Gene
KWMTBOMO05864
Annotation
PREDICTED:_LOW_QUALITY_PROTEIN:_uncharacterized_protein_LOC105841899?_partial_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 1.402
Sequence
CDS
ATGGACGCCACGTGGGGCACGCCTCCGCGTCGCCCCAGCGTTCCCGCTGCCGAGGAGCCCCCTCCCCCTATCACGGGGGCGGAGATCCATGCGGCCGTGTCCAGAATGAGAGCGAAGGACGCGGCCCCCGGCCCTGACGGTGTTCACGGCCGGGTCTGGAGCTTGGCCTTCGAGGCCCTAGGGGACCGGCTCGGGGGGCTTTTCGAAGCGTGCCTCGAGTCGGGACGGTTCCCGTCGAAATGGAAGACGGGCAGACTGGTCCTTCTGCGGAAGGACGGGCGCCCCGCGGACTCACCCGCGGGCTATCGCCCCATCGTGTTGCTGGACGAAGCGGGCAAGATGCTCGAGAGGATCGTCGCGGCCCGCATCGTCCGGCACCTGACCGAGACGGGTCCCGATCTTTCGGCGGAGCAATACGGCTTCAGAGAGGGCCGCTCGACTATCGACGCAATTCTGCGCGTGCGGGCCCTCTCCGATGAAGCCGTATCCCGGGGCGGGGTGGCGATGGCGTTATCCTTAGATGTAAGGAACGCCTTCAACACCCTGCCCTGGTCGGTGATCGCGGGGGCGCTGGAGTATCACGGCGTCCCCGCATACCTCCGCCGACTGATCGGCTCCTACCTCGAGGGCAGGTCGATCCAGTGCATCGGACACGGTGGGGCGATGTACCGCTTCCCCGTCGAGCGCGGTGTTCCGCAGGGGTCCGTCCTGGGCCCCCTGCTGTGGAACATCGGCTACGACTGGGTCCTGCGGGGCGTCATTCGGGGACCTCTCCCCGGGCTCAGCGTAATTTGTTACGCTGACGACACGTTGGTCGTAGCTCCGGGGAGGGACTACCGGGAGTCTGCCCGTCTGGCGTGCGCAGGCGTGGCACACGTCGTCACCAGGATCCGACGGTTGGGGCTCGAGGTGGCGCTCGACAAAACCCAGGCCCTGCTTTTTCACGGGCCGGGACGAGCGCCGCCTCTGGGTGCCCACCTCGTGATCGGAGGCGTCTGCGTCGGGGTCGGGGTGACCGGTCTTCGGTACCTCGGCCTCGAACTGGACGGTCGGTGGAACTTCCGCGCTCACTTTGAGAAGTTGGGCCCTCGGCTGATGGCAACGGCCGGCTCTTTGAGCCGGCTCCTTCCAAACGTCGGGGGTCCCGATCAGGTGGCGCGCCGCCTCTACATGGGGGTGGTGCGGTCGATAGCACTATACGGCGCGCCCGTATGGTGCCACGCCCTGACCCGCCAGAACGTCGCCGCGCTGCGACGTCCGCAGCGCGCGATCGCGGTCAGGGCGATCCGAGGATACCGCACCGTCTCCTTCGAGGCGGCGTGCTTGCTCGCCGGGGCGCCACCCTGGGACCTGGAGGCGGAGGCGCTCGCTGCCGATTACCGGTGGCGTAGCGACCTCCGCTCTAGGGGGGAAGGGCGCCCCAGCGAAGGTGTAGTTCGAGCGCGGAGGCTCCACTCTCGGCGGTCCGTGCTGGAGGCGTGGTCTCGCCGCTTGGCGGACCCGTCGGCCGGCCTCCGAACCGTCGAGGCGGTCCGCCCGGTTCTCGCGGACTGGGTGGGCCGCGACCGCGGATGCCTCACCTTCCGGCTAACGCAGATGCTTACCGGCCATGGTTGTTTCGGCCGGTACCTGCACAAGATAGCCGGAAGGGAACCGACGGCGCAGTGCCACCATTGCGCGGACCGCGACGAGGATGACACAGCGGAACACACGCTGGCGCGTTGCTCTGGGTTTGACGAGCAGCGCGCCGCCCTCGTCGCGGTCATTGGAGAGGACCTCTCGCTGCCGCGCGTCGTGGCTACGATGCTCGGCAGCGACGCGTCCTGGAAGGCGATGCTCGACTTCTGCGAGTCCACCATCTCGCAGAAGGAGGCGGCGGAGCGAGAGAGGGAGAGCTCTTCCCTTTCCGCACCGATCCGTCGCCGTCGAGCCGGGGGCCGGAGGCGGGAATACGCCCGCACGTCCCGGCCCCTGTAG
Protein
MDATWGTPPRRPSVPAAEEPPPPITGAEIHAAVSRMRAKDAAPGPDGVHGRVWSLAFEALGDRLGGLFEACLESGRFPSKWKTGRLVLLRKDGRPADSPAGYRPIVLLDEAGKMLERIVAARIVRHLTETGPDLSAEQYGFREGRSTIDAILRVRALSDEAVSRGGVAMALSLDVRNAFNTLPWSVIAGALEYHGVPAYLRRLIGSYLEGRSIQCIGHGGAMYRFPVERGVPQGSVLGPLLWNIGYDWVLRGVIRGPLPGLSVICYADDTLVVAPGRDYRESARLACAGVAHVVTRIRRLGLEVALDKTQALLFHGPGRAPPLGAHLVIGGVCVGVGVTGLRYLGLELDGRWNFRAHFEKLGPRLMATAGSLSRLLPNVGGPDQVARRLYMGVVRSIALYGAPVWCHALTRQNVAALRRPQRAIAVRAIRGYRTVSFEAACLLAGAPPWDLEAEALAADYRWRSDLRSRGEGRPSEGVVRARRLHSRRSVLEAWSRRLADPSAGLRTVEAVRPVLADWVGRDRGCLTFRLTQMLTGHGCFGRYLHKIAGREPTAQCHHCADRDEDDTAEHTLARCSGFDEQRAALVAVIGEDLSLPRVVATMLGSDASWKAMLDFCESTISQKEAAERERESSSLSAPIRRRRAGGRRREYARTSRPL
Summary
Similarity
Belongs to the AB hydrolase superfamily. Lipase family.
Uniprot
Q868Q4
Q8MY31
Q8MY33
Q8MY35
Q8MY27
A0A0J7N1D9
+ More
A0A2W1C3J5 O01419 A0A194R5D3 A0A3S2N998 A0A0J7KJG6 A0A0J7KIY5 A0A3S2LNH8 A0A0J7KL06 A0A0J7KF37 A0A0J7KJH4 A0A0J7KIM6 A0A0J7KLT9 A0A0J7KQK3 A0A0J7K3R6 A0A0J7KLH8 A0A0J7KYD2 A0A0J7KPR0 A0A0J7K3F1 A0A0J7KSR8 A0A0J7NAV8 A0A0J7KWU5 A0A0J7NMN5 A0A0J7KPZ3 A0A0J7KRB8 A0A0J7JZ13 A0A2S2N8I2 A0A0J7KIW1 A0A2S2PBP5 A0A0J7KF40 A0A0J7KW81 A0A0J7JZF7 A0A0J7N7S8 J9LJ80 A0A0J7N0R3 A0A0J7KMG5 A0A0J7KFH2 X1WJY4 A0A0J7MY17 A0A2S2PEZ7 A0A0J7NFZ0 A0A0J7KQY4 A0A0J7KMU1 A0A142LX45 A0A0J7NJI3 J9KJG8 J9KV04 A0A034WR48 A0A2S2NJ82 A0A2H8TIE5 A0A2H8TR36 A0A0J7MWN5 J9LVU7 X1WZ64 J9KNN8 A0A0J7K3R2 A0A2S2P488 J9KM84 J9KUM6 J9JWJ3 V5I8F3 W8ANK7 W8ADE4 J9LM37 X1WMW4 A0A2S2PEU2 A0A2M4ADW9 D6WC19 Q9N9Z1 A0A3L8D4C5 A0A139W9J0 J9KPP8 A0A2S2NQQ6 F7IYU8 J9JY94 A0A2M4BDI3 T1DG89 A0A2M4BBV9 A0A0J7MYL9 J9KNG8 A0A2S2P7D1 J9KNS1 A0A139W940 F7IYV2
A0A2W1C3J5 O01419 A0A194R5D3 A0A3S2N998 A0A0J7KJG6 A0A0J7KIY5 A0A3S2LNH8 A0A0J7KL06 A0A0J7KF37 A0A0J7KJH4 A0A0J7KIM6 A0A0J7KLT9 A0A0J7KQK3 A0A0J7K3R6 A0A0J7KLH8 A0A0J7KYD2 A0A0J7KPR0 A0A0J7K3F1 A0A0J7KSR8 A0A0J7NAV8 A0A0J7KWU5 A0A0J7NMN5 A0A0J7KPZ3 A0A0J7KRB8 A0A0J7JZ13 A0A2S2N8I2 A0A0J7KIW1 A0A2S2PBP5 A0A0J7KF40 A0A0J7KW81 A0A0J7JZF7 A0A0J7N7S8 J9LJ80 A0A0J7N0R3 A0A0J7KMG5 A0A0J7KFH2 X1WJY4 A0A0J7MY17 A0A2S2PEZ7 A0A0J7NFZ0 A0A0J7KQY4 A0A0J7KMU1 A0A142LX45 A0A0J7NJI3 J9KJG8 J9KV04 A0A034WR48 A0A2S2NJ82 A0A2H8TIE5 A0A2H8TR36 A0A0J7MWN5 J9LVU7 X1WZ64 J9KNN8 A0A0J7K3R2 A0A2S2P488 J9KM84 J9KUM6 J9JWJ3 V5I8F3 W8ANK7 W8ADE4 J9LM37 X1WMW4 A0A2S2PEU2 A0A2M4ADW9 D6WC19 Q9N9Z1 A0A3L8D4C5 A0A139W9J0 J9KPP8 A0A2S2NQQ6 F7IYU8 J9JY94 A0A2M4BDI3 T1DG89 A0A2M4BBV9 A0A0J7MYL9 J9KNG8 A0A2S2P7D1 J9KNS1 A0A139W940 F7IYV2
Pubmed
EMBL
AB090825
BAC57926.1
AB078931
BAC06456.1
AB078930
BAC06454.1
+ More
AB078929 BAC06452.1 AB078935 BAC06462.1 LBMM01012027 KMQ86470.1 KZ149896 PZC78733.1 D85594 BAA19776.1 KQ460685 KPJ12887.1 RSAL01000481 RVE41577.1 LBMM01006671 KMQ90407.1 LBMM01006954 KMQ90166.1 RSAL01003504 RVE40273.1 LBMM01006063 KMQ90967.1 LBMM01008448 KMQ88867.1 LBMM01006657 KMQ90417.1 LBMM01007036 KMQ90099.1 LBMM01005733 KMQ91252.1 LBMM01004272 KMQ92618.1 LBMM01015018 KMQ84954.1 LBMM01005948 KMQ91066.1 LBMM01002052 KMQ95336.1 LBMM01004634 KMQ92224.1 LBMM01015025 KMQ84953.1 LBMM01003575 KMQ93366.1 LBMM01007425 KMQ89740.1 LBMM01002480 KMQ94776.1 LBMM01003245 KMQ93755.1 LBMM01004505 KMQ92346.1 LBMM01003988 KMQ92922.1 LBMM01019839 KMQ83438.1 GGMR01000779 MBY13398.1 LBMM01006725 KMQ90373.1 GGMR01014195 MBY26814.1 LBMM01008422 KMQ88894.1 KMQ94772.1 LBMM01019838 KMQ83439.1 LBMM01008682 KMQ88705.1 ABLF02015311 ABLF02015320 ABLF02034009 ABLF02043484 LBMM01012412 KMQ86235.1 LBMM01005465 KMQ91492.1 LBMM01008278 KMQ89009.1 ABLF02011238 LBMM01014122 KMQ85315.1 GGMR01015159 MBY27778.1 LBMM01005474 KMQ91485.1 LBMM01004262 KMQ92649.1 LBMM01005216 KMQ91703.1 KU543683 AMS38371.1 LBMM01004261 KMQ92650.1 ABLF02041886 ABLF02004674 ABLF02004675 ABLF02004676 ABLF02011637 GAKP01002292 JAC56660.1 GGMR01004600 MBY17219.1 GFXV01002090 MBW13895.1 GFXV01003903 MBW15708.1 LBMM01015285 KMQ84870.1 ABLF02034925 ABLF02011183 ABLF02041884 ABLF02023257 LBMM01014956 KMQ84982.1 GGMR01011682 MBY24301.1 ABLF02018942 ABLF02041885 ABLF02041557 ABLF02013358 ABLF02013361 ABLF02054869 GALX01004785 JAB63681.1 GAMC01020517 JAB86038.1 GAMC01020519 GAMC01020515 JAB86040.1 ABLF02014650 ABLF02059310 GGMR01015354 MBY27973.1 GGFK01005660 MBW38981.1 KQ971309 EEZ99146.1 AJ278684 CAB99192.1 QOIP01000014 RLU15074.1 KQ972226 KYB24577.1 ABLF02009991 ABLF02009992 ABLF02054952 GGMR01006838 MBY19457.1 AB593321 BAK38639.1 ABLF02007989 GGFJ01001727 MBW50868.1 GALA01001776 JAA93076.1 GGFJ01001369 MBW50510.1 LBMM01013668 KMQ85535.1 ABLF02005203 ABLF02041316 GGMR01012748 MBY25367.1 ABLF02018808 KQ972691 KXZ75806.1 AB593323 BAK38643.1
AB078929 BAC06452.1 AB078935 BAC06462.1 LBMM01012027 KMQ86470.1 KZ149896 PZC78733.1 D85594 BAA19776.1 KQ460685 KPJ12887.1 RSAL01000481 RVE41577.1 LBMM01006671 KMQ90407.1 LBMM01006954 KMQ90166.1 RSAL01003504 RVE40273.1 LBMM01006063 KMQ90967.1 LBMM01008448 KMQ88867.1 LBMM01006657 KMQ90417.1 LBMM01007036 KMQ90099.1 LBMM01005733 KMQ91252.1 LBMM01004272 KMQ92618.1 LBMM01015018 KMQ84954.1 LBMM01005948 KMQ91066.1 LBMM01002052 KMQ95336.1 LBMM01004634 KMQ92224.1 LBMM01015025 KMQ84953.1 LBMM01003575 KMQ93366.1 LBMM01007425 KMQ89740.1 LBMM01002480 KMQ94776.1 LBMM01003245 KMQ93755.1 LBMM01004505 KMQ92346.1 LBMM01003988 KMQ92922.1 LBMM01019839 KMQ83438.1 GGMR01000779 MBY13398.1 LBMM01006725 KMQ90373.1 GGMR01014195 MBY26814.1 LBMM01008422 KMQ88894.1 KMQ94772.1 LBMM01019838 KMQ83439.1 LBMM01008682 KMQ88705.1 ABLF02015311 ABLF02015320 ABLF02034009 ABLF02043484 LBMM01012412 KMQ86235.1 LBMM01005465 KMQ91492.1 LBMM01008278 KMQ89009.1 ABLF02011238 LBMM01014122 KMQ85315.1 GGMR01015159 MBY27778.1 LBMM01005474 KMQ91485.1 LBMM01004262 KMQ92649.1 LBMM01005216 KMQ91703.1 KU543683 AMS38371.1 LBMM01004261 KMQ92650.1 ABLF02041886 ABLF02004674 ABLF02004675 ABLF02004676 ABLF02011637 GAKP01002292 JAC56660.1 GGMR01004600 MBY17219.1 GFXV01002090 MBW13895.1 GFXV01003903 MBW15708.1 LBMM01015285 KMQ84870.1 ABLF02034925 ABLF02011183 ABLF02041884 ABLF02023257 LBMM01014956 KMQ84982.1 GGMR01011682 MBY24301.1 ABLF02018942 ABLF02041885 ABLF02041557 ABLF02013358 ABLF02013361 ABLF02054869 GALX01004785 JAB63681.1 GAMC01020517 JAB86038.1 GAMC01020519 GAMC01020515 JAB86040.1 ABLF02014650 ABLF02059310 GGMR01015354 MBY27973.1 GGFK01005660 MBW38981.1 KQ971309 EEZ99146.1 AJ278684 CAB99192.1 QOIP01000014 RLU15074.1 KQ972226 KYB24577.1 ABLF02009991 ABLF02009992 ABLF02054952 GGMR01006838 MBY19457.1 AB593321 BAK38639.1 ABLF02007989 GGFJ01001727 MBW50868.1 GALA01001776 JAA93076.1 GGFJ01001369 MBW50510.1 LBMM01013668 KMQ85535.1 ABLF02005203 ABLF02041316 GGMR01012748 MBY25367.1 ABLF02018808 KQ972691 KXZ75806.1 AB593323 BAK38643.1
Proteomes
Pfam
Interpro
IPR000477
RT_dom
+ More
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR002123 Plipid/glycerol_acylTrfase
IPR012934 Znf_AD
IPR013087 Znf_C2H2_type
IPR036236 Znf_C2H2_sf
IPR024079 MetalloPept_cat_dom_sf
IPR001590 Peptidase_M12B
IPR001878 Znf_CCHC
IPR036875 Znf_CCHC_sf
IPR021896 Transposase_37
IPR029058 AB_hydrolase
IPR013818 Lipase/vitellogenin
IPR000734 TAG_lipase
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR002123 Plipid/glycerol_acylTrfase
IPR012934 Znf_AD
IPR013087 Znf_C2H2_type
IPR036236 Znf_C2H2_sf
IPR024079 MetalloPept_cat_dom_sf
IPR001590 Peptidase_M12B
IPR001878 Znf_CCHC
IPR036875 Znf_CCHC_sf
IPR021896 Transposase_37
IPR029058 AB_hydrolase
IPR013818 Lipase/vitellogenin
IPR000734 TAG_lipase
Gene 3D
ProteinModelPortal
Q868Q4
Q8MY31
Q8MY33
Q8MY35
Q8MY27
A0A0J7N1D9
+ More
A0A2W1C3J5 O01419 A0A194R5D3 A0A3S2N998 A0A0J7KJG6 A0A0J7KIY5 A0A3S2LNH8 A0A0J7KL06 A0A0J7KF37 A0A0J7KJH4 A0A0J7KIM6 A0A0J7KLT9 A0A0J7KQK3 A0A0J7K3R6 A0A0J7KLH8 A0A0J7KYD2 A0A0J7KPR0 A0A0J7K3F1 A0A0J7KSR8 A0A0J7NAV8 A0A0J7KWU5 A0A0J7NMN5 A0A0J7KPZ3 A0A0J7KRB8 A0A0J7JZ13 A0A2S2N8I2 A0A0J7KIW1 A0A2S2PBP5 A0A0J7KF40 A0A0J7KW81 A0A0J7JZF7 A0A0J7N7S8 J9LJ80 A0A0J7N0R3 A0A0J7KMG5 A0A0J7KFH2 X1WJY4 A0A0J7MY17 A0A2S2PEZ7 A0A0J7NFZ0 A0A0J7KQY4 A0A0J7KMU1 A0A142LX45 A0A0J7NJI3 J9KJG8 J9KV04 A0A034WR48 A0A2S2NJ82 A0A2H8TIE5 A0A2H8TR36 A0A0J7MWN5 J9LVU7 X1WZ64 J9KNN8 A0A0J7K3R2 A0A2S2P488 J9KM84 J9KUM6 J9JWJ3 V5I8F3 W8ANK7 W8ADE4 J9LM37 X1WMW4 A0A2S2PEU2 A0A2M4ADW9 D6WC19 Q9N9Z1 A0A3L8D4C5 A0A139W9J0 J9KPP8 A0A2S2NQQ6 F7IYU8 J9JY94 A0A2M4BDI3 T1DG89 A0A2M4BBV9 A0A0J7MYL9 J9KNG8 A0A2S2P7D1 J9KNS1 A0A139W940 F7IYV2
A0A2W1C3J5 O01419 A0A194R5D3 A0A3S2N998 A0A0J7KJG6 A0A0J7KIY5 A0A3S2LNH8 A0A0J7KL06 A0A0J7KF37 A0A0J7KJH4 A0A0J7KIM6 A0A0J7KLT9 A0A0J7KQK3 A0A0J7K3R6 A0A0J7KLH8 A0A0J7KYD2 A0A0J7KPR0 A0A0J7K3F1 A0A0J7KSR8 A0A0J7NAV8 A0A0J7KWU5 A0A0J7NMN5 A0A0J7KPZ3 A0A0J7KRB8 A0A0J7JZ13 A0A2S2N8I2 A0A0J7KIW1 A0A2S2PBP5 A0A0J7KF40 A0A0J7KW81 A0A0J7JZF7 A0A0J7N7S8 J9LJ80 A0A0J7N0R3 A0A0J7KMG5 A0A0J7KFH2 X1WJY4 A0A0J7MY17 A0A2S2PEZ7 A0A0J7NFZ0 A0A0J7KQY4 A0A0J7KMU1 A0A142LX45 A0A0J7NJI3 J9KJG8 J9KV04 A0A034WR48 A0A2S2NJ82 A0A2H8TIE5 A0A2H8TR36 A0A0J7MWN5 J9LVU7 X1WZ64 J9KNN8 A0A0J7K3R2 A0A2S2P488 J9KM84 J9KUM6 J9JWJ3 V5I8F3 W8ANK7 W8ADE4 J9LM37 X1WMW4 A0A2S2PEU2 A0A2M4ADW9 D6WC19 Q9N9Z1 A0A3L8D4C5 A0A139W9J0 J9KPP8 A0A2S2NQQ6 F7IYU8 J9JY94 A0A2M4BDI3 T1DG89 A0A2M4BBV9 A0A0J7MYL9 J9KNG8 A0A2S2P7D1 J9KNS1 A0A139W940 F7IYV2
PDB
6AR3
E-value=7.78633e-05,
Score=112
Ontologies
GO
PANTHER
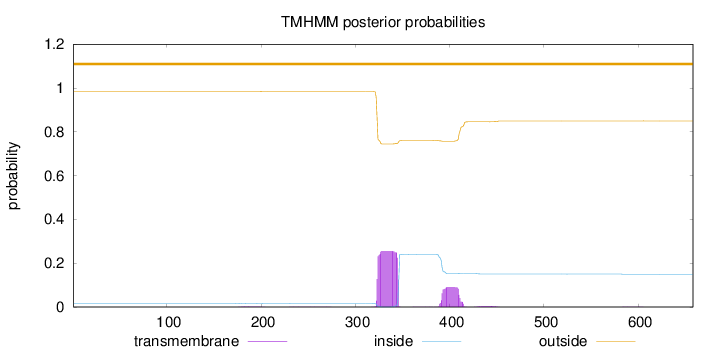
Topology
Subcellular location
Secreted
Length:
658
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
7.75467999999999
Exp number, first 60 AAs:
6e-05
Total prob of N-in:
0.01681
outside
1 - 658
Population Genetic Test Statistics
Pi
41.196992
Theta
2.716632
Tajima's D
2.344017
CLR
0.73485
CSRT
0.942652867356632
Interpretation
Uncertain
