Gene
KWMTBOMO05757
Pre Gene Modal
BGIBMGA006804
Annotation
PREDICTED:_tyrosine_decarboxylase_[Amyelois_transitella]
Location in the cell
Cytoplasmic Reliability : 1.065 PlasmaMembrane Reliability : 1.412
Sequence
CDS
ATGGATGTCGAAGAGTTTCGTGTGCGCGGTAAAGAGATGGTAGATTACATATGCACTTACATGAACACGTTGCGTACCCGGCGCGTTGTGCCCAGCGTGGAGCCAGGATACCTACGAACAGCACTTCCGGCCGAAGCTCCTCATCACCCCGAACCCTGGGACGATGTGATGAACGATGTCGAGCAAAAGATAATGCCAGGTGTAACACATTGGCAGCATCCACGTTTTCACGCATACTTTCCGGCAGGCAACGGGTACCCTTCGATACTCGGAGATATGTTGTCTGCTGGCATCGGATGCATTGGTTTTTCCTGGGCAGCAAGTCCCGCCTGTACTGAGCTAGAAATAATCATGTTGGATTGGATGGGAAAAGCGATTGGACTACCGCCTGCCTTTTTAGCGCTTGAAGAAGGTAGCAAAGGGGGCGGTGTTATCCAGGGATCTGCTAGTGAATGCGTTCTTGTATCTATGTTGGCTGCCAGAGCCAATGGAATTAAAAGATTAAAGCATCAATTTCCCAATGTTGACGAGGGACTTTTGCTATCAAAATTAATAGCATATTGTTCTAAGGAAGCCCATTCATGCGTCGAAAAGGCAGCCATGATTTGCTTTGTGAAACTTCGTATCTTACAGCCGGATGACAACGGTTCTCTTAGAGGTGAAACACTAAAGGAGGCTATGGAAGAAGATGAAGAGGCAGGGCGAGTACCGTTCTTTGTCGCTACGACATTGGGGACTACTTCATGTTGCTCCTTCGATAATTTGCCTGAAATAGGTCCCATGGTGCGGAAATTCCCTTGCGTCTGGCTTCACGTGGACGCCGCATATGCTGGAAGCTCATTCCTCTGCCCAGAACACAAGTATCACTTAGCTGGAGTTGAATATGCTGACTCCTTCAATACTAATCCTAATAAATTAATGCTTGCGAGTTTTGACTGTTCACTATTGTGGGTCACTAACCGCTATCTTCTAACATCTGCCCTCGTGGTTGATCCGTTGTATCTACAACATTGCTACGACCACACTGCTATCGATTATCGACACTGGGGTATATCGCTTAGTCGTCGTTTCCGTTCGCTTAAGCTATGGTTCATGTTGAGAAGTTACGGTATTAGCGGCTTGCAAAAATATGTCAGAAAGCATTGCGAACTGGCCAAATATTTTGAAAGGCTTGTCAAGAAAGACGATAGATTCCAAGTCTGCAATCAAGTAAGGTTGGGTTTGGTTTGCTTCCGCCTCGTTGGAGGTCCCGACGAGACTCCGGAGCAAATTGATGAGATGAACAAGAAACTCCTCACTAATATAAATGCATCTGGAAAACTGCATATGGTCCCGGCCTCTGTTCGGGAACGTTTTGTGATACGTTTCTGTGTGGTCGCTCAGCACGCTACACGTGAAGATATTGAAGTTGCTTGGGATATAATATCCGACTTTGCTACGGAGTTATTAGAGGGTCCGGATAAGGAAAGGGATCTAAACGAGGAACGTACTCGTCGCAATCGTGCCGCTCTAGCCCACAAACGTTCGTTCTTCGTACGTATGGTTAGCGATCCAAAGATCTATAATCCGGCTATCAACAAAACTCCACCACCGGTAGCTACATCTCCTCCTGGGTCACCACCCGCTGCTGCACCTTCCACGCCAGATACTCCATCCGACTCTGGCTCCATTCAACAGCCCACTACGCCGAAGCAATCGTCTTGGATAAGCTGGCCTCTTGGATTCTTTTTCCAGAGCGTTGATCACGACTCTAACGATTTGCCACTGAGATTTCGTCATCTCGACACGATGGTGAGGTTGAAGAGTCCCGGAGCAAGTCGTCGGGGCTCCACGCCGTGCGCCTCACCCGAACATCGGGCCTCGTCGCCCTCCAACAATCACTAA
Protein
MDVEEFRVRGKEMVDYICTYMNTLRTRRVVPSVEPGYLRTALPAEAPHHPEPWDDVMNDVEQKIMPGVTHWQHPRFHAYFPAGNGYPSILGDMLSAGIGCIGFSWAASPACTELEIIMLDWMGKAIGLPPAFLALEEGSKGGGVIQGSASECVLVSMLAARANGIKRLKHQFPNVDEGLLLSKLIAYCSKEAHSCVEKAAMICFVKLRILQPDDNGSLRGETLKEAMEEDEEAGRVPFFVATTLGTTSCCSFDNLPEIGPMVRKFPCVWLHVDAAYAGSSFLCPEHKYHLAGVEYADSFNTNPNKLMLASFDCSLLWVTNRYLLTSALVVDPLYLQHCYDHTAIDYRHWGISLSRRFRSLKLWFMLRSYGISGLQKYVRKHCELAKYFERLVKKDDRFQVCNQVRLGLVCFRLVGGPDETPEQIDEMNKKLLTNINASGKLHMVPASVRERFVIRFCVVAQHATREDIEVAWDIISDFATELLEGPDKERDLNEERTRRNRAALAHKRSFFVRMVSDPKIYNPAINKTPPPVATSPPGSPPAAAPSTPDTPSDSGSIQQPTTPKQSSWISWPLGFFFQSVDHDSNDLPLRFRHLDTMVRLKSPGASRRGSTPCASPEHRASSPSNNH
Summary
Cofactor
pyridoxal 5'-phosphate
Similarity
Belongs to the group II decarboxylase family.
Uniprot
A0A2A4JQI7
A0A2W1BU38
A0A0G3VFZ2
A0A2H1WFA6
H9JBA9
A0A194RC18
+ More
A0A194Q8T4 A0A212F653 A0A0L7LQ13 A0A3S2P4I8 D6X389 B0WT43 Q17F96 B4NMH2 A0A1S4G848 B4J6J2 B4KNI7 A0A0M4EAN4 B4LK06 A0A1W4VF47 A0A0J9R6G5 B4P0Z8 A1Z6N4 B3NB37 A0A084VQQ1 A0A3B0JNG0 B4HQA4 B4GC74 B3MIP9 Q292Z5 A0A0R3NKV6 A0A182PZH8 A0A182NKM3 A0A0R1DSU5 A0A182IWE7 A0A336MRT6 A0A182XYJ0 A0A182RU01 A0A182VJL2 A0A182WZ30 A0A182KQV6 Q7PTH4 A0A182WJK6 A0A182UH83 A0A182P203 A0A0A1WWU3 A0A1I8N2R7 A0A0L0C2L5 A0A182FG49 A0A1I8P932 W8VJW9 A0A034V1P5 T1I5A1 W8BSM1 X2KSD2 A0A1J1IUG8 A0A182I2X6 A0A1A9YC03 E0VK92 W8C9H5 U4UGN6 N6TFJ6 E2ALW7 A0A088AVR5 A0A2A3ERL5 A0A195BWL8 A0A158NCJ3 A0A182MSY0 E9IM88 A0A195CQE1 A0A158NCJ2 A0A336MRS6 A0A0M8ZRE3 A0A026WVU1 A0A3L8DI55 E2BHY3 A0A1A9ZUY7 A0A1B0FC75 A0A1B0GJF5 A0A1A9W101 J9M312 A0A1A9UU25 A0A195F967 A0A151X5T7 A0A0L7RJA9 A0A3R7PEH3 F4WX53 A0A067QQI2 A0A1D1VF32 A0A2P8ZK26 T1IJC5 A0A2J7QH10 A0A1Y1KDP2 A0A0T6BC19 A0A1W4XJ75 A0A1Y3BWD0 A0A1W0WSW7 A0A0P5BGP2 A0A0P6GS82 A0A0P6I7Q2
A0A194Q8T4 A0A212F653 A0A0L7LQ13 A0A3S2P4I8 D6X389 B0WT43 Q17F96 B4NMH2 A0A1S4G848 B4J6J2 B4KNI7 A0A0M4EAN4 B4LK06 A0A1W4VF47 A0A0J9R6G5 B4P0Z8 A1Z6N4 B3NB37 A0A084VQQ1 A0A3B0JNG0 B4HQA4 B4GC74 B3MIP9 Q292Z5 A0A0R3NKV6 A0A182PZH8 A0A182NKM3 A0A0R1DSU5 A0A182IWE7 A0A336MRT6 A0A182XYJ0 A0A182RU01 A0A182VJL2 A0A182WZ30 A0A182KQV6 Q7PTH4 A0A182WJK6 A0A182UH83 A0A182P203 A0A0A1WWU3 A0A1I8N2R7 A0A0L0C2L5 A0A182FG49 A0A1I8P932 W8VJW9 A0A034V1P5 T1I5A1 W8BSM1 X2KSD2 A0A1J1IUG8 A0A182I2X6 A0A1A9YC03 E0VK92 W8C9H5 U4UGN6 N6TFJ6 E2ALW7 A0A088AVR5 A0A2A3ERL5 A0A195BWL8 A0A158NCJ3 A0A182MSY0 E9IM88 A0A195CQE1 A0A158NCJ2 A0A336MRS6 A0A0M8ZRE3 A0A026WVU1 A0A3L8DI55 E2BHY3 A0A1A9ZUY7 A0A1B0FC75 A0A1B0GJF5 A0A1A9W101 J9M312 A0A1A9UU25 A0A195F967 A0A151X5T7 A0A0L7RJA9 A0A3R7PEH3 F4WX53 A0A067QQI2 A0A1D1VF32 A0A2P8ZK26 T1IJC5 A0A2J7QH10 A0A1Y1KDP2 A0A0T6BC19 A0A1W4XJ75 A0A1Y3BWD0 A0A1W0WSW7 A0A0P5BGP2 A0A0P6GS82 A0A0P6I7Q2
Pubmed
28756777
19121390
26354079
22118469
26227816
18362917
+ More
19820115 17510324 17994087 22936249 17550304 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 24438588 15632085 23185243 25244985 20966253 12364791 14747013 17210077 25830018 25315136 26108605 25348373 24495485 24646461 20566863 23537049 20798317 21347285 21282665 24508170 30249741 21719571 24845553 27649274 29403074 28004739
19820115 17510324 17994087 22936249 17550304 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 24438588 15632085 23185243 25244985 20966253 12364791 14747013 17210077 25830018 25315136 26108605 25348373 24495485 24646461 20566863 23537049 20798317 21347285 21282665 24508170 30249741 21719571 24845553 27649274 29403074 28004739
EMBL
NWSH01000887
PCG73682.1
KZ149900
PZC78592.1
KP657629
AKL78854.1
+ More
ODYU01008299 SOQ51765.1 BABH01030961 BABH01030962 KQ460398 KPJ15019.1 KQ459302 KPJ01814.1 AGBW02010077 OWR49204.1 JTDY01000364 KOB77527.1 RSAL01000493 RVE41539.1 KQ971372 EFA10348.1 DS232080 EDS34186.1 CH477272 EAT45251.1 CH964282 EDW85561.1 CH916367 EDW01993.1 CH933808 EDW08946.2 CP012524 ALC42171.1 CH940648 EDW61660.1 CM002911 KMY91772.1 CM000157 EDW89072.1 AE013599 BT083414 AAM70812.1 ACQ89823.1 CH954177 EDV59802.1 ATLV01015292 KE525006 KFB40295.1 OUUW01000001 SPP74181.1 CH480816 EDW46643.1 CH479181 EDW31392.1 CH902619 EDV38125.1 CM000071 EAL24715.2 KRT01490.1 KRT01491.1 KRT01492.1 AXCN02000551 KRJ98161.1 UFQT01001786 SSX31809.1 AAAB01008807 EAA03914.4 GBXI01011171 JAD03121.1 JRES01000975 KNC26593.1 AB897807 BAO52000.1 GAKP01023480 JAC35478.1 ACPB03016005 GAMC01006647 JAB99908.1 KJ152556 AHN85839.1 CVRI01000059 CRL03346.1 APCN01000188 DS235239 EEB13798.1 GAMC01006646 JAB99909.1 KB632136 ERL89055.1 APGK01029452 APGK01029453 APGK01029454 APGK01029455 APGK01029456 APGK01029457 KB740735 ENN79134.1 GL440682 EFN65559.1 KZ288198 PBC33681.1 KQ976396 KYM92977.1 ADTU01011942 AXCM01001173 GL764129 EFZ18462.1 KQ977408 KYN02931.1 UFQT01002012 SSX32405.1 KQ435922 KOX68436.1 KK107088 EZA59791.1 QOIP01000008 RLU19843.1 GL448390 EFN84699.1 CCAG010011030 AJWK01020251 ABLF02041125 ABLF02041147 KQ981727 KYN36983.1 KQ982490 KYQ55775.1 KQ414582 KOC70868.1 QCYY01003481 ROT63152.1 GL888417 EGI61296.1 KK853042 KDR12158.1 BDGG01000004 GAU97503.1 PYGN01000033 PSN56849.1 JH430253 NEVH01014358 PNF27872.1 GEZM01089377 JAV57516.1 LJIG01002050 KRT84890.1 MUJZ01000708 OTF84103.1 MTYJ01000051 OQV18280.1 GDIP01190175 JAJ33227.1 GDIQ01029522 JAN65215.1 GDIQ01010107 JAN84630.1
ODYU01008299 SOQ51765.1 BABH01030961 BABH01030962 KQ460398 KPJ15019.1 KQ459302 KPJ01814.1 AGBW02010077 OWR49204.1 JTDY01000364 KOB77527.1 RSAL01000493 RVE41539.1 KQ971372 EFA10348.1 DS232080 EDS34186.1 CH477272 EAT45251.1 CH964282 EDW85561.1 CH916367 EDW01993.1 CH933808 EDW08946.2 CP012524 ALC42171.1 CH940648 EDW61660.1 CM002911 KMY91772.1 CM000157 EDW89072.1 AE013599 BT083414 AAM70812.1 ACQ89823.1 CH954177 EDV59802.1 ATLV01015292 KE525006 KFB40295.1 OUUW01000001 SPP74181.1 CH480816 EDW46643.1 CH479181 EDW31392.1 CH902619 EDV38125.1 CM000071 EAL24715.2 KRT01490.1 KRT01491.1 KRT01492.1 AXCN02000551 KRJ98161.1 UFQT01001786 SSX31809.1 AAAB01008807 EAA03914.4 GBXI01011171 JAD03121.1 JRES01000975 KNC26593.1 AB897807 BAO52000.1 GAKP01023480 JAC35478.1 ACPB03016005 GAMC01006647 JAB99908.1 KJ152556 AHN85839.1 CVRI01000059 CRL03346.1 APCN01000188 DS235239 EEB13798.1 GAMC01006646 JAB99909.1 KB632136 ERL89055.1 APGK01029452 APGK01029453 APGK01029454 APGK01029455 APGK01029456 APGK01029457 KB740735 ENN79134.1 GL440682 EFN65559.1 KZ288198 PBC33681.1 KQ976396 KYM92977.1 ADTU01011942 AXCM01001173 GL764129 EFZ18462.1 KQ977408 KYN02931.1 UFQT01002012 SSX32405.1 KQ435922 KOX68436.1 KK107088 EZA59791.1 QOIP01000008 RLU19843.1 GL448390 EFN84699.1 CCAG010011030 AJWK01020251 ABLF02041125 ABLF02041147 KQ981727 KYN36983.1 KQ982490 KYQ55775.1 KQ414582 KOC70868.1 QCYY01003481 ROT63152.1 GL888417 EGI61296.1 KK853042 KDR12158.1 BDGG01000004 GAU97503.1 PYGN01000033 PSN56849.1 JH430253 NEVH01014358 PNF27872.1 GEZM01089377 JAV57516.1 LJIG01002050 KRT84890.1 MUJZ01000708 OTF84103.1 MTYJ01000051 OQV18280.1 GDIP01190175 JAJ33227.1 GDIQ01029522 JAN65215.1 GDIQ01010107 JAN84630.1
Proteomes
UP000218220
UP000005204
UP000053240
UP000053268
UP000007151
UP000037510
+ More
UP000283053 UP000007266 UP000002320 UP000008820 UP000007798 UP000001070 UP000009192 UP000092553 UP000008792 UP000192221 UP000002282 UP000000803 UP000008711 UP000030765 UP000268350 UP000001292 UP000008744 UP000007801 UP000001819 UP000075886 UP000075884 UP000075880 UP000076408 UP000075900 UP000075903 UP000076407 UP000075882 UP000007062 UP000075920 UP000075902 UP000075885 UP000095301 UP000037069 UP000069272 UP000095300 UP000015103 UP000183832 UP000075840 UP000092443 UP000009046 UP000030742 UP000019118 UP000000311 UP000005203 UP000242457 UP000078540 UP000005205 UP000075883 UP000078542 UP000053105 UP000053097 UP000279307 UP000008237 UP000092445 UP000092444 UP000092461 UP000091820 UP000007819 UP000078200 UP000078541 UP000075809 UP000053825 UP000283509 UP000007755 UP000027135 UP000186922 UP000245037 UP000235965 UP000192223
UP000283053 UP000007266 UP000002320 UP000008820 UP000007798 UP000001070 UP000009192 UP000092553 UP000008792 UP000192221 UP000002282 UP000000803 UP000008711 UP000030765 UP000268350 UP000001292 UP000008744 UP000007801 UP000001819 UP000075886 UP000075884 UP000075880 UP000076408 UP000075900 UP000075903 UP000076407 UP000075882 UP000007062 UP000075920 UP000075902 UP000075885 UP000095301 UP000037069 UP000069272 UP000095300 UP000015103 UP000183832 UP000075840 UP000092443 UP000009046 UP000030742 UP000019118 UP000000311 UP000005203 UP000242457 UP000078540 UP000005205 UP000075883 UP000078542 UP000053105 UP000053097 UP000279307 UP000008237 UP000092445 UP000092444 UP000092461 UP000091820 UP000007819 UP000078200 UP000078541 UP000075809 UP000053825 UP000283509 UP000007755 UP000027135 UP000186922 UP000245037 UP000235965 UP000192223
Interpro
Gene 3D
ProteinModelPortal
A0A2A4JQI7
A0A2W1BU38
A0A0G3VFZ2
A0A2H1WFA6
H9JBA9
A0A194RC18
+ More
A0A194Q8T4 A0A212F653 A0A0L7LQ13 A0A3S2P4I8 D6X389 B0WT43 Q17F96 B4NMH2 A0A1S4G848 B4J6J2 B4KNI7 A0A0M4EAN4 B4LK06 A0A1W4VF47 A0A0J9R6G5 B4P0Z8 A1Z6N4 B3NB37 A0A084VQQ1 A0A3B0JNG0 B4HQA4 B4GC74 B3MIP9 Q292Z5 A0A0R3NKV6 A0A182PZH8 A0A182NKM3 A0A0R1DSU5 A0A182IWE7 A0A336MRT6 A0A182XYJ0 A0A182RU01 A0A182VJL2 A0A182WZ30 A0A182KQV6 Q7PTH4 A0A182WJK6 A0A182UH83 A0A182P203 A0A0A1WWU3 A0A1I8N2R7 A0A0L0C2L5 A0A182FG49 A0A1I8P932 W8VJW9 A0A034V1P5 T1I5A1 W8BSM1 X2KSD2 A0A1J1IUG8 A0A182I2X6 A0A1A9YC03 E0VK92 W8C9H5 U4UGN6 N6TFJ6 E2ALW7 A0A088AVR5 A0A2A3ERL5 A0A195BWL8 A0A158NCJ3 A0A182MSY0 E9IM88 A0A195CQE1 A0A158NCJ2 A0A336MRS6 A0A0M8ZRE3 A0A026WVU1 A0A3L8DI55 E2BHY3 A0A1A9ZUY7 A0A1B0FC75 A0A1B0GJF5 A0A1A9W101 J9M312 A0A1A9UU25 A0A195F967 A0A151X5T7 A0A0L7RJA9 A0A3R7PEH3 F4WX53 A0A067QQI2 A0A1D1VF32 A0A2P8ZK26 T1IJC5 A0A2J7QH10 A0A1Y1KDP2 A0A0T6BC19 A0A1W4XJ75 A0A1Y3BWD0 A0A1W0WSW7 A0A0P5BGP2 A0A0P6GS82 A0A0P6I7Q2
A0A194Q8T4 A0A212F653 A0A0L7LQ13 A0A3S2P4I8 D6X389 B0WT43 Q17F96 B4NMH2 A0A1S4G848 B4J6J2 B4KNI7 A0A0M4EAN4 B4LK06 A0A1W4VF47 A0A0J9R6G5 B4P0Z8 A1Z6N4 B3NB37 A0A084VQQ1 A0A3B0JNG0 B4HQA4 B4GC74 B3MIP9 Q292Z5 A0A0R3NKV6 A0A182PZH8 A0A182NKM3 A0A0R1DSU5 A0A182IWE7 A0A336MRT6 A0A182XYJ0 A0A182RU01 A0A182VJL2 A0A182WZ30 A0A182KQV6 Q7PTH4 A0A182WJK6 A0A182UH83 A0A182P203 A0A0A1WWU3 A0A1I8N2R7 A0A0L0C2L5 A0A182FG49 A0A1I8P932 W8VJW9 A0A034V1P5 T1I5A1 W8BSM1 X2KSD2 A0A1J1IUG8 A0A182I2X6 A0A1A9YC03 E0VK92 W8C9H5 U4UGN6 N6TFJ6 E2ALW7 A0A088AVR5 A0A2A3ERL5 A0A195BWL8 A0A158NCJ3 A0A182MSY0 E9IM88 A0A195CQE1 A0A158NCJ2 A0A336MRS6 A0A0M8ZRE3 A0A026WVU1 A0A3L8DI55 E2BHY3 A0A1A9ZUY7 A0A1B0FC75 A0A1B0GJF5 A0A1A9W101 J9M312 A0A1A9UU25 A0A195F967 A0A151X5T7 A0A0L7RJA9 A0A3R7PEH3 F4WX53 A0A067QQI2 A0A1D1VF32 A0A2P8ZK26 T1IJC5 A0A2J7QH10 A0A1Y1KDP2 A0A0T6BC19 A0A1W4XJ75 A0A1Y3BWD0 A0A1W0WSW7 A0A0P5BGP2 A0A0P6GS82 A0A0P6I7Q2
PDB
3RCH
E-value=2.28233e-153,
Score=1393
Ontologies
KEGG
PATHWAY
GO
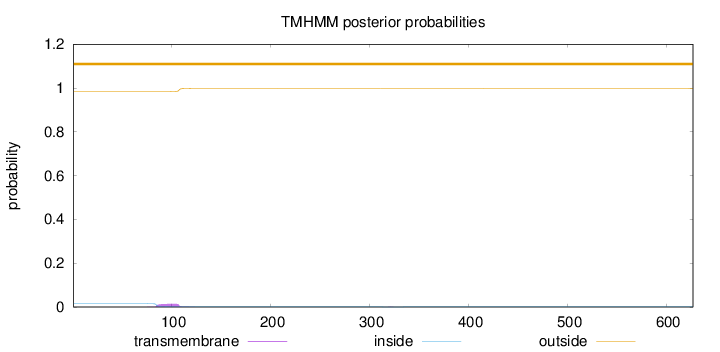
Topology
Length:
627
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.341990000000001
Exp number, first 60 AAs:
0
Total prob of N-in:
0.01523
outside
1 - 627
Population Genetic Test Statistics
Pi
297.010825
Theta
195.034378
Tajima's D
1.712947
CLR
0.165922
CSRT
0.831158442077896
Interpretation
Uncertain
