Gene
KWMTBOMO05151
Annotation
PREDICTED:_uncharacterized_protein_LOC101747137_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 2.775
Sequence
CDS
ATGGCAAGGTGCTTCTCGTGCTGCAACACTGAGGGGGAGCTGCACACGTTCGACGACGCCAGCGAGAGGGCCTACGCGTGTGCAGCCTATTGGCGTCAGAAGACAGACGACAGCACCTACCACGTCACACTACTAGCAGGGAAAGCCAGGGTGACCCCACTGAGACCAGTCTCTATCCCAAGGCTGGAGCTGCAAGCTGCACTGCTGGGGACAAGGATGGCGCAGGCGATAGCGAACGAATTGGACATCGCGGTCGGCAGAAGGACGTACTGGACTGACTCCAGTACAGTTCTGACATGGATAAAGACCGACCCACGCACGTTCAAACCTTTCGTTGCGCACCGACTCGCCGAGATAGAAGAGTCAACGAAGCCCCAAGAATGGCGATGGGTGCCCGGCTCGCAAAACCCAGCAGACGACGCAACTAGGGAGGCACCGGCGGATTTCGACCACACGCATCGGTGGTTTAATGGACCCGAGTTCCTGTCGTGGGATGAATCGCGCTGGCCCAAGCCGCGAACCTTCAAGCAAGAACCGTCTGGAGAAGAGAAAGAGGCCTTCCTGGTCGCCACGGCGAGGACCGCTGACGCTCGACCCACGCCCGATCCGCACAGATTCTCGAGCTGGGTCAGGTTATTGAGAGCAACAGCCAGGGTCCTCCAATTTATCGAGCTGTGTCGACCCCGGAAGGAGAGCGCTTGCGTGTCCAGACAATCGAAGCAGCAAGATCCCACGTGGAGGACGACACGAACGAAGCAACCCCGATCTACATGGAAGTTAAGGACCCCGGAAGCACCTACAGAAGCATGGCTGCCGCTTGATCCGCCCCACTTGAAGAAGGCGGAGAAGATCCTACTGAGAAGCAGCCAAGGAGAGAGCTTCGGTGAAAAAGATCCCGAACGTCACCCCAAGTTGCGACGGCTCGACGTCGTGATGGAAGATGGCCTGTTGCGCCTGCGCGGACGTATCGACGCAGCGCAGTACATAGATGCAGGCTGCAAGCGACCGATCGTGCTGGACGGGAAGCACGTGATAGCGAGATTATTGATCAAACACTATCACGAGGCGTTCCAGCACGGTAACCACGCAACGGTGATGAACGAGGTGCGGCAGCGCTACTGGATCCTAGGTCTAAGATCGATCATTCGAGCGACGGCCGTCCGATGCCAGTGGTGCAAGGTCTATCGGAGTACACCACGACTACCGCCCACCGGAGACCTGCCGATAGAGCGGCTACGACATGGAGAGCCGCCGTTCACCTGTGCTGCCGTCGATTACTTCGGGCCGATGACGGTGACTGTAGGACGACGTCACGAGAAGAGAACGAGCGGCGGTCTGCTTAAGAGACCTGCCTCCAAGATGATCCTGCTGGTGCCTGCAACGAGCGACGAGCCCACGCCCATAGAGTCGCCACGACAAGGAGTCGGTGCTACGCACGAGGGGGAGGATGTTGGCGACAGCGGCGTCGCCCCATAA
Protein
MARCFSCCNTEGELHTFDDASERAYACAAYWRQKTDDSTYHVTLLAGKARVTPLRPVSIPRLELQAALLGTRMAQAIANELDIAVGRRTYWTDSSTVLTWIKTDPRTFKPFVAHRLAEIEESTKPQEWRWVPGSQNPADDATREAPADFDHTHRWFNGPEFLSWDESRWPKPRTFKQEPSGEEKEAFLVATARTADARPTPDPHRFSSWVRLLRATARVLQFIELCRPRKESACVSRQSKQQDPTWRTTRTKQPRSTWKLRTPEAPTEAWLPLDPPHLKKAEKILLRSSQGESFGEKDPERHPKLRRLDVVMEDGLLRLRGRIDAAQYIDAGCKRPIVLDGKHVIARLLIKHYHEAFQHGNHATVMNEVRQRYWILGLRSIIRATAVRCQWCKVYRSTPRLPPTGDLPIERLRHGEPPFTCAAVDYFGPMTVTVGRRHEKRTSGGLLKRPASKMILLVPATSDEPTPIESPRQGVGATHEGEDVGDSGVAP
Summary
Uniprot
Q2MGA5
Q9NDM9
Q9NDN0
A0A0N1IGH3
K7JA41
A0A3S2N5J7
+ More
A0A2A4IRQ3 A0A226EKF8 A0A226EFR1 A0A226D351 A0A226DGY5 A0A226DM06 A0A226DH22 A0A182GL35 A0A1W7R6H4 W8AYL5 A0A226E2P7 W8B882 A0A182HCB4 W8AJC8 A0A182G158 A0A182GRX9 A0A226DTU1 A0A1W7R681 A0A2M4CKZ0 A0A2M4CKI9 A0A1W7R6F2 A0A182YRW4 A0A182H5B8 A0A0L7LE26 Q76C95 Q76C94 A0A2M4CVY8 A0A146SP51 A0A146S4B8 A0A0N5E779 A0A182G7Z0 A0A226EEI4 W4YU11 A0A1S3SW27 A0A1B6LK77 A0A0P6C5E3 A0A0P6AZF3 W4XIU4 A0A1W7R6M1 W4XIU5 A0A034V9M7 A0A146S1B6 W4XIU3 A0A146Q7X0 W4XIU6 A0A2G8KVJ2 A0A2M4AHJ4 A0A182G4R3 A0A146PP93 A0A146QCZ7 W4YQI7 A0A085NF60 W4Y9V5 W4ZKP7 A0A0P6IXJ2 A0A2B4SX75 A0A0S7LT68 A0A146T8G6 A0A1A8V6P1 A0A0N8BUP4 A0A2I4AL07 A0A0N5E596 A0A0P6IQY0 A0A146QI09 A0A146QJ75 A0A182G5I1 A0A1S3HN42 A0A146TD72 W4ZDM7 W4Z813 A0A182H4A2 W4XFV2 A0A0P5QIW7 A0A182HEF9 A0A0S7F0U2 W4XBE0 W4XFV4 W4Z812 A0A146T9I5 A0A0S7G7U6 A0A0P4VNG8 A0A0P4VX58 A0A0N0PBK0 A0A226EDU9 A0A0N5DTJ9 A0A146S218 W4Z815 A0A146RUC7 A0A0S7FZ14 A0A3R7QG24 A0A0P6DVP0 W4XFV1 W4XFV3 A0A0P6DZM7
A0A2A4IRQ3 A0A226EKF8 A0A226EFR1 A0A226D351 A0A226DGY5 A0A226DM06 A0A226DH22 A0A182GL35 A0A1W7R6H4 W8AYL5 A0A226E2P7 W8B882 A0A182HCB4 W8AJC8 A0A182G158 A0A182GRX9 A0A226DTU1 A0A1W7R681 A0A2M4CKZ0 A0A2M4CKI9 A0A1W7R6F2 A0A182YRW4 A0A182H5B8 A0A0L7LE26 Q76C95 Q76C94 A0A2M4CVY8 A0A146SP51 A0A146S4B8 A0A0N5E779 A0A182G7Z0 A0A226EEI4 W4YU11 A0A1S3SW27 A0A1B6LK77 A0A0P6C5E3 A0A0P6AZF3 W4XIU4 A0A1W7R6M1 W4XIU5 A0A034V9M7 A0A146S1B6 W4XIU3 A0A146Q7X0 W4XIU6 A0A2G8KVJ2 A0A2M4AHJ4 A0A182G4R3 A0A146PP93 A0A146QCZ7 W4YQI7 A0A085NF60 W4Y9V5 W4ZKP7 A0A0P6IXJ2 A0A2B4SX75 A0A0S7LT68 A0A146T8G6 A0A1A8V6P1 A0A0N8BUP4 A0A2I4AL07 A0A0N5E596 A0A0P6IQY0 A0A146QI09 A0A146QJ75 A0A182G5I1 A0A1S3HN42 A0A146TD72 W4ZDM7 W4Z813 A0A182H4A2 W4XFV2 A0A0P5QIW7 A0A182HEF9 A0A0S7F0U2 W4XBE0 W4XFV4 W4Z812 A0A146T9I5 A0A0S7G7U6 A0A0P4VNG8 A0A0P4VX58 A0A0N0PBK0 A0A226EDU9 A0A0N5DTJ9 A0A146S218 W4Z815 A0A146RUC7 A0A0S7FZ14 A0A3R7QG24 A0A0P6DVP0 W4XFV1 W4XFV3 A0A0P6DZM7
Pubmed
EMBL
AF530470
AAQ09229.1
AB042119
BAA95570.1
AB042118
BAA95569.1
+ More
KQ459986 KPJ18861.1 AAZX01023278 RSAL01000375 RVE42106.1 NWSH01008670 PCG62455.1 LNIX01000003 OXA57597.1 LNIX01000004 OXA56382.1 LNIX01000039 OXA39294.1 LNIX01000019 OXA44812.1 LNIX01000015 OXA46575.1 OXA44855.1 JXUM01013493 KQ560378 KXJ82704.1 GEHC01000914 JAV46731.1 GAMC01020594 JAB85961.1 LNIX01000007 OXA52015.1 GAMC01020598 GAMC01020596 GAMC01020592 JAB85959.1 JXUM01127084 KQ567147 KXJ69602.1 GAMC01017930 JAB88625.1 JXUM01136032 KQ568367 KXJ69026.1 JXUM01083542 KQ563383 KXJ73941.1 LNIX01000012 OXA48117.1 GEHC01000967 JAV46678.1 GGFL01001842 MBW66020.1 GGFL01001682 MBW65860.1 GEHC01000913 JAV46732.1 JXUM01111221 KQ565482 KXJ70922.1 JTDY01001530 KOB73634.1 AB110069 BAD01589.1 BAD01590.1 GGFL01005193 MBW69371.1 GCES01103975 JAQ82347.1 GCES01110894 JAQ75428.1 JXUM01151198 KQ571164 KXJ68235.1 OXA55241.1 AAGJ04142769 AAGJ04142770 GEBQ01015876 JAT24101.1 GDIP01004568 JAM99147.1 GDIP01036233 JAM67482.1 AAGJ04091218 GEHC01000872 JAV46773.1 AAGJ04001285 GAKP01020709 JAC38243.1 GCES01111900 JAQ74422.1 GCES01134385 JAQ51937.1 MRZV01000348 PIK51982.1 GGFK01006929 MBW40250.1 JXUM01144298 KQ569759 KXJ68523.1 GCES01140631 JAQ45691.1 GCES01132273 JAQ54049.1 AAGJ04146728 AAGJ04146729 KL367508 KFD68106.1 AAGJ04125735 AAGJ04095911 GDIQ01002920 JAN91817.1 LSMT01000016 PFX33015.1 GBYX01150372 JAO92643.1 GCES01097275 JAQ89047.1 HAEJ01014775 SBS55232.1 GDIQ01143107 JAL08619.1 GDIQ01002921 JAN91816.1 GCES01130483 JAQ55839.1 GCES01130050 JAQ56272.1 JXUM01146045 JXUM01146046 GCES01095441 JAQ90881.1 AAGJ04011066 AAGJ04020014 JXUM01109099 KQ565290 KXJ71123.1 AAGJ04008210 GDIQ01114015 JAL37711.1 JXUM01005879 KQ560203 KXJ83865.1 GBYX01472248 GBYX01472237 GBYX01472235 GBYX01472234 JAO09406.1 JAO09417.1 JAO09419.1 JAO09420.1 GCES01096813 JAQ89509.1 GBYX01459357 JAO22202.1 GDRN01111557 JAI56761.1 GDRN01111556 JAI56762.1 KQ460970 KPJ10166.1 OXA55408.1 GCES01112016 JAQ74306.1 GCES01114344 JAQ71978.1 GBYX01459355 JAO22204.1 QCYY01001488 ROT77729.1 GDIQ01071585 JAN23152.1 GDIQ01071584 JAN23153.1
KQ459986 KPJ18861.1 AAZX01023278 RSAL01000375 RVE42106.1 NWSH01008670 PCG62455.1 LNIX01000003 OXA57597.1 LNIX01000004 OXA56382.1 LNIX01000039 OXA39294.1 LNIX01000019 OXA44812.1 LNIX01000015 OXA46575.1 OXA44855.1 JXUM01013493 KQ560378 KXJ82704.1 GEHC01000914 JAV46731.1 GAMC01020594 JAB85961.1 LNIX01000007 OXA52015.1 GAMC01020598 GAMC01020596 GAMC01020592 JAB85959.1 JXUM01127084 KQ567147 KXJ69602.1 GAMC01017930 JAB88625.1 JXUM01136032 KQ568367 KXJ69026.1 JXUM01083542 KQ563383 KXJ73941.1 LNIX01000012 OXA48117.1 GEHC01000967 JAV46678.1 GGFL01001842 MBW66020.1 GGFL01001682 MBW65860.1 GEHC01000913 JAV46732.1 JXUM01111221 KQ565482 KXJ70922.1 JTDY01001530 KOB73634.1 AB110069 BAD01589.1 BAD01590.1 GGFL01005193 MBW69371.1 GCES01103975 JAQ82347.1 GCES01110894 JAQ75428.1 JXUM01151198 KQ571164 KXJ68235.1 OXA55241.1 AAGJ04142769 AAGJ04142770 GEBQ01015876 JAT24101.1 GDIP01004568 JAM99147.1 GDIP01036233 JAM67482.1 AAGJ04091218 GEHC01000872 JAV46773.1 AAGJ04001285 GAKP01020709 JAC38243.1 GCES01111900 JAQ74422.1 GCES01134385 JAQ51937.1 MRZV01000348 PIK51982.1 GGFK01006929 MBW40250.1 JXUM01144298 KQ569759 KXJ68523.1 GCES01140631 JAQ45691.1 GCES01132273 JAQ54049.1 AAGJ04146728 AAGJ04146729 KL367508 KFD68106.1 AAGJ04125735 AAGJ04095911 GDIQ01002920 JAN91817.1 LSMT01000016 PFX33015.1 GBYX01150372 JAO92643.1 GCES01097275 JAQ89047.1 HAEJ01014775 SBS55232.1 GDIQ01143107 JAL08619.1 GDIQ01002921 JAN91816.1 GCES01130483 JAQ55839.1 GCES01130050 JAQ56272.1 JXUM01146045 JXUM01146046 GCES01095441 JAQ90881.1 AAGJ04011066 AAGJ04020014 JXUM01109099 KQ565290 KXJ71123.1 AAGJ04008210 GDIQ01114015 JAL37711.1 JXUM01005879 KQ560203 KXJ83865.1 GBYX01472248 GBYX01472237 GBYX01472235 GBYX01472234 JAO09406.1 JAO09417.1 JAO09419.1 JAO09420.1 GCES01096813 JAQ89509.1 GBYX01459357 JAO22202.1 GDRN01111557 JAI56761.1 GDRN01111556 JAI56762.1 KQ460970 KPJ10166.1 OXA55408.1 GCES01112016 JAQ74306.1 GCES01114344 JAQ71978.1 GBYX01459355 JAO22204.1 QCYY01001488 ROT77729.1 GDIQ01071585 JAN23152.1 GDIQ01071584 JAN23153.1
Proteomes
Pfam
Interpro
IPR040676
DUF5641
+ More
IPR036397 RNaseH_sf
IPR005312 DUF1759
IPR001584 Integrase_cat-core
IPR008042 Retrotrans_Pao
IPR012337 RNaseH-like_sf
IPR041588 Integrase_H2C2
IPR008737 Peptidase_asp_put
IPR001995 Peptidase_A2_cat
IPR001878 Znf_CCHC
IPR006094 Oxid_FAD_bind_N
IPR016166 FAD-bd_PCMH
IPR016164 FAD-linked_Oxase-like_C
IPR016170 Cytok_DH_C_sf
IPR015213 Cholesterol_OX_subst-bd
IPR036318 FAD-bd_PCMH-like_sf
IPR016169 FAD-bd_PCMH_sub2
IPR016167 FAD-bd_PCMH_sub1
IPR011011 Znf_FYVE_PHD
IPR013083 Znf_RING/FYVE/PHD
IPR001965 Znf_PHD
IPR019787 Znf_PHD-finger
IPR019786 Zinc_finger_PHD-type_CS
IPR000477 RT_dom
IPR001841 Znf_RING
IPR000253 FHA_dom
IPR021109 Peptidase_aspartic_dom_sf
IPR001969 Aspartic_peptidase_AS
IPR036875 Znf_CCHC_sf
IPR036397 RNaseH_sf
IPR005312 DUF1759
IPR001584 Integrase_cat-core
IPR008042 Retrotrans_Pao
IPR012337 RNaseH-like_sf
IPR041588 Integrase_H2C2
IPR008737 Peptidase_asp_put
IPR001995 Peptidase_A2_cat
IPR001878 Znf_CCHC
IPR006094 Oxid_FAD_bind_N
IPR016166 FAD-bd_PCMH
IPR016164 FAD-linked_Oxase-like_C
IPR016170 Cytok_DH_C_sf
IPR015213 Cholesterol_OX_subst-bd
IPR036318 FAD-bd_PCMH-like_sf
IPR016169 FAD-bd_PCMH_sub2
IPR016167 FAD-bd_PCMH_sub1
IPR011011 Znf_FYVE_PHD
IPR013083 Znf_RING/FYVE/PHD
IPR001965 Znf_PHD
IPR019787 Znf_PHD-finger
IPR019786 Zinc_finger_PHD-type_CS
IPR000477 RT_dom
IPR001841 Znf_RING
IPR000253 FHA_dom
IPR021109 Peptidase_aspartic_dom_sf
IPR001969 Aspartic_peptidase_AS
IPR036875 Znf_CCHC_sf
SUPFAM
ProteinModelPortal
Q2MGA5
Q9NDM9
Q9NDN0
A0A0N1IGH3
K7JA41
A0A3S2N5J7
+ More
A0A2A4IRQ3 A0A226EKF8 A0A226EFR1 A0A226D351 A0A226DGY5 A0A226DM06 A0A226DH22 A0A182GL35 A0A1W7R6H4 W8AYL5 A0A226E2P7 W8B882 A0A182HCB4 W8AJC8 A0A182G158 A0A182GRX9 A0A226DTU1 A0A1W7R681 A0A2M4CKZ0 A0A2M4CKI9 A0A1W7R6F2 A0A182YRW4 A0A182H5B8 A0A0L7LE26 Q76C95 Q76C94 A0A2M4CVY8 A0A146SP51 A0A146S4B8 A0A0N5E779 A0A182G7Z0 A0A226EEI4 W4YU11 A0A1S3SW27 A0A1B6LK77 A0A0P6C5E3 A0A0P6AZF3 W4XIU4 A0A1W7R6M1 W4XIU5 A0A034V9M7 A0A146S1B6 W4XIU3 A0A146Q7X0 W4XIU6 A0A2G8KVJ2 A0A2M4AHJ4 A0A182G4R3 A0A146PP93 A0A146QCZ7 W4YQI7 A0A085NF60 W4Y9V5 W4ZKP7 A0A0P6IXJ2 A0A2B4SX75 A0A0S7LT68 A0A146T8G6 A0A1A8V6P1 A0A0N8BUP4 A0A2I4AL07 A0A0N5E596 A0A0P6IQY0 A0A146QI09 A0A146QJ75 A0A182G5I1 A0A1S3HN42 A0A146TD72 W4ZDM7 W4Z813 A0A182H4A2 W4XFV2 A0A0P5QIW7 A0A182HEF9 A0A0S7F0U2 W4XBE0 W4XFV4 W4Z812 A0A146T9I5 A0A0S7G7U6 A0A0P4VNG8 A0A0P4VX58 A0A0N0PBK0 A0A226EDU9 A0A0N5DTJ9 A0A146S218 W4Z815 A0A146RUC7 A0A0S7FZ14 A0A3R7QG24 A0A0P6DVP0 W4XFV1 W4XFV3 A0A0P6DZM7
A0A2A4IRQ3 A0A226EKF8 A0A226EFR1 A0A226D351 A0A226DGY5 A0A226DM06 A0A226DH22 A0A182GL35 A0A1W7R6H4 W8AYL5 A0A226E2P7 W8B882 A0A182HCB4 W8AJC8 A0A182G158 A0A182GRX9 A0A226DTU1 A0A1W7R681 A0A2M4CKZ0 A0A2M4CKI9 A0A1W7R6F2 A0A182YRW4 A0A182H5B8 A0A0L7LE26 Q76C95 Q76C94 A0A2M4CVY8 A0A146SP51 A0A146S4B8 A0A0N5E779 A0A182G7Z0 A0A226EEI4 W4YU11 A0A1S3SW27 A0A1B6LK77 A0A0P6C5E3 A0A0P6AZF3 W4XIU4 A0A1W7R6M1 W4XIU5 A0A034V9M7 A0A146S1B6 W4XIU3 A0A146Q7X0 W4XIU6 A0A2G8KVJ2 A0A2M4AHJ4 A0A182G4R3 A0A146PP93 A0A146QCZ7 W4YQI7 A0A085NF60 W4Y9V5 W4ZKP7 A0A0P6IXJ2 A0A2B4SX75 A0A0S7LT68 A0A146T8G6 A0A1A8V6P1 A0A0N8BUP4 A0A2I4AL07 A0A0N5E596 A0A0P6IQY0 A0A146QI09 A0A146QJ75 A0A182G5I1 A0A1S3HN42 A0A146TD72 W4ZDM7 W4Z813 A0A182H4A2 W4XFV2 A0A0P5QIW7 A0A182HEF9 A0A0S7F0U2 W4XBE0 W4XFV4 W4Z812 A0A146T9I5 A0A0S7G7U6 A0A0P4VNG8 A0A0P4VX58 A0A0N0PBK0 A0A226EDU9 A0A0N5DTJ9 A0A146S218 W4Z815 A0A146RUC7 A0A0S7FZ14 A0A3R7QG24 A0A0P6DVP0 W4XFV1 W4XFV3 A0A0P6DZM7
Ontologies
GO
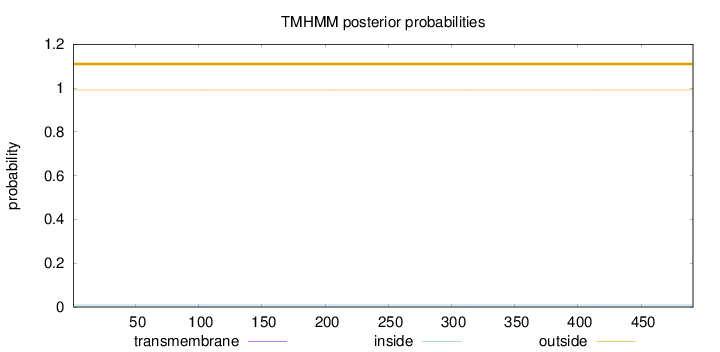
Topology
Length:
491
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00347000000000001
Exp number, first 60 AAs:
0.00124
Total prob of N-in:
0.00905
outside
1 - 491
Population Genetic Test Statistics
Pi
425.699012
Theta
220.27077
Tajima's D
2.667333
CLR
80.776148
CSRT
0.958302084895755
Interpretation
Uncertain
