Gene
KWMTBOMO04189
Pre Gene Modal
BGIBMGA000182
Annotation
PREDICTED:_organic_cation_transporter_protein-like_[Bombyx_mori]
Location in the cell
PlasmaMembrane Reliability : 4.878
Sequence
CDS
ATGCCGGAATCCCCAAAATGGCTAATCTCAGTTGGACGGCTTGAAGAGGCCAGTATTGTTATGACCTCTGCGGCTAAATTCAACAACATCGATCTCCCATCATCAGGTATGTTGTCAGTCGTCGAAACATTGGCCAAAAAGGGAAAAGCTGAAATAAAGACTCAAAGAAAGGCTACTTATATGGACTTGGTCAGATTACCAAGTTTGAGGATTAAGAACTTGTGCTGCTGCGTCGTTTGGCTGGTGATTGGGATAAGCTTTTATGGCGGTACCCAATACTTTGGACAGACCAGTATAAATACATTCTTATCTGTCGCTTTGGGAGGTCTGGTGCAGATCCCTGGGCTACCACTGAGCGCGTATTTCTCGAAAAGATTTGGACGCCGTGTCGCCATCATTGGCATGTTCGTGATTAACTGCATTTGTAACCTGCTATTAGTTCTCCCTGACGCATGGTTTTATCTGAAGCTCGTGGCTGGTTCCATAGCCAACGCTTGCGGAGCTGCTGCGTTTTCTGTAGTTTCTGACATGGACGGTACGAAACGAAAATCAGTTACTATTTCTGAAAAAATAGAAGAGACACCGAAACAAGATGAAGATAAACTGATTCAAGCGATAGGTGAATTCGGGAGATGGCACGTGATGCTAACTATGACGCTATCTGCGCCGATCCGAATGACGGCAGTCTGGAATCATTTGGGAATCATATTTCTGGCACCGAAAACTGAATTTATTTGCGCGGAGAGACGTAATAACAACGAAACCATCCAAGAATCGGAGTGCTACAGCGATTGTACTGAATACGAATACAAAACGGATTTCGTGAATACAATCATTTCTGAATGGGACCTAGTTTGCGATCGAGCCTGGCTGACAAACTTCAATCAGACAGCTTGCATGTCCGGTGTTTTATTGGGCAGTATAGTTTTTGGATTTTTCGCTGATAGATTTGGCCGTCGTCCCGCTCTCCTGACGGCCTGCACCACGCAACTGATAGCTGGATTCACAGCGCCGTTTTGCCCAAATTACATCACATTCACCTTGGTCCGATTCTTCAACGGCGCAGCTGTTGCTGGCGCATTACTGACTTCTTTTATTCTAATGATTGAAACCACGGGCATATCGAAAAGGGAACTAATGAACTGTATCGTTTCCGTGCCTCTGTCGGTGGGAGAAATGATAATGCCAATGTTTTCATATTACTTGAGGTCTTGGACTAGATATAGTTTGGGTCTCGCCCTACCAAATTTGGTATTTTTGGTTTTATTCTTCATTGTACCAGAGTCTCCGAAGTGGTTGATTTCCGTTGGAAGGCTCGAGGAGGCGAGTTTGATCATGACGAAAATAGCGAAATGGAATAGACTTCCAACAGAAGGCATGATGGACAGAGTAAAGATTATAGCATTTGAATCCAATGTTGACGAGCACAAAAACGTCAAGGCAACTTACTTAGACCTGATCAAAGTGCAGGAGATAAGGTGGAAGAATATCGGATGCTGCCTTACGTGGTTTATCCTCGGGCTGAGCTTCTATGGCAGCAATCAGTACATAGGACAGACCAGCCCTAATGTTTTCGTTGCTATTGCAATGGCTGGAGCTATTCAGACTACCTTTGACCCATTCCTTGTTCTATTACTGGCCATAGAGATCGACTTAGCTGGAGGTCTTGAGGCTAATAAACTAATCATAATCGGCCACACGTTAACAGCGAAAAGCTACATGTTCACAACAGAACTCTTCCCGACGGTAGCCAGGAATATGGCGATGGGAGCTTCATCAACGGTCTCCAGAATCGGTTCCATGCTGGCTCCTTTCATCGCTGGTCTTACCGCCACTGGACCTTGGATTCCACCGACTGTGTTCGGCCTTGTACCATTCGCCGCGGCAGTAGCTTGTTACTGTCTACCAGAAACCAGGGGCAAAAAGCTCAAAGATCACTTCGGAGAAACATAA
Protein
MPESPKWLISVGRLEEASIVMTSAAKFNNIDLPSSGMLSVVETLAKKGKAEIKTQRKATYMDLVRLPSLRIKNLCCCVVWLVIGISFYGGTQYFGQTSINTFLSVALGGLVQIPGLPLSAYFSKRFGRRVAIIGMFVINCICNLLLVLPDAWFYLKLVAGSIANACGAAAFSVVSDMDGTKRKSVTISEKIEETPKQDEDKLIQAIGEFGRWHVMLTMTLSAPIRMTAVWNHLGIIFLAPKTEFICAERRNNNETIQESECYSDCTEYEYKTDFVNTIISEWDLVCDRAWLTNFNQTACMSGVLLGSIVFGFFADRFGRRPALLTACTTQLIAGFTAPFCPNYITFTLVRFFNGAAVAGALLTSFILMIETTGISKRELMNCIVSVPLSVGEMIMPMFSYYLRSWTRYSLGLALPNLVFLVLFFIVPESPKWLISVGRLEEASLIMTKIAKWNRLPTEGMMDRVKIIAFESNVDEHKNVKATYLDLIKVQEIRWKNIGCCLTWFILGLSFYGSNQYIGQTSPNVFVAIAMAGAIQTTFDPFLVLLLAIEIDLAGGLEANKLIIIGHTLTAKSYMFTTELFPTVARNMAMGASSTVSRIGSMLAPFIAGLTATGPWIPPTVFGLVPFAAAVACYCLPETRGKKLKDHFGET
Summary
Uniprot
H9ISF5
A0A2W1BFH6
A0A2H1VMD7
A0A2A4K5L0
A0A2A4J3B3
A0A2W1BP27
+ More
A0A194RHR5 A0A2H1VGH7 A0A2A4K9Y1 A0A2A4KA95 I4DNF7 A0A194PMC3 H9JNB3 A0A194QN55 V5IAH1 A0A1B0C9J9 A0A2H1W3H1 A0A194PY14 A0A2A4K9Y5 A0A1Q3FH65 D6WKJ1 A0A1J1IHN1 A0A1Y1KD57 A0A1Y1K9J5 D6WKJ0 B0X4Q4 A0A0L7LUK1 Q17CS1 A0A182H4N7 A0A023EUG3 A0A182Y4D6 A0A182PAB8 A0A2H1VUP3 A0A182VI17 D6X4T8 A0A182TEQ3 A0A182QLT3 Q7QDB3 A0A182HUA7 A0A182WRH0 K7ITG9 A0A182W090 A0A212FFR4 A0A182K794 A0A088ALA3 A0A194QK42 A0A154P8V9 A0A0L7QRN1 A0A182MBN7 A0A182NFM5 U4UII8 A0A182RNN4 A0A2W1BCH7 A0A182JCT1 A0A2J7RBY0 A0A310SN29 E2BIH0 A0A2M4CU13 A0A2M4CTR9 A0A2A4J566 A0A084VXP9 K7JAK3 A0A2A3ESR9 E1ZZY2 A0A088A3V4 A0A194QIQ7 A0A182F7G4 A0A0M9A1C2 A0A232EL12 N6T164 A0A2W1BI79 B4PAQ3 B3NK59 A0A195DRF1 E9J9G6 A0A232F564 A0A310SUT5 B4HPT1 A0A158NPS8 A0A195ATZ5 A0A1W4X9I8 A0A2S2QUM7 Q7K3M6 B4QEI8 A0A2A3ELA5 A0A1W4WYZ1 A0A151IP62 A0A151JT16 A0A195C6E8 A0A195FCZ0 A0A034WDR5 B4MRV4 A0A3S2LHN8 E2A1J5 E9IX20
A0A194RHR5 A0A2H1VGH7 A0A2A4K9Y1 A0A2A4KA95 I4DNF7 A0A194PMC3 H9JNB3 A0A194QN55 V5IAH1 A0A1B0C9J9 A0A2H1W3H1 A0A194PY14 A0A2A4K9Y5 A0A1Q3FH65 D6WKJ1 A0A1J1IHN1 A0A1Y1KD57 A0A1Y1K9J5 D6WKJ0 B0X4Q4 A0A0L7LUK1 Q17CS1 A0A182H4N7 A0A023EUG3 A0A182Y4D6 A0A182PAB8 A0A2H1VUP3 A0A182VI17 D6X4T8 A0A182TEQ3 A0A182QLT3 Q7QDB3 A0A182HUA7 A0A182WRH0 K7ITG9 A0A182W090 A0A212FFR4 A0A182K794 A0A088ALA3 A0A194QK42 A0A154P8V9 A0A0L7QRN1 A0A182MBN7 A0A182NFM5 U4UII8 A0A182RNN4 A0A2W1BCH7 A0A182JCT1 A0A2J7RBY0 A0A310SN29 E2BIH0 A0A2M4CU13 A0A2M4CTR9 A0A2A4J566 A0A084VXP9 K7JAK3 A0A2A3ESR9 E1ZZY2 A0A088A3V4 A0A194QIQ7 A0A182F7G4 A0A0M9A1C2 A0A232EL12 N6T164 A0A2W1BI79 B4PAQ3 B3NK59 A0A195DRF1 E9J9G6 A0A232F564 A0A310SUT5 B4HPT1 A0A158NPS8 A0A195ATZ5 A0A1W4X9I8 A0A2S2QUM7 Q7K3M6 B4QEI8 A0A2A3ELA5 A0A1W4WYZ1 A0A151IP62 A0A151JT16 A0A195C6E8 A0A195FCZ0 A0A034WDR5 B4MRV4 A0A3S2LHN8 E2A1J5 E9IX20
Pubmed
19121390
28756777
26354079
22651552
18362917
19820115
+ More
28004739 26227816 17510324 26483478 24945155 25244985 12364791 14747013 17210077 20075255 22118469 23537049 20798317 24438588 28648823 17994087 17550304 21282665 21347285 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 22936249 25348373
28004739 26227816 17510324 26483478 24945155 25244985 12364791 14747013 17210077 20075255 22118469 23537049 20798317 24438588 28648823 17994087 17550304 21282665 21347285 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 22936249 25348373
EMBL
BABH01044534
BABH01044535
KZ150304
PZC71520.1
ODYU01003353
SOQ42005.1
+ More
NWSH01000158 PCG78950.1 NWSH01003717 PCG65892.1 KZ149948 PZC76718.1 KQ460367 KPJ15486.1 ODYU01002452 SOQ39959.1 NWSH01000020 PCG80714.1 PCG80713.1 AK402884 BAM19447.1 KQ459600 KPI94138.1 BABH01019600 KQ461195 KPJ06789.1 GALX01001195 JAB67271.1 AJWK01002434 AJWK01002435 AJWK01002436 ODYU01006026 SOQ47506.1 KQ459589 KPI97639.1 PCG80708.1 GFDL01008134 JAV26911.1 KQ971342 EFA04007.2 CVRI01000052 CRK99712.1 GEZM01088532 JAV58130.1 GEZM01088533 JAV58129.1 EFA04006.1 DS232350 EDS40471.1 JTDY01000054 KOB79137.1 CH477305 EAT44124.1 JXUM01109993 KQ565372 KXJ71040.1 GAPW01001011 JAC12587.1 ODYU01004551 SOQ44555.1 KQ971381 EEZ97586.2 AXCN02000238 AAAB01008859 EAA07935.5 APCN01000614 AAZX01006832 AGBW02008774 OWR52572.1 KQ461198 KPJ05943.1 KQ434827 KZC07658.1 KQ414775 KOC61282.1 AXCM01003853 KB632314 ERL92208.1 KZ150301 PZC71534.1 NEVH01005889 PNF38331.1 KQ762190 OAD56165.1 GL448504 EFN84558.1 GGFL01004591 MBW68769.1 GGFL01004566 MBW68744.1 NWSH01003004 PCG67121.1 ATLV01018145 KE525215 KFB42743.1 AAZX01002446 KZ288189 PBC34564.1 GL435531 EFN73254.1 KQ459185 KPJ03336.1 KQ435794 KOX73779.1 NNAY01003667 OXU19035.1 APGK01048466 KB741110 ENN73834.1 KZ150029 PZC74769.1 CM000158 EDW91444.1 KRJ99843.1 CH954179 EDV55264.1 KQ980581 KYN15387.1 GL769313 EFZ10529.1 NNAY01000983 OXU25618.1 KQ760085 OAD61975.1 CH480816 EDW48649.1 ADTU01022726 KQ976745 KYM75454.1 GGMS01012221 MBY81424.1 AE013599 AY058478 BT120184 AAF57511.1 AAL13707.1 ADB91432.1 CM000362 CM002911 EDX07862.1 KMY95206.1 KZ288225 PBC31939.1 KQ976914 KYN07128.1 KQ982014 KYN30660.1 KQ978219 KYM96439.1 KQ981673 KYN38256.1 GAKP01006475 JAC52477.1 CH963850 EDW74843.1 RSAL01000008 RVE54031.1 GL435766 EFN72668.1 GL766616 EFZ14896.1
NWSH01000158 PCG78950.1 NWSH01003717 PCG65892.1 KZ149948 PZC76718.1 KQ460367 KPJ15486.1 ODYU01002452 SOQ39959.1 NWSH01000020 PCG80714.1 PCG80713.1 AK402884 BAM19447.1 KQ459600 KPI94138.1 BABH01019600 KQ461195 KPJ06789.1 GALX01001195 JAB67271.1 AJWK01002434 AJWK01002435 AJWK01002436 ODYU01006026 SOQ47506.1 KQ459589 KPI97639.1 PCG80708.1 GFDL01008134 JAV26911.1 KQ971342 EFA04007.2 CVRI01000052 CRK99712.1 GEZM01088532 JAV58130.1 GEZM01088533 JAV58129.1 EFA04006.1 DS232350 EDS40471.1 JTDY01000054 KOB79137.1 CH477305 EAT44124.1 JXUM01109993 KQ565372 KXJ71040.1 GAPW01001011 JAC12587.1 ODYU01004551 SOQ44555.1 KQ971381 EEZ97586.2 AXCN02000238 AAAB01008859 EAA07935.5 APCN01000614 AAZX01006832 AGBW02008774 OWR52572.1 KQ461198 KPJ05943.1 KQ434827 KZC07658.1 KQ414775 KOC61282.1 AXCM01003853 KB632314 ERL92208.1 KZ150301 PZC71534.1 NEVH01005889 PNF38331.1 KQ762190 OAD56165.1 GL448504 EFN84558.1 GGFL01004591 MBW68769.1 GGFL01004566 MBW68744.1 NWSH01003004 PCG67121.1 ATLV01018145 KE525215 KFB42743.1 AAZX01002446 KZ288189 PBC34564.1 GL435531 EFN73254.1 KQ459185 KPJ03336.1 KQ435794 KOX73779.1 NNAY01003667 OXU19035.1 APGK01048466 KB741110 ENN73834.1 KZ150029 PZC74769.1 CM000158 EDW91444.1 KRJ99843.1 CH954179 EDV55264.1 KQ980581 KYN15387.1 GL769313 EFZ10529.1 NNAY01000983 OXU25618.1 KQ760085 OAD61975.1 CH480816 EDW48649.1 ADTU01022726 KQ976745 KYM75454.1 GGMS01012221 MBY81424.1 AE013599 AY058478 BT120184 AAF57511.1 AAL13707.1 ADB91432.1 CM000362 CM002911 EDX07862.1 KMY95206.1 KZ288225 PBC31939.1 KQ976914 KYN07128.1 KQ982014 KYN30660.1 KQ978219 KYM96439.1 KQ981673 KYN38256.1 GAKP01006475 JAC52477.1 CH963850 EDW74843.1 RSAL01000008 RVE54031.1 GL435766 EFN72668.1 GL766616 EFZ14896.1
Proteomes
UP000005204
UP000218220
UP000053240
UP000053268
UP000092461
UP000007266
+ More
UP000183832 UP000002320 UP000037510 UP000008820 UP000069940 UP000249989 UP000076408 UP000075885 UP000075903 UP000075902 UP000075886 UP000007062 UP000075840 UP000076407 UP000002358 UP000075920 UP000007151 UP000075881 UP000005203 UP000076502 UP000053825 UP000075883 UP000075884 UP000030742 UP000075900 UP000075880 UP000235965 UP000008237 UP000030765 UP000242457 UP000000311 UP000069272 UP000053105 UP000215335 UP000019118 UP000002282 UP000008711 UP000078492 UP000001292 UP000005205 UP000078540 UP000192223 UP000000803 UP000000304 UP000078542 UP000078541 UP000007798 UP000283053
UP000183832 UP000002320 UP000037510 UP000008820 UP000069940 UP000249989 UP000076408 UP000075885 UP000075903 UP000075902 UP000075886 UP000007062 UP000075840 UP000076407 UP000002358 UP000075920 UP000007151 UP000075881 UP000005203 UP000076502 UP000053825 UP000075883 UP000075884 UP000030742 UP000075900 UP000075880 UP000235965 UP000008237 UP000030765 UP000242457 UP000000311 UP000069272 UP000053105 UP000215335 UP000019118 UP000002282 UP000008711 UP000078492 UP000001292 UP000005205 UP000078540 UP000192223 UP000000803 UP000000304 UP000078542 UP000078541 UP000007798 UP000283053
Pfam
Interpro
IPR020846
MFS_dom
+ More
IPR036259 MFS_trans_sf
IPR005828 MFS_sugar_transport-like
IPR005829 Sugar_transporter_CS
IPR011701 MFS
IPR015943 WD40/YVTN_repeat-like_dom_sf
IPR022100 Mcl1_mid
IPR017986 WD40_repeat_dom
IPR019775 WD40_repeat_CS
IPR001680 WD40_repeat
IPR036322 WD40_repeat_dom_sf
IPR024977 Apc4_WD40_dom
IPR011041 Quinoprot_gluc/sorb_DH
IPR017996 Royal_jelly/protein_yellow
IPR011042 6-blade_b-propeller_TolB-like
IPR036259 MFS_trans_sf
IPR005828 MFS_sugar_transport-like
IPR005829 Sugar_transporter_CS
IPR011701 MFS
IPR015943 WD40/YVTN_repeat-like_dom_sf
IPR022100 Mcl1_mid
IPR017986 WD40_repeat_dom
IPR019775 WD40_repeat_CS
IPR001680 WD40_repeat
IPR036322 WD40_repeat_dom_sf
IPR024977 Apc4_WD40_dom
IPR011041 Quinoprot_gluc/sorb_DH
IPR017996 Royal_jelly/protein_yellow
IPR011042 6-blade_b-propeller_TolB-like
Gene 3D
CDD
ProteinModelPortal
H9ISF5
A0A2W1BFH6
A0A2H1VMD7
A0A2A4K5L0
A0A2A4J3B3
A0A2W1BP27
+ More
A0A194RHR5 A0A2H1VGH7 A0A2A4K9Y1 A0A2A4KA95 I4DNF7 A0A194PMC3 H9JNB3 A0A194QN55 V5IAH1 A0A1B0C9J9 A0A2H1W3H1 A0A194PY14 A0A2A4K9Y5 A0A1Q3FH65 D6WKJ1 A0A1J1IHN1 A0A1Y1KD57 A0A1Y1K9J5 D6WKJ0 B0X4Q4 A0A0L7LUK1 Q17CS1 A0A182H4N7 A0A023EUG3 A0A182Y4D6 A0A182PAB8 A0A2H1VUP3 A0A182VI17 D6X4T8 A0A182TEQ3 A0A182QLT3 Q7QDB3 A0A182HUA7 A0A182WRH0 K7ITG9 A0A182W090 A0A212FFR4 A0A182K794 A0A088ALA3 A0A194QK42 A0A154P8V9 A0A0L7QRN1 A0A182MBN7 A0A182NFM5 U4UII8 A0A182RNN4 A0A2W1BCH7 A0A182JCT1 A0A2J7RBY0 A0A310SN29 E2BIH0 A0A2M4CU13 A0A2M4CTR9 A0A2A4J566 A0A084VXP9 K7JAK3 A0A2A3ESR9 E1ZZY2 A0A088A3V4 A0A194QIQ7 A0A182F7G4 A0A0M9A1C2 A0A232EL12 N6T164 A0A2W1BI79 B4PAQ3 B3NK59 A0A195DRF1 E9J9G6 A0A232F564 A0A310SUT5 B4HPT1 A0A158NPS8 A0A195ATZ5 A0A1W4X9I8 A0A2S2QUM7 Q7K3M6 B4QEI8 A0A2A3ELA5 A0A1W4WYZ1 A0A151IP62 A0A151JT16 A0A195C6E8 A0A195FCZ0 A0A034WDR5 B4MRV4 A0A3S2LHN8 E2A1J5 E9IX20
A0A194RHR5 A0A2H1VGH7 A0A2A4K9Y1 A0A2A4KA95 I4DNF7 A0A194PMC3 H9JNB3 A0A194QN55 V5IAH1 A0A1B0C9J9 A0A2H1W3H1 A0A194PY14 A0A2A4K9Y5 A0A1Q3FH65 D6WKJ1 A0A1J1IHN1 A0A1Y1KD57 A0A1Y1K9J5 D6WKJ0 B0X4Q4 A0A0L7LUK1 Q17CS1 A0A182H4N7 A0A023EUG3 A0A182Y4D6 A0A182PAB8 A0A2H1VUP3 A0A182VI17 D6X4T8 A0A182TEQ3 A0A182QLT3 Q7QDB3 A0A182HUA7 A0A182WRH0 K7ITG9 A0A182W090 A0A212FFR4 A0A182K794 A0A088ALA3 A0A194QK42 A0A154P8V9 A0A0L7QRN1 A0A182MBN7 A0A182NFM5 U4UII8 A0A182RNN4 A0A2W1BCH7 A0A182JCT1 A0A2J7RBY0 A0A310SN29 E2BIH0 A0A2M4CU13 A0A2M4CTR9 A0A2A4J566 A0A084VXP9 K7JAK3 A0A2A3ESR9 E1ZZY2 A0A088A3V4 A0A194QIQ7 A0A182F7G4 A0A0M9A1C2 A0A232EL12 N6T164 A0A2W1BI79 B4PAQ3 B3NK59 A0A195DRF1 E9J9G6 A0A232F564 A0A310SUT5 B4HPT1 A0A158NPS8 A0A195ATZ5 A0A1W4X9I8 A0A2S2QUM7 Q7K3M6 B4QEI8 A0A2A3ELA5 A0A1W4WYZ1 A0A151IP62 A0A151JT16 A0A195C6E8 A0A195FCZ0 A0A034WDR5 B4MRV4 A0A3S2LHN8 E2A1J5 E9IX20
PDB
4LDS
E-value=8.1236e-09,
Score=146
Ontologies
GO
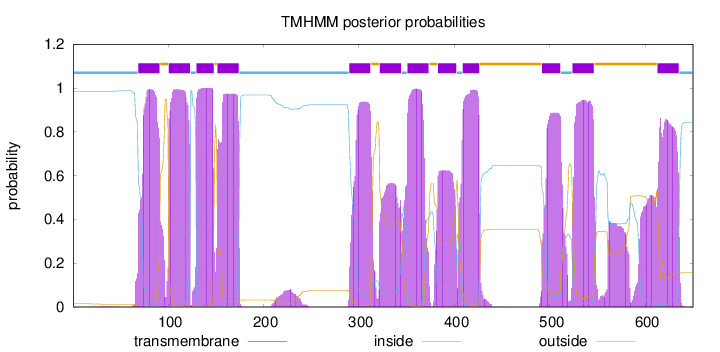
Topology
Length:
650
Number of predicted TMHs:
12
Exp number of AAs in TMHs:
251.67201
Exp number, first 60 AAs:
0.04299
Total prob of N-in:
0.98521
inside
1 - 68
TMhelix
69 - 91
outside
92 - 100
TMhelix
101 - 123
inside
124 - 129
TMhelix
130 - 148
outside
149 - 151
TMhelix
152 - 174
inside
175 - 289
TMhelix
290 - 312
outside
313 - 321
TMhelix
322 - 344
inside
345 - 350
TMhelix
351 - 373
outside
374 - 382
TMhelix
383 - 402
inside
403 - 408
TMhelix
409 - 426
outside
427 - 491
TMhelix
492 - 511
inside
512 - 523
TMhelix
524 - 546
outside
547 - 612
TMhelix
613 - 635
inside
636 - 650
Population Genetic Test Statistics
Pi
268.132969
Theta
203.128772
Tajima's D
1.317097
CLR
0.403903
CSRT
0.748512574371281
Interpretation
Uncertain
