Gene
KWMTBOMO05121
Pre Gene Modal
BGIBMGA001459
Annotation
reverse_transcriptase_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 2.465
Sequence
CDS
ATGGACGCTGTATTCGCGGAATTCCTCCGACTTCGCCACCCACAGCTCGCCTCGGAGTTTTTGGCCTTCAAGGCCAATCACACTGCGAGCCCACTCGAGGACTCCGCCGCGCTCGCTGCTCCTGCGTCGCCTGTACCTGCGTGCAGAGCTTCTGCATTGAGCACCGCAGCCTCTGTCGTGCCTGCCGCTCCCGTGTCGCCTATACTGGCGAGAAAAGCTGCTGCGTCGTCCGCCGTGACCATCGTTTCAGCTGAGCGATCATCCGCGGCCTCCGTCGCGCCCTCTAAAACACCTACACTTGCTCGTAGGTCGCCTGCACCCGCCTCCTCGTGCTCCGACTCTGACTCGGACATGGAGGTCGACCTCGCCCCCGCCTCATCGACGGATGGATTCACCCTGGTACAGAAGGGTAAGAAGCGTGCCGCGGAGTCTCGAGCTCCCGCGGCCGCTAAAATTAGCAAAGCCGTGAACGCGTCGCGCCCCCGCCCTCAGACTCCCGTTGCGCCCCCAGCCCGTGCCACTCCGTCGCCGCGTCCGGTGGCACAAAATAAAACCCAGACCCCTCCCCCGGTTATCCTTCAGGAGAAGGCAGCTTGGGATCGAGTTTGCCTGGCCCTTAAGGCCAAAAATATAAATTTCACGAATGCCCGTAACCTCGCGAACGGCATTCAAATTAAGGTTCAAACACCCGACGACCATAGGGCCCTCTCTTCTTACCTCCGTAAGGAGCGTATAAGTTTCCATACGTATACGCTCCAGGAGGAGCGCGAACTCCGCGTTGTAATACGCGGAATCCCTAAAGAGTTAGATGTAGAGCTCGTAAAAGCCGACCTGTTAGAACAAGGCCTACCAGTGAATTCTGTGCACCGTATGCACACCGGTCGCGGTAGGGAGCCATATAATATGGTTCTAGTCGCTCTCCAGCCTACCCCCGAGGTGCAGGAGACCCTACTTAAGCCCGCGCGCCGTGACCCTAAAATCGCGAACTATAACATGGTCAGGAACGACAGGCTCTCTGCCCGTGGTGGTGGTACCGTCATTTACTATAGAAGAGCCCTGCATTGCGTCCCGCTCGATCCTCCCGCGCTCGCTAATATCGAAGCATCAGTGTGCCGAATCTCACTGACGGGACACGCGCCGATCGTTATCGCGTCCGTTTATCTTCCACCGGATAAGATCGTTCTAAGCAGTGATATCGAGGCGCTGCTCGGTATGGGGAGCTCTGTCATTCTGGCGGGCGACCTAAATTGTAAACACATCAGGTGGAACTCACACACCACAACCCCGAATGGCAGGCGGCTTGACGCGTTAGTCGATGATCTCGCCTTCGATATCGTCGCTCCGCTAACCCCGACTCACTACCCGCTAAATATCGCGCATCGCCCGGATATACTCGACATAGCGTTATTAAAAAACGTAACTCTGCGCTTACACTCGATCGAAGTAGTTTCAGAGTTAGATTCAGACCACCGTCCCGTCGTTATGAAGCTCGGTCGCGCTCCCGATTCCGTTCCCGTCACGAGGACTGTGGTGGATTGGCACACGCTGGGCATCAGCCTGGCTGAATCTGATCCACCATCGCTTCCGTTTAGTCCGGACTCTATCCCGTCTCCTCAGGATACCGCTGAAGCCATAGACATCGTAACCTCACACATCACCTCGACATTAGATAGGTCATCGAAGCAAGTTGTAGCGGAGGACTTCCTTCACCGCTTCAAATTGCCCGACGATATTAGGGAACTCCTTAGAGCTAAGAACGCCTCGATCCGTGCGTACGATAGGTATCCTACCGCGGAAAATCGTATTCGAATGCGTGCCCTACAACGCGACGTAAAGTCTCGCATCACCGAAGTCCGAGATGCCAGATGGTCTGATTTCTTAGAAGGACTCGCGCCCTCTCAAAGGTCTTACTACCGCTTAGCTCGTACTCTCAAATCGGATACGGTAGTAACTATGCCCCCCCTCGTAGGCCCCTCAGGCCGACTCGCGGCGTTCGATGATGACGAAAAAGCAGAACTGCTGGCCGATACATTGCAAACCCAGTGCACGCCCAGCACTCAATCCGTGGACCCTGTTCATGTAGAATTAGTAGACAGTGAGGTAGAACGCAGAGCCTCCTTGCCACCCTCTGATGCGTTACCACCCGTCACCCCGATGGAAGTTAAAGACTTGATCAAAGACCTACGTCCTCGCAAGGCTCCCGGTTCCGACGGTATATCCAACCGCGTTATTAAACTTCTACCCGTCCAACTCATCGTGATGTTGGCATCTATTTTCAATGCCGCTATGGCGAACTGTATCTTTCCCGCGGTGTGGAAAGAAGCGGACGTTATCGGCATACATAAACCCGGTAAACCAAAAAATCATCCGACGAGCTACCGCCCGATTAGCCTCCTCATGTCTCTAGGCAAACTGTATGAGCGTCTGCTCTACAAACGCCTCAGAGACTTCGTCTCATCCAAGGGCATTCTTATCGATGAACAATTCGGATTCCGTACAAATCACTCATGCGTTCAACAGGTGCACCGCCTCACGGAGCACATTCTTGTGGGGCTTAATCGACCAAAACCGTTATACACGGGAGCTCTCTTCTTCGACGTCGCAAAAGCGTTCGACAAAGTCTGGCACAATGGTTTGATTTTCAAACTATTCAACATGGGCGTGCCGGATAGTCTCGTGCTCATCATACGGGACTTCTTGTCGAACCGCTCTTTTCGATATCGAGTCGAGGGAACCCGCTCCTCCCCACGACCTCTCACAGCTGGAGTCCCGCAAGGCTCTGTCCTCTCACCCCTCCTATTTAGCTTATTCGTCAACGATATTCCCCGGTCGCCGCCGACCCATTTAGCTTTATTCGCCGACGACACGACTGTTTACTATTCTAGTAGAAATAAGTCCCTAATCGCGAAGAAGCTTCAGAGCGCAGCCCTAGCTCTAGGACAGTGGTTCCGAAAATGGCGCATAGACATCAACCCAGCGAAAAGTACTGCGGTGCTATTTCAGAGGGGAAGCTCCACACGGATTTCCTCCCGGATTAGGAGGAGGAATCTCACACCCCCGATTACTCTCTTTAGACAACCCATACCCTGGGCCAGGAAGGTCAAGTACCTGGGCGTTACCCTGGATGCATCGATGACATTCCGCCCGCATATAAAATCAGTCCGTGACCGTGCCGCGTTTATTCTCGGTAGACTCTACCCCATGATCTGTAAGCGGAGTAAAATGTCCCTTCGGAACAAGGTGACACTTTACAAAACTTGCATAAGGCCCGTCATGACTTACGCGAGTGTGGTGTTCGCTCACGCGGCCCGCACACACATAGACACCCTCCAATCCCTACAATCCCGCTTTTGCAGGTTAGCTGTCGGGGCTCCGTGGTTCGTGAGGAACGTTGACCTACACGACGACCTGGGCCTCGAATCAATTCGGAAATACATGAAGTCAGCGTCGGAACGATACTTCGATAAGGCTATGCGTCATGATAATCGCCTTATCGTTGCCGCCGCTGACTACTCCCCGAATCCTGATCATGCAGGAGCCAGTCACCGTCGACGCCCTAGACACGTCCTTACGGATCCATCAGATCCAATAACCTTTGCATTAGATGCCTTCAGCTCTAATACTAGGGGCAGGCTTAGGGACCCCGACAAAGTCTGGCACAACGGTTTGATATACAAACTGTATAACATGGGAGTGCCAGACAGACTCGTGCTCATCATACGAGACTACTTGTCGAACCGTTCGTTCCGATATCGAGTCGAGGGAACGCGTTCCCGGCCCCGTCACGTCACAGCCGGAGTCCCGCAAGGCTCCGCCCTCTCCCCGTTACTATTCAGTTTGTATATCAACGATATACCCCGGTCTCCGGAGACCCATCTGGCGCTCTTCGCCGATGACACCGCCATCTACTACTCGTGTAGGAAGAAGGCGTTGCTTCATCGACGACTTCAGACCGCAGCTACCACCATGGGACAGTGGTTCCGGAAGTGGCGCATCGACATTAACCCCACGAAAAGCACAGCGGTGCTCTTCAAAAGGGGTCGCCCTCCGAACACCACGCTGAGCATCCCTCTCCCGACTAGGCGCGTCAACACCCCCGCCCCCGCCGTTCGCCCAATCACGATGTACGACCAGCCCATACCGTGGGCCCCGAAGGTCAAATATTTAGGCGTCACCCTCGACAGTAGGATGACATTCCGCCCCCACATCAAGACGGTACGCGATCGTGCCGCCTTCATCCTAGGACGTCTCTACCCGATGATATGTAGGCGAAGTAAAATGTCCCTTAGAAATAAGGTGACACTCTACAAAACTTGCATACGCCCCGTCATGACCTATGCAAGTGTAGTGTTCGCTCACGCGGCCCGCATACACTTAAAATCCTTTCAAATCATTCAATCCCGTTTTTGCAGGATAGCCGTCGGAGCCCCGTGGTTCGTCAGGAACGTCGACCTCCATGACGACCTGGACTTAGAGTCCATCAGTAAGTATCTTCAGTCGGCGTCCATGCGCCACTTCGATAAAGCGGCACGACACGAGAACCCTCTCATCGTGGCCGCCGGTAACTACATTCCCGATCCTGCGGACAGAATGGAAAGCAGTCGACGTCGCCCAAAACACGTCATCTCGGATCCTCCCGATCCACTAACGGTGCTTTTAGATCAGAATACTAGTAGTGGTACGCCACAAGAGCGTGAATGCAGCATACGGAAGTCGTGTAATAACGTCTCTGTTACCTCGATGGAAGATCTGAGCCACGAAGAAATCGTCACATCTTACGTTCTGGCTCATGTTGCCCAGTTTGACTGTCGACGTCATCACATGGCGTTTACTAACGGAAATACAACACGTATCAAGCTATTGCAAGAAATTAAGGCAAGTGCAAATCTCAAGTGA
Protein
MDAVFAEFLRLRHPQLASEFLAFKANHTASPLEDSAALAAPASPVPACRASALSTAASVVPAAPVSPILARKAAASSAVTIVSAERSSAASVAPSKTPTLARRSPAPASSCSDSDSDMEVDLAPASSTDGFTLVQKGKKRAAESRAPAAAKISKAVNASRPRPQTPVAPPARATPSPRPVAQNKTQTPPPVILQEKAAWDRVCLALKAKNINFTNARNLANGIQIKVQTPDDHRALSSYLRKERISFHTYTLQEERELRVVIRGIPKELDVELVKADLLEQGLPVNSVHRMHTGRGREPYNMVLVALQPTPEVQETLLKPARRDPKIANYNMVRNDRLSARGGGTVIYYRRALHCVPLDPPALANIEASVCRISLTGHAPIVIASVYLPPDKIVLSSDIEALLGMGSSVILAGDLNCKHIRWNSHTTTPNGRRLDALVDDLAFDIVAPLTPTHYPLNIAHRPDILDIALLKNVTLRLHSIEVVSELDSDHRPVVMKLGRAPDSVPVTRTVVDWHTLGISLAESDPPSLPFSPDSIPSPQDTAEAIDIVTSHITSTLDRSSKQVVAEDFLHRFKLPDDIRELLRAKNASIRAYDRYPTAENRIRMRALQRDVKSRITEVRDARWSDFLEGLAPSQRSYYRLARTLKSDTVVTMPPLVGPSGRLAAFDDDEKAELLADTLQTQCTPSTQSVDPVHVELVDSEVERRASLPPSDALPPVTPMEVKDLIKDLRPRKAPGSDGISNRVIKLLPVQLIVMLASIFNAAMANCIFPAVWKEADVIGIHKPGKPKNHPTSYRPISLLMSLGKLYERLLYKRLRDFVSSKGILIDEQFGFRTNHSCVQQVHRLTEHILVGLNRPKPLYTGALFFDVAKAFDKVWHNGLIFKLFNMGVPDSLVLIIRDFLSNRSFRYRVEGTRSSPRPLTAGVPQGSVLSPLLFSLFVNDIPRSPPTHLALFADDTTVYYSSRNKSLIAKKLQSAALALGQWFRKWRIDINPAKSTAVLFQRGSSTRISSRIRRRNLTPPITLFRQPIPWARKVKYLGVTLDASMTFRPHIKSVRDRAAFILGRLYPMICKRSKMSLRNKVTLYKTCIRPVMTYASVVFAHAARTHIDTLQSLQSRFCRLAVGAPWFVRNVDLHDDLGLESIRKYMKSASERYFDKAMRHDNRLIVAAADYSPNPDHAGASHRRRPRHVLTDPSDPITFALDAFSSNTRGRLRDPDKVWHNGLIYKLYNMGVPDRLVLIIRDYLSNRSFRYRVEGTRSRPRHVTAGVPQGSALSPLLFSLYINDIPRSPETHLALFADDTAIYYSCRKKALLHRRLQTAATTMGQWFRKWRIDINPTKSTAVLFKRGRPPNTTLSIPLPTRRVNTPAPAVRPITMYDQPIPWAPKVKYLGVTLDSRMTFRPHIKTVRDRAAFILGRLYPMICRRSKMSLRNKVTLYKTCIRPVMTYASVVFAHAARIHLKSFQIIQSRFCRIAVGAPWFVRNVDLHDDLDLESISKYLQSASMRHFDKAARHENPLIVAAGNYIPDPADRMESSRRRPKHVISDPPDPLTVLLDQNTSSGTPQERECSIRKSCNNVSVTSMEDLSHEEIVTSYVLAHVAQFDCRRHHMAFTNGNTTRIKLLQEIKASANLK
Summary
Uniprot
ProteinModelPortal
Ontologies
GO
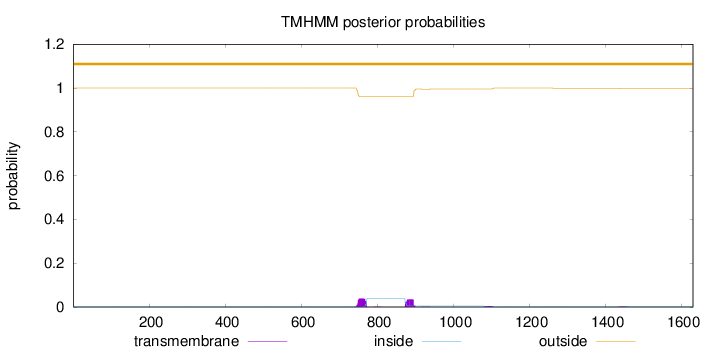
Topology
Length:
1631
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
1.78225
Exp number, first 60 AAs:
0.00071
Total prob of N-in:
0.00011
outside
1 - 1631
Population Genetic Test Statistics
Pi
110.435223
Theta
1.124601
Tajima's D
1.297822
CLR
199.591884
CSRT
0.8027598620069
Interpretation
Uncertain
