Gene
KWMTBOMO04831
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 1.157 Nuclear Reliability : 1.008 PlasmaMembrane Reliability : 1.458
Sequence
CDS
ATGAAAATCCGACTTATAACTCGTCAATTTAAGCCAAAGACTTGGGCTATTGAGAATGCGTCTGGAGATACTGTCACTGAGATCAAAAAGATTGTTAGCGCTTGGAAAGACTATTGCAAGTCTCTGTTTACTGACACCACAACAAAGATCTCAATGACCTACAGTGAACCCTCGGAACTGGAGCCGATGCCTCTAAAAGACGAGGTGCGAGCAGCTATCAACCACCTTAAACGGAATAAGGCTGTAGGTTTTGATGAAATCCCCATCGAGACAATAAAAGCCATGGGGGAAATCGGAGTGGATATACTGCACATCATCTGTTGTCGAATTTGGGTCACTGGAGTTTGGCCGAGCGATTGGTCACATTCGATATTCATTCCACTTCACAAAAAGGGATCAACGAAGAAGTGCAATAACTACAGACTAATTTCTCTCGTATCCCACGCCAGTAAAGTCATGCTGCACATAATTAACATGAGGCTCCAAGATTACTTGTCCCGAGAGATCGCCCTCGAGCAAGCAGGTTTTGTAAAAGGTCGAGCCACAAGAGAGCAACTACTCGTAATGCGGCAAATTGTAGAAAAGGCCAGAGAATTTAATATTAGCCTATATGTGTGCTTCGTTAATTTCCGAAAGGCTTTTGATACCGTCAAGTGGTGGAAGCTGTGGTTGGTACTGACGGAAATGGGAGTACCGCAACATCTTGTACACACCCTCAGACGTCTGTACGAAGATGGCACGGCCGCTGTTCGTATTGACAGTATTGACTCCGAACGCTTCAGCACACAGGCAGGCGTTAGGCAGGGCGGCATACTATCTCCTCTTTTATTTAATATATACACAGAAGACGAAATCGAGGTGCTTCTTCGCCGGCTGGAGACTACAGCGTTGGATTTTGGACTTGCGATCAATCGTGATAAGACCAAAATGATGATCGTAGATCGTGCTAACATCAACCAACCGGAAGTCCAACATATAGCGGGATGCGAGGTAGTTAATAGCTATGTATATCTAGGCTCCACTATCACCAACGCTGGAGGCTGCGAAGATGAGATCCGAAGACGTTGCGCCGTGACAAGATCCTTAGTTGAGAGGCTAACAAAGATTTGGAGAGACCGCAGAATAACCAAAAACACAAAAGTTCGGTTAATGAGATGCCTGGTCTTTCCAACATTTCTGTATGGGGCTGAGACGTGGACGCTGAGGATGCAAGACCGTCGTAAGATTGACGCACTGAAAATGTGGTATTGGAGACGTCTCCTCAGAATTCCGTGTACCGCGCACCGTACCAACATTTCGATCCTGAAAGAGCTCAACATCAAGGAGAGACTATCGTCAACAGTCCACTTACAAATCCTCAAGTTCTTCGGGCACATCACTCGTAATGAGGACTCTATTGAGCGGTTGGTGGTGCAAGGGAAAGTCGAGGGCAAAAGATCTCGCGGACGATCTCCAACACGATGGACCGACCTCATAAAATCTGTGACTCACTCCAACATAAATGTTTGCTCTCACTCTGCGAAATATCGTGCCACATGGCGTCGCATCGCGAGAGCATCAGTATCGCAGGATCCGACTGATGCATGA
Protein
MKIRLITRQFKPKTWAIENASGDTVTEIKKIVSAWKDYCKSLFTDTTTKISMTYSEPSELEPMPLKDEVRAAINHLKRNKAVGFDEIPIETIKAMGEIGVDILHIICCRIWVTGVWPSDWSHSIFIPLHKKGSTKKCNNYRLISLVSHASKVMLHIINMRLQDYLSREIALEQAGFVKGRATREQLLVMRQIVEKAREFNISLYVCFVNFRKAFDTVKWWKLWLVLTEMGVPQHLVHTLRRLYEDGTAAVRIDSIDSERFSTQAGVRQGGILSPLLFNIYTEDEIEVLLRRLETTALDFGLAINRDKTKMMIVDRANINQPEVQHIAGCEVVNSYVYLGSTITNAGGCEDEIRRRCAVTRSLVERLTKIWRDRRITKNTKVRLMRCLVFPTFLYGAETWTLRMQDRRKIDALKMWYWRRLLRIPCTAHRTNISILKELNIKERLSSTVHLQILKFFGHITRNEDSIERLVVQGKVEGKRSRGRSPTRWTDLIKSVTHSNINVCSHSAKYRATWRRIARASVSQDPTDA
Summary
Uniprot
D7F162
D7F167
D7F175
D7F163
D7F161
D7F172
+ More
D7F168 D5LB39 A0A2W1BGD9 D7F174 A0A3Q1LQN1 A0A2W1BNH4 A0A2H6KKM9 A0A3S3P648 A0A3Q1LR18 A0A3Q1MJK8 A0A3Q1M1T7 A0A3Q1MQ02 A0A3Q1MMW6 A0A3Q1MI63 A0A3Q1ML42 H3B2Y3 A0A3Q1LXQ5 A0A3Q1N3Y9 A0A3Q1MA23 A0A3Q1LUA6 A0A3Q1MXT5 A0A3Q1MDY8 A0A3Q1MLI2 A0A3Q1LWU3 A0A3Q1MTQ8 A0A3Q1NE48 A0A3Q1MCA0 A0A3Q1MWB3 A0A3Q1N1P1 A0A3Q1LNZ2 A0A3Q1MLZ9 A0A3Q1LI58 A0A3Q1LRD2 A0A3Q1LXF5 A0A3Q1MP18 A0A3Q1MLV4 A0A3Q1N2X2 A0A3Q1M8D1 A0A3Q1MZJ0 A0A3Q1M6J7 A0A3Q1LR52 A0A3Q1LIW6 A0A3Q1LWH0 A0A3Q1MGH6 A0A3Q1LX64 A0A3Q1LYT3 A0A3Q1MNB9 A0A3Q1MSP2 A0A3Q1MJ53 A0A3Q1LWK8 A0A3Q1MS12 A0A3Q1M527 A0A3Q1LVS6 A0A3S2L0R7 A0A3Q1M0W5 A0A3Q1MM54 A0A3Q1NL67 A0A0J7KGF7 A0A3Q1MKR5 A0A3Q1MCP5 A0A3Q1NFT8 A0A3Q1M6I4 A0A3Q1LWQ9 A0A3Q1MKS2 A0A3Q1MEQ0 A0A3Q1LR83 A0A3Q1NAZ9 A0A3Q1M090 A0A3Q1M6V6 A0A3Q1MZD1 A0A3Q1MLF3 A0A3Q1N9U5 A0A3Q1MGI7 W5N9W4 A0A3Q1M4W4 A0A3Q1N5E6 D5LB38 A0A3Q1N0I8 W5MYT3 A0A3Q1LWK2 A0A3Q1LUF2 H3A382 A0A3Q1MLX8 A0A3Q1MD43 A0A0U4HM41 A0A3Q1LNR6 A0A3Q1MML0 A0A3Q1LWF9 A0A3Q1LLN0
D7F168 D5LB39 A0A2W1BGD9 D7F174 A0A3Q1LQN1 A0A2W1BNH4 A0A2H6KKM9 A0A3S3P648 A0A3Q1LR18 A0A3Q1MJK8 A0A3Q1M1T7 A0A3Q1MQ02 A0A3Q1MMW6 A0A3Q1MI63 A0A3Q1ML42 H3B2Y3 A0A3Q1LXQ5 A0A3Q1N3Y9 A0A3Q1MA23 A0A3Q1LUA6 A0A3Q1MXT5 A0A3Q1MDY8 A0A3Q1MLI2 A0A3Q1LWU3 A0A3Q1MTQ8 A0A3Q1NE48 A0A3Q1MCA0 A0A3Q1MWB3 A0A3Q1N1P1 A0A3Q1LNZ2 A0A3Q1MLZ9 A0A3Q1LI58 A0A3Q1LRD2 A0A3Q1LXF5 A0A3Q1MP18 A0A3Q1MLV4 A0A3Q1N2X2 A0A3Q1M8D1 A0A3Q1MZJ0 A0A3Q1M6J7 A0A3Q1LR52 A0A3Q1LIW6 A0A3Q1LWH0 A0A3Q1MGH6 A0A3Q1LX64 A0A3Q1LYT3 A0A3Q1MNB9 A0A3Q1MSP2 A0A3Q1MJ53 A0A3Q1LWK8 A0A3Q1MS12 A0A3Q1M527 A0A3Q1LVS6 A0A3S2L0R7 A0A3Q1M0W5 A0A3Q1MM54 A0A3Q1NL67 A0A0J7KGF7 A0A3Q1MKR5 A0A3Q1MCP5 A0A3Q1NFT8 A0A3Q1M6I4 A0A3Q1LWQ9 A0A3Q1MKS2 A0A3Q1MEQ0 A0A3Q1LR83 A0A3Q1NAZ9 A0A3Q1M090 A0A3Q1M6V6 A0A3Q1MZD1 A0A3Q1MLF3 A0A3Q1N9U5 A0A3Q1MGI7 W5N9W4 A0A3Q1M4W4 A0A3Q1N5E6 D5LB38 A0A3Q1N0I8 W5MYT3 A0A3Q1LWK2 A0A3Q1LUF2 H3A382 A0A3Q1MLX8 A0A3Q1MD43 A0A0U4HM41 A0A3Q1LNR6 A0A3Q1MML0 A0A3Q1LWF9 A0A3Q1LLN0
EMBL
FJ265547
ADI61815.1
FJ265552
ADI61820.1
FJ265560
ADI61828.1
+ More
FJ265548 ADI61816.1 FJ265546 ADI61814.1 FJ265557 ADI61825.1 FJ265553 ADI61821.1 GU815090 ADF18553.1 KZ150203 PZC72267.1 FJ265559 ADI61827.1 KZ150025 PZC74827.1 BDSA01000081 GBE63537.1 NCKU01004395 RWS06003.1 AFYH01119920 RSAL01000289 RVE42931.1 LBMM01007694 KMQ89503.1 AHAT01022314 GU815089 ADF18552.1 AHAT01017643 AFYH01204777 AFYH01204778 AFYH01204779 KU311054 ALX81665.1
FJ265548 ADI61816.1 FJ265546 ADI61814.1 FJ265557 ADI61825.1 FJ265553 ADI61821.1 GU815090 ADF18553.1 KZ150203 PZC72267.1 FJ265559 ADI61827.1 KZ150025 PZC74827.1 BDSA01000081 GBE63537.1 NCKU01004395 RWS06003.1 AFYH01119920 RSAL01000289 RVE42931.1 LBMM01007694 KMQ89503.1 AHAT01022314 GU815089 ADF18552.1 AHAT01017643 AFYH01204777 AFYH01204778 AFYH01204779 KU311054 ALX81665.1
Proteomes
Interpro
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
D7F162
D7F167
D7F175
D7F163
D7F161
D7F172
+ More
D7F168 D5LB39 A0A2W1BGD9 D7F174 A0A3Q1LQN1 A0A2W1BNH4 A0A2H6KKM9 A0A3S3P648 A0A3Q1LR18 A0A3Q1MJK8 A0A3Q1M1T7 A0A3Q1MQ02 A0A3Q1MMW6 A0A3Q1MI63 A0A3Q1ML42 H3B2Y3 A0A3Q1LXQ5 A0A3Q1N3Y9 A0A3Q1MA23 A0A3Q1LUA6 A0A3Q1MXT5 A0A3Q1MDY8 A0A3Q1MLI2 A0A3Q1LWU3 A0A3Q1MTQ8 A0A3Q1NE48 A0A3Q1MCA0 A0A3Q1MWB3 A0A3Q1N1P1 A0A3Q1LNZ2 A0A3Q1MLZ9 A0A3Q1LI58 A0A3Q1LRD2 A0A3Q1LXF5 A0A3Q1MP18 A0A3Q1MLV4 A0A3Q1N2X2 A0A3Q1M8D1 A0A3Q1MZJ0 A0A3Q1M6J7 A0A3Q1LR52 A0A3Q1LIW6 A0A3Q1LWH0 A0A3Q1MGH6 A0A3Q1LX64 A0A3Q1LYT3 A0A3Q1MNB9 A0A3Q1MSP2 A0A3Q1MJ53 A0A3Q1LWK8 A0A3Q1MS12 A0A3Q1M527 A0A3Q1LVS6 A0A3S2L0R7 A0A3Q1M0W5 A0A3Q1MM54 A0A3Q1NL67 A0A0J7KGF7 A0A3Q1MKR5 A0A3Q1MCP5 A0A3Q1NFT8 A0A3Q1M6I4 A0A3Q1LWQ9 A0A3Q1MKS2 A0A3Q1MEQ0 A0A3Q1LR83 A0A3Q1NAZ9 A0A3Q1M090 A0A3Q1M6V6 A0A3Q1MZD1 A0A3Q1MLF3 A0A3Q1N9U5 A0A3Q1MGI7 W5N9W4 A0A3Q1M4W4 A0A3Q1N5E6 D5LB38 A0A3Q1N0I8 W5MYT3 A0A3Q1LWK2 A0A3Q1LUF2 H3A382 A0A3Q1MLX8 A0A3Q1MD43 A0A0U4HM41 A0A3Q1LNR6 A0A3Q1MML0 A0A3Q1LWF9 A0A3Q1LLN0
D7F168 D5LB39 A0A2W1BGD9 D7F174 A0A3Q1LQN1 A0A2W1BNH4 A0A2H6KKM9 A0A3S3P648 A0A3Q1LR18 A0A3Q1MJK8 A0A3Q1M1T7 A0A3Q1MQ02 A0A3Q1MMW6 A0A3Q1MI63 A0A3Q1ML42 H3B2Y3 A0A3Q1LXQ5 A0A3Q1N3Y9 A0A3Q1MA23 A0A3Q1LUA6 A0A3Q1MXT5 A0A3Q1MDY8 A0A3Q1MLI2 A0A3Q1LWU3 A0A3Q1MTQ8 A0A3Q1NE48 A0A3Q1MCA0 A0A3Q1MWB3 A0A3Q1N1P1 A0A3Q1LNZ2 A0A3Q1MLZ9 A0A3Q1LI58 A0A3Q1LRD2 A0A3Q1LXF5 A0A3Q1MP18 A0A3Q1MLV4 A0A3Q1N2X2 A0A3Q1M8D1 A0A3Q1MZJ0 A0A3Q1M6J7 A0A3Q1LR52 A0A3Q1LIW6 A0A3Q1LWH0 A0A3Q1MGH6 A0A3Q1LX64 A0A3Q1LYT3 A0A3Q1MNB9 A0A3Q1MSP2 A0A3Q1MJ53 A0A3Q1LWK8 A0A3Q1MS12 A0A3Q1M527 A0A3Q1LVS6 A0A3S2L0R7 A0A3Q1M0W5 A0A3Q1MM54 A0A3Q1NL67 A0A0J7KGF7 A0A3Q1MKR5 A0A3Q1MCP5 A0A3Q1NFT8 A0A3Q1M6I4 A0A3Q1LWQ9 A0A3Q1MKS2 A0A3Q1MEQ0 A0A3Q1LR83 A0A3Q1NAZ9 A0A3Q1M090 A0A3Q1M6V6 A0A3Q1MZD1 A0A3Q1MLF3 A0A3Q1N9U5 A0A3Q1MGI7 W5N9W4 A0A3Q1M4W4 A0A3Q1N5E6 D5LB38 A0A3Q1N0I8 W5MYT3 A0A3Q1LWK2 A0A3Q1LUF2 H3A382 A0A3Q1MLX8 A0A3Q1MD43 A0A0U4HM41 A0A3Q1LNR6 A0A3Q1MML0 A0A3Q1LWF9 A0A3Q1LLN0
Ontologies
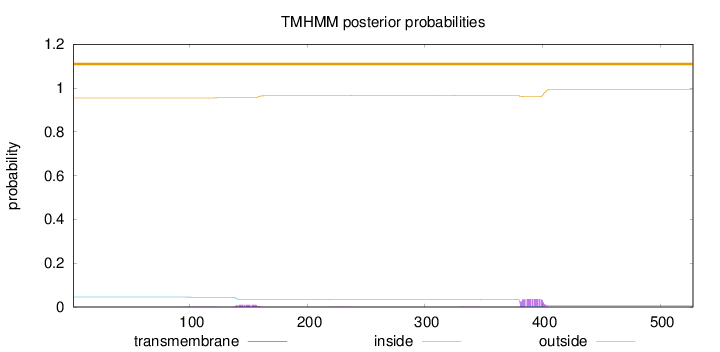
Topology
Length:
528
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.96905
Exp number, first 60 AAs:
0
Total prob of N-in:
0.04585
outside
1 - 528
Population Genetic Test Statistics
Pi
0.201365
Theta
0.946273
Tajima's D
-0.805225
CLR
8.638389
CSRT
0.182490875456227
Interpretation
Uncertain
