Gene
KWMTBOMO04280
Pre Gene Modal
BGIBMGA012647
Annotation
PREDICTED:_lipase_1-like_[Bombyx_mori]
Full name
Lipase
Location in the cell
PlasmaMembrane Reliability : 3.78
Sequence
CDS
ATGTCGTTCAAAACATTCAAATTAATAAGTGCTCTCGTATTCCTAACATTTAAGCGCGGTGTAGCCACGGAAGACTCGTCCGGGACCGTACCAAGCTTTTCATCAATAGCTGAAGCCCTCGGATACGATTGCATTGTTCACACGATCACAACGAAAGACGGTTACATACTCACTCTGTTCAATATACCGGGTAACAAAACCAGGCCGATTTTTTTTATGCACGGATTCGGAGATTCCTCACGCACGTTCATAGTCCGAGGGAACACATCTATGTTGGTATCCGTGGCCAAAGCCGGGTATGACGTGTGGGCCGGGAACATGAGGGGGAATGAATACTCCCGCGACCACGTGACGCTGAACGCAGATATAGACGTCGCGGCCTACTGGAATTTTTCGTACCACGAAGTCGGCGTGTACGATCTGCCCGCCCAAATCGATTACATACTGAACAAGACCCAACAAAATCGACTGAGTGTCATAAGTCATTCCCAAGGAAGCGCCGTATTCTTCGTGCTCTTATCCACTTACCCAGAGTACAATGAAAAGATTCGACCCCTAATAACACTAGGGCCTTTATCATACGCTCAAAACATACGACCTCCAGTTAGATTATTGATTCTAATCGCGCCCGTTTTGGAGGTAATATTCAAGGCATTAAACCAGTACGAGCTGTTTGATAGAACTACGAGACTCATCGTAGAATTGCTTTGCAGTCTCCGACCGTTCGGTGGTGAATTGTGCGAAGCCATACTGCTGAGCACGGCGGGACAAAGCGACGACACGTTCAAAGGAGGATTCATTCGCACGGTCATAAATTACTACCCGGTGAGCACTTCAGTTAAGAACAGCCTGCACGTTGCCCAAAACAGTAACAGCAGAAGGTTCGCCCAATACGATTACGGTCCGGCGGCGAACATCGCTAGATACGGCAACGCTCTTCCACCGGCTTACGATCTGAGCAAAGTGAACTGCAGCGTAGCAATTCTCATGGCCAAAAACGACAAGCTGTCGCCAATCGAAGATAACGTGATGTTAAATAATTCACTGCCTAATGTTGTCGTATATGAGTTTGTTGAACGGAGCCCAATGAATCATTACGACTTCGTGTGGGGCGAAGATATGTATTTGATCTTACATCCGTTTATTTTGAAATTACTAGAGCGGTATCAGTAA
Protein
MSFKTFKLISALVFLTFKRGVATEDSSGTVPSFSSIAEALGYDCIVHTITTKDGYILTLFNIPGNKTRPIFFMHGFGDSSRTFIVRGNTSMLVSVAKAGYDVWAGNMRGNEYSRDHVTLNADIDVAAYWNFSYHEVGVYDLPAQIDYILNKTQQNRLSVISHSQGSAVFFVLLSTYPEYNEKIRPLITLGPLSYAQNIRPPVRLLILIAPVLEVIFKALNQYELFDRTTRLIVELLCSLRPFGGELCEAILLSTAGQSDDTFKGGFIRTVINYYPVSTSVKNSLHVAQNSNSRRFAQYDYGPAANIARYGNALPPAYDLSKVNCSVAILMAKNDKLSPIEDNVMLNNSLPNVVVYEFVERSPMNHYDFVWGEDMYLILHPFILKLLERYQ
Summary
Similarity
Belongs to the AB hydrolase superfamily. Lipase family.
Feature
chain Lipase
Uniprot
H9JSY5
A0A0N1IGF0
H9JT83
A0A194Q1A5
A0A0L7L277
H9JYF4
+ More
H9JSY7 A0A1E1WS39 H9JYF5 A0A0L7KQV5 A0A212F2S9 A0A2H1VHX0 A0A194QRY3 A0A2W1BHX8 H9JXH3 H9JXK1 H9JSX2 I4DP30 H9JSY6 A0A194Q1X8 A0A212FGM8 A0A0L7L5K9 A0A212F0W8 S4PBS0 A0A212F2S4 A0A2A4JBG8 A0A1Y1MRZ1 A0A232EX80 A0A067R7B4 A0A2A4J9X8 A0A067RHQ8 A0A1D2MRG2 A0A2J7QW32 W5J548 A0A2J7PY44 A0A0K8UT86 A0A182H3N2 A0A2J7QG16 V5I953 R4WRP2 A0A1D2MVM7 Q7QH37 A0A226E1Q5 A0A182QF27 B4MWR0 A0A182UTW8 A0A182QBZ9 A0A182FL00 A0A1Y9J0S8 A0A182KX99 A0A1I8JTB8 B0W6G8 B0WDK6 A0A2A4JCC5 D2A3G9 A0A182ML09 A0A1I8NAV5 K7J8S8 A0A0M5J1K3 A0A194R3I6 A0A1Y9H2R7 A0A182G6M0 F4W6F8 A0A1W4UT82 A0A182TMH4 A0A2A4K633 A0A1B6C389 A0A182VB02 A0A182HBR9 A0A151X224 A0A1B0A014 A0A1B0G0B6 B4KKR3 A0A1W4W6G4 A0A0Q5WBH0 B0W6G4 A0A195FI51 K7J2Y2 A0A1I8PCV0 T1EA03 A0A182R885 A0A1B0B8Z4 A0A1Y9H2S0 A0A1W4UT02 E9I8R3 A0A158NSS7 B4HWS6 W5J6X8 K7ITV1 B4LUK9 A0A0L7LCZ5 A0A182U574 U5ERJ2 B3N5D9 A0A084VDU8 A0A232F2D1 B4LUJ8 B4JE79 B4LUL0
H9JSY7 A0A1E1WS39 H9JYF5 A0A0L7KQV5 A0A212F2S9 A0A2H1VHX0 A0A194QRY3 A0A2W1BHX8 H9JXH3 H9JXK1 H9JSX2 I4DP30 H9JSY6 A0A194Q1X8 A0A212FGM8 A0A0L7L5K9 A0A212F0W8 S4PBS0 A0A212F2S4 A0A2A4JBG8 A0A1Y1MRZ1 A0A232EX80 A0A067R7B4 A0A2A4J9X8 A0A067RHQ8 A0A1D2MRG2 A0A2J7QW32 W5J548 A0A2J7PY44 A0A0K8UT86 A0A182H3N2 A0A2J7QG16 V5I953 R4WRP2 A0A1D2MVM7 Q7QH37 A0A226E1Q5 A0A182QF27 B4MWR0 A0A182UTW8 A0A182QBZ9 A0A182FL00 A0A1Y9J0S8 A0A182KX99 A0A1I8JTB8 B0W6G8 B0WDK6 A0A2A4JCC5 D2A3G9 A0A182ML09 A0A1I8NAV5 K7J8S8 A0A0M5J1K3 A0A194R3I6 A0A1Y9H2R7 A0A182G6M0 F4W6F8 A0A1W4UT82 A0A182TMH4 A0A2A4K633 A0A1B6C389 A0A182VB02 A0A182HBR9 A0A151X224 A0A1B0A014 A0A1B0G0B6 B4KKR3 A0A1W4W6G4 A0A0Q5WBH0 B0W6G4 A0A195FI51 K7J2Y2 A0A1I8PCV0 T1EA03 A0A182R885 A0A1B0B8Z4 A0A1Y9H2S0 A0A1W4UT02 E9I8R3 A0A158NSS7 B4HWS6 W5J6X8 K7ITV1 B4LUK9 A0A0L7LCZ5 A0A182U574 U5ERJ2 B3N5D9 A0A084VDU8 A0A232F2D1 B4LUJ8 B4JE79 B4LUL0
Pubmed
EMBL
BABH01038896
KQ460480
KPJ14361.1
BABH01034815
KQ459580
KPI99336.1
+ More
JTDY01008561 JTDY01003454 KOB64514.1 KOB69530.1 BABH01043411 BABH01038893 BABH01038894 GDQN01001224 JAT89830.1 JTDY01007194 KOB65389.1 AGBW02010690 OWR48036.1 ODYU01002476 SOQ40012.1 KQ461196 KPJ06301.1 KZ150080 PZC73881.1 BABH01032763 BABH01032765 BABH01038949 AK403268 BAM19670.1 KPI99333.1 AGBW02008642 OWR52889.1 JTDY01002846 KOB70611.1 AGBW02011015 OWR47367.1 GAIX01002709 JAA89851.1 OWR48034.1 NWSH01002215 PCG68894.1 GEZM01023396 JAV88411.1 NNAY01001790 OXU22918.1 KK852827 KDR15390.1 PCG68895.1 KK852673 KDR18742.1 LJIJ01000666 ODM95481.1 NEVH01009768 PNF32784.1 ADMH02002130 ETN58533.1 NEVH01020850 PNF21255.1 GDHF01022533 JAI29781.1 JXUM01107700 KQ565165 KXJ71265.1 NEVH01014836 PNF27530.1 GALX01003535 JAB64931.1 AK417362 BAN20577.1 LJIJ01000475 ODM97066.1 AAAB01008820 EAA05393.4 LNIX01000009 OXA50396.1 AXCN02000090 CH963857 EDW76549.2 APCN01002227 DS231848 EDS36642.1 DS231898 EDS44615.1 NWSH01002126 PCG69082.1 KQ971338 EFA02315.1 AXCM01003911 CP012523 ALC39218.1 KQ460949 KPJ10406.1 JXUM01045142 KQ561427 KXJ78623.1 GL887707 EGI70293.1 NWSH01000119 PCG79374.1 GEDC01029317 GEDC01024313 GEDC01024096 GEDC01020251 JAS07981.1 JAS12985.1 JAS13202.1 JAS17047.1 JXUM01125660 JXUM01125661 JXUM01125662 KQ566971 KXJ69694.1 KQ982580 KYQ54457.1 CCAG010008921 CH933807 EDW12727.1 KRG03395.1 CH954177 KQS70596.1 EDS36638.1 KQ981523 KYN40355.1 GAMD01001453 JAB00138.1 JXJN01010151 JXJN01010152 JXJN01010153 JXJN01010154 GL761642 EFZ23043.1 ADTU01025110 ADTU01025111 ADTU01025112 CH480818 EDW52471.1 ETN58534.1 CH940649 EDW64195.2 KRF81489.1 JTDY01001637 KOB73274.1 GANO01002781 JAB57090.1 EDV59018.1 ATLV01011799 KE524723 KFB36142.1 NNAY01001167 OXU24914.1 EDW64184.1 KRF81481.1 CH916368 EDW03599.1 EDW64196.1
JTDY01008561 JTDY01003454 KOB64514.1 KOB69530.1 BABH01043411 BABH01038893 BABH01038894 GDQN01001224 JAT89830.1 JTDY01007194 KOB65389.1 AGBW02010690 OWR48036.1 ODYU01002476 SOQ40012.1 KQ461196 KPJ06301.1 KZ150080 PZC73881.1 BABH01032763 BABH01032765 BABH01038949 AK403268 BAM19670.1 KPI99333.1 AGBW02008642 OWR52889.1 JTDY01002846 KOB70611.1 AGBW02011015 OWR47367.1 GAIX01002709 JAA89851.1 OWR48034.1 NWSH01002215 PCG68894.1 GEZM01023396 JAV88411.1 NNAY01001790 OXU22918.1 KK852827 KDR15390.1 PCG68895.1 KK852673 KDR18742.1 LJIJ01000666 ODM95481.1 NEVH01009768 PNF32784.1 ADMH02002130 ETN58533.1 NEVH01020850 PNF21255.1 GDHF01022533 JAI29781.1 JXUM01107700 KQ565165 KXJ71265.1 NEVH01014836 PNF27530.1 GALX01003535 JAB64931.1 AK417362 BAN20577.1 LJIJ01000475 ODM97066.1 AAAB01008820 EAA05393.4 LNIX01000009 OXA50396.1 AXCN02000090 CH963857 EDW76549.2 APCN01002227 DS231848 EDS36642.1 DS231898 EDS44615.1 NWSH01002126 PCG69082.1 KQ971338 EFA02315.1 AXCM01003911 CP012523 ALC39218.1 KQ460949 KPJ10406.1 JXUM01045142 KQ561427 KXJ78623.1 GL887707 EGI70293.1 NWSH01000119 PCG79374.1 GEDC01029317 GEDC01024313 GEDC01024096 GEDC01020251 JAS07981.1 JAS12985.1 JAS13202.1 JAS17047.1 JXUM01125660 JXUM01125661 JXUM01125662 KQ566971 KXJ69694.1 KQ982580 KYQ54457.1 CCAG010008921 CH933807 EDW12727.1 KRG03395.1 CH954177 KQS70596.1 EDS36638.1 KQ981523 KYN40355.1 GAMD01001453 JAB00138.1 JXJN01010151 JXJN01010152 JXJN01010153 JXJN01010154 GL761642 EFZ23043.1 ADTU01025110 ADTU01025111 ADTU01025112 CH480818 EDW52471.1 ETN58534.1 CH940649 EDW64195.2 KRF81489.1 JTDY01001637 KOB73274.1 GANO01002781 JAB57090.1 EDV59018.1 ATLV01011799 KE524723 KFB36142.1 NNAY01001167 OXU24914.1 EDW64184.1 KRF81481.1 CH916368 EDW03599.1 EDW64196.1
Proteomes
UP000005204
UP000053240
UP000053268
UP000037510
UP000007151
UP000218220
+ More
UP000215335 UP000027135 UP000094527 UP000235965 UP000000673 UP000069940 UP000249989 UP000007062 UP000198287 UP000075886 UP000007798 UP000075903 UP000069272 UP000076407 UP000075882 UP000075840 UP000002320 UP000007266 UP000075883 UP000095301 UP000002358 UP000092553 UP000075884 UP000007755 UP000192221 UP000075902 UP000075809 UP000092445 UP000092444 UP000009192 UP000192223 UP000008711 UP000078541 UP000095300 UP000075900 UP000092460 UP000005205 UP000001292 UP000008792 UP000030765 UP000001070
UP000215335 UP000027135 UP000094527 UP000235965 UP000000673 UP000069940 UP000249989 UP000007062 UP000198287 UP000075886 UP000007798 UP000075903 UP000069272 UP000076407 UP000075882 UP000075840 UP000002320 UP000007266 UP000075883 UP000095301 UP000002358 UP000092553 UP000075884 UP000007755 UP000192221 UP000075902 UP000075809 UP000092445 UP000092444 UP000009192 UP000192223 UP000008711 UP000078541 UP000095300 UP000075900 UP000092460 UP000005205 UP000001292 UP000008792 UP000030765 UP000001070
Interpro
SUPFAM
SSF53474
SSF53474
Gene 3D
ProteinModelPortal
H9JSY5
A0A0N1IGF0
H9JT83
A0A194Q1A5
A0A0L7L277
H9JYF4
+ More
H9JSY7 A0A1E1WS39 H9JYF5 A0A0L7KQV5 A0A212F2S9 A0A2H1VHX0 A0A194QRY3 A0A2W1BHX8 H9JXH3 H9JXK1 H9JSX2 I4DP30 H9JSY6 A0A194Q1X8 A0A212FGM8 A0A0L7L5K9 A0A212F0W8 S4PBS0 A0A212F2S4 A0A2A4JBG8 A0A1Y1MRZ1 A0A232EX80 A0A067R7B4 A0A2A4J9X8 A0A067RHQ8 A0A1D2MRG2 A0A2J7QW32 W5J548 A0A2J7PY44 A0A0K8UT86 A0A182H3N2 A0A2J7QG16 V5I953 R4WRP2 A0A1D2MVM7 Q7QH37 A0A226E1Q5 A0A182QF27 B4MWR0 A0A182UTW8 A0A182QBZ9 A0A182FL00 A0A1Y9J0S8 A0A182KX99 A0A1I8JTB8 B0W6G8 B0WDK6 A0A2A4JCC5 D2A3G9 A0A182ML09 A0A1I8NAV5 K7J8S8 A0A0M5J1K3 A0A194R3I6 A0A1Y9H2R7 A0A182G6M0 F4W6F8 A0A1W4UT82 A0A182TMH4 A0A2A4K633 A0A1B6C389 A0A182VB02 A0A182HBR9 A0A151X224 A0A1B0A014 A0A1B0G0B6 B4KKR3 A0A1W4W6G4 A0A0Q5WBH0 B0W6G4 A0A195FI51 K7J2Y2 A0A1I8PCV0 T1EA03 A0A182R885 A0A1B0B8Z4 A0A1Y9H2S0 A0A1W4UT02 E9I8R3 A0A158NSS7 B4HWS6 W5J6X8 K7ITV1 B4LUK9 A0A0L7LCZ5 A0A182U574 U5ERJ2 B3N5D9 A0A084VDU8 A0A232F2D1 B4LUJ8 B4JE79 B4LUL0
H9JSY7 A0A1E1WS39 H9JYF5 A0A0L7KQV5 A0A212F2S9 A0A2H1VHX0 A0A194QRY3 A0A2W1BHX8 H9JXH3 H9JXK1 H9JSX2 I4DP30 H9JSY6 A0A194Q1X8 A0A212FGM8 A0A0L7L5K9 A0A212F0W8 S4PBS0 A0A212F2S4 A0A2A4JBG8 A0A1Y1MRZ1 A0A232EX80 A0A067R7B4 A0A2A4J9X8 A0A067RHQ8 A0A1D2MRG2 A0A2J7QW32 W5J548 A0A2J7PY44 A0A0K8UT86 A0A182H3N2 A0A2J7QG16 V5I953 R4WRP2 A0A1D2MVM7 Q7QH37 A0A226E1Q5 A0A182QF27 B4MWR0 A0A182UTW8 A0A182QBZ9 A0A182FL00 A0A1Y9J0S8 A0A182KX99 A0A1I8JTB8 B0W6G8 B0WDK6 A0A2A4JCC5 D2A3G9 A0A182ML09 A0A1I8NAV5 K7J8S8 A0A0M5J1K3 A0A194R3I6 A0A1Y9H2R7 A0A182G6M0 F4W6F8 A0A1W4UT82 A0A182TMH4 A0A2A4K633 A0A1B6C389 A0A182VB02 A0A182HBR9 A0A151X224 A0A1B0A014 A0A1B0G0B6 B4KKR3 A0A1W4W6G4 A0A0Q5WBH0 B0W6G4 A0A195FI51 K7J2Y2 A0A1I8PCV0 T1EA03 A0A182R885 A0A1B0B8Z4 A0A1Y9H2S0 A0A1W4UT02 E9I8R3 A0A158NSS7 B4HWS6 W5J6X8 K7ITV1 B4LUK9 A0A0L7LCZ5 A0A182U574 U5ERJ2 B3N5D9 A0A084VDU8 A0A232F2D1 B4LUJ8 B4JE79 B4LUL0
PDB
1K8Q
E-value=3.02335e-43,
Score=441
Ontologies
GO
Topology
SignalP
Position: 1 - 22,
Likelihood: 0.926054

Length:
390
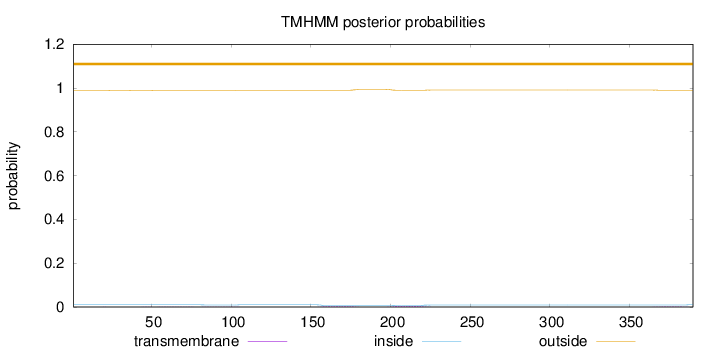
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.22055
Exp number, first 60 AAs:
0.00664
Total prob of N-in:
0.01051
outside
1 - 390
Population Genetic Test Statistics
Pi
2.836442
Theta
6.067833
Tajima's D
-1.49831
CLR
138.475359
CSRT
0.055847207639618
Interpretation
Uncertain
