620 KPU73915.1 AE014134 AAN10754.1 KQS70602.1 SPP81716.1 SPP81714.1 KQ415041 KOC58867.1 AF145601 AAD38576.1 AAF52958.2 EDV32357.1 CM000157 KRJ97497.1 CM000361 EDX04554.1 KMY89555.1 GBXI01009935 JAD04357.1 UFQS01000218 UFQT01000218 SSX01504.1 SSX21884.1 SSX01503.1 SSX21883.1 GANO01001972 JAB57899.1 NEVH01002545 PNF42230.1 GDHF01003182 JAI49132.1 GAMC01015917 GAMC01015916 GAMC01015915 GAMC01015914 JAB90640.1 GDHF01010271 GDHF01002021 JAI42043.1 JAI50293.1 GDHF01030990 JAI21324.1 GDHF01027225 GDHF01012263 JAI25089.1 JAI40051.1 GDHF01010454 JAI41860.1 GBXI01004996 JAD09296.1 KK107648 QOIP01000008 EZA48346.1 RLU19783.1 CVRI01000038 CRK93802.1 GAKP01021245 JAC37707.1 CH940649 EDW63296.2 PNF42229.1 CH933807 EDW13347.1 JXJN01006781 GFDL01006180 JAV28865.1 KRG03704.1 GL762576 EFZ21090.1 GFDL01006179 JAV28866.1 ODYU01002940 SOQ41072.1 CP012523 ALC39282.1 CH916372 EDV98984.1 CRK93801.1 PYGN01000082 PSN54844.1 CH963857 EDW76035.2 AAAB01008944 CH477358 EAT42739.1 KQ435775 KOX74960.1 KRF81022.1 GAPW01000098 JAC13500.1 NWSH01004815 PCG64677.1 GFDF01005288 JAV08796.1 AJVK01005894 AJVK01005895 GFDF01005289 JAV08795.1 AXCM01004170 ATLV01012389 KE524787 KFB36708.1 JXUM01075505 KQ562889 KXJ74905.1 MK075237 AYV99640.1 EAL40242.3 GAPW01000077 JAC13521.1 KK853110 KDR11229.1 APGK01026882 KB740648 ENN79713.1 ADMH02000752 ETN65164.1
Proteomes
UP000005204
UP000218220
UP000007151
UP000008237
UP000076502
UP000078492
+ More
UP000078541 UP000005203 UP000008744 UP000008711 UP000078542 UP000005205 UP000268350 UP000192221 UP000000311 UP000007801 UP000000803 UP000075881 UP000053825 UP000002282 UP000095301 UP000000304 UP000091820 UP000235965 UP000075901 UP000095300 UP000053097 UP000279307 UP000183832 UP000008792 UP000009192 UP000092445 UP000092460 UP000092443 UP000075880 UP000092553 UP000001070 UP000245037 UP000007798 UP000008820 UP000053105 UP000192223 UP000092462 UP000075883 UP000030765 UP000075900 UP000075920 UP000075903 UP000069940 UP000249989 UP000075902 UP000075882 UP000007062 UP000027135 UP000019118 UP000000673
UP000078541 UP000005203 UP000008744 UP000008711 UP000078542 UP000005205 UP000268350 UP000192221 UP000000311 UP000007801 UP000000803 UP000075881 UP000053825 UP000002282 UP000095301 UP000000304 UP000091820 UP000235965 UP000075901 UP000095300 UP000053097 UP000279307 UP000183832 UP000008792 UP000009192 UP000092445 UP000092460 UP000092443 UP000075880 UP000092553 UP000001070 UP000245037 UP000007798 UP000008820 UP000053105 UP000192223 UP000092462 UP000075883 UP000030765 UP000075900 UP000075920 UP000075903 UP000069940 UP000249989 UP000075902 UP000075882 UP000007062 UP000027135 UP000019118 UP000000673
Pfam
Interpro
Gene 3D
ProteinModelPortal
H9IWU4
A0A2A4JTB7
A0A212F2R7
A0A2A4JU30
A0A2A4JTC5
A0A2W1BD84
+ More
E2B6B9 A0A2H1VPQ4 A0A154P4N7 A0A195DWW0 A0A195F0Z8 A0A087ZZ79 B4G755 B3N5F1 A0A151IBC4 A0A158NP11 V5GCU3 A0A3B0JHP2 A0A0J9R086 A0A1W4UUP0 E2ABS8 A0A3G5BIG9 A0A0P8XGR3 Q8IPB7 A0A0Q5WN53 A0A3B0K7V8 A0A3B0JH41 A0A182KEA5 A0A0L7QJS0 Q9VKV1 B3MJI9 A0A0R1DR73 A0A1I8M2X1 B4Q9B2 A0A0A1WZV2 A0A1I8M2X3 A0A336LVA7 A0A336LYZ7 U5EV98 A0A1W4UG04 A0A1A9W6T1 A0A2J7RN15 A0A0K8WDF9 W8APD9 A0A182TAD7 A0A0K8VT60 A0A0K8U3N4 A0A0K8UEK7 A0A0K8VTI9 A0A0A1XEF2 A0A1I8QCY1 A0A026VXZ2 A0A1J1I212 A0A034V6I7 B4LVB7 A0A2J7RN33 B4KHR0 A0A1A9ZH71 A0A1B0B0M4 A0A1Q3FMS6 A0A0Q9XIT5 E9IED8 A0A1Q3FMY3 A0A1I8QD07 A0A1A9XCE3 A0A2H1VKW2 A0A182IPC8 A0A0M4ER29 B4JQ04 A0A1J1I5Z3 A0A2P8ZEB4 B4MUA9 A0A1S4HD42 Q178W0 A0A0M9A0P7 A0A0Q9WJC0 A0A023EWA7 A0A1W4WS90 A0A2A4IXJ1 A0A1S4FBS2 A0A1L8DQQ1 A0A1B0GPJ0 A0A1L8DR39 A0A182MB09 A0A084VFG4 A0A182RK94 A0A182WFL0 A0A1S4GYN8 A0A182VJW0 A0A182GN05 A0A3G5BIM3 A0A182UGZ7 A0A182KMH3 Q5TS83 A0A023EX89 A0A067R124 N6TNX4 W5JQY8
E2B6B9 A0A2H1VPQ4 A0A154P4N7 A0A195DWW0 A0A195F0Z8 A0A087ZZ79 B4G755 B3N5F1 A0A151IBC4 A0A158NP11 V5GCU3 A0A3B0JHP2 A0A0J9R086 A0A1W4UUP0 E2ABS8 A0A3G5BIG9 A0A0P8XGR3 Q8IPB7 A0A0Q5WN53 A0A3B0K7V8 A0A3B0JH41 A0A182KEA5 A0A0L7QJS0 Q9VKV1 B3MJI9 A0A0R1DR73 A0A1I8M2X1 B4Q9B2 A0A0A1WZV2 A0A1I8M2X3 A0A336LVA7 A0A336LYZ7 U5EV98 A0A1W4UG04 A0A1A9W6T1 A0A2J7RN15 A0A0K8WDF9 W8APD9 A0A182TAD7 A0A0K8VT60 A0A0K8U3N4 A0A0K8UEK7 A0A0K8VTI9 A0A0A1XEF2 A0A1I8QCY1 A0A026VXZ2 A0A1J1I212 A0A034V6I7 B4LVB7 A0A2J7RN33 B4KHR0 A0A1A9ZH71 A0A1B0B0M4 A0A1Q3FMS6 A0A0Q9XIT5 E9IED8 A0A1Q3FMY3 A0A1I8QD07 A0A1A9XCE3 A0A2H1VKW2 A0A182IPC8 A0A0M4ER29 B4JQ04 A0A1J1I5Z3 A0A2P8ZEB4 B4MUA9 A0A1S4HD42 Q178W0 A0A0M9A0P7 A0A0Q9WJC0 A0A023EWA7 A0A1W4WS90 A0A2A4IXJ1 A0A1S4FBS2 A0A1L8DQQ1 A0A1B0GPJ0 A0A1L8DR39 A0A182MB09 A0A084VFG4 A0A182RK94 A0A182WFL0 A0A1S4GYN8 A0A182VJW0 A0A182GN05 A0A3G5BIM3 A0A182UGZ7 A0A182KMH3 Q5TS83 A0A023EX89 A0A067R124 N6TNX4 W5JQY8
PDB
1O7D
E-value=1.71805e-38,
Score=398
Ontologies
PATHWAY
GO
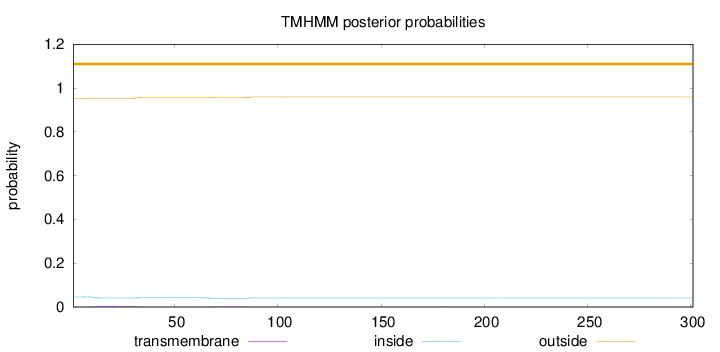
Topology
Length:
301
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.164310000000001
Exp number, first 60 AAs:
0.11201
Total prob of N-in:
0.04607
outside
1 - 301
Population Genetic Test Statistics
Pi
364.680997
Theta
196.743048
Tajima's D
2.560326
CLR
0.010169
CSRT
0.951402429878506
Interpretation
Uncertain
