Gene
KWMTBOMO04166
Annotation
PREDICTED:_uncharacterized_protein_LOC105842971_[Bombyx_mori]
Transcription factor
Location in the cell
Nuclear Reliability : 3.348
Sequence
CDS
ATGCCGCGTGTTTATAAACCTGATCCTAGAGGGAAAAGATATAGAAAGTACGATGAAAATATTATAAATGAAGCTGTTGCCGCATATGCGAATGGCAAATTCTCTTTAAAGGCCATAGCAGAGAAATATGATATTGATAAATCTGTACTTTATAGACACAGTGTCAAAACTATGAAAAAACAAGGAGGGCAAACTATTTTAACTAACGAGACTGAAGAATGCATGATAAAGTATATCAATATATGTGCAGAATGGGGATGCCCTTTAGATGCTTTAGATTTGAGATACATTGTTAAGACTTATCTTGACCAAGTGGGTCGGACAGTACTCAAATTTAAAAATAATTTACCAGGCCCCGACTTTGTCGCGAGTTTTCTAAAACGTCACAAAAACCAAATTTCCCAAAGATATAGCCAAAACATTAAGAAAAAAAGGGCAGAAGTTTCACCAAACTTGATTAAAGAATATTTCAAAGAGTTAGAAGTGTCATTGGCAGGTATTAATCCGAGTTCTATAATTAATTACGATGAAACTAATTTGACTGATGACCCAGGACGCAAAAAAATCATTACTAAACGAGGAACGAAGTACCCAGAGAGGGTAATGAATAGTTCAAAGTCTTCAGTTTCATTGATGATAGCCGCTACAGCTGACGGCACTTTACTCCCGCCGTATGTGGTATACAAGGCTCAAAACTTGTATAACACATGGACTGAAAATGGACCACCAGGAGCAAGGTATAATCGTACGCAATCCGGTTGGTTTGATGCAACCATTTTTGAAGATTGGATTAAATCCATCATTATACCTCACTTCAAAGATAAAGTTGGGAAAAAGTGCCTAATAGGCGATAATCTCTCGTCGCATTTATCGATTGAGACAATAAAACTGTGCCACGAACAAAATATATCGTTCATTTTTTTACCGGCAAATAGTACACACCTCACACAACCTCTTGATGTAGCATTCTTCAGGCCTATGAAGATAGCATGGCGAAACGTAGTATTGAAGTGGAAAAAAACCGATGGGAAAGCCCAATCAACGATTCCAAAGGGATGTTTTCCTAGACTTTTAAATAAATTAATGGAAGAGCTTCGTAACAACAGTAAGGCTAATATAATAGCAGGATTCCGGAAAACAGGAATATACCCGATTAATGAAGAAGAGGTTTTAAGCAGACTACCTGAAGTTCAAAACAATGAAAAGAAAACTGAAATAGAAAAATCCGTTCTAGATTTGCTCAGGGAGATGAGATATGGAACCATGAACATAACCGAACCAAAAAGGAAAAAAAAGTTGATAGTAATACCTGGACGAAGCGTTGGTTCTGAAAGTGAAGACTACATAGAAACAGAACAAGAAGAAAGCCAACAAAAACCAAAAAAGGCAGATCTGAGATCCAAAATAACACAAAACAAAGAAAATACTACAACTAATACTTACGATGGCAAAGAAAACAATAAATATAAGGGAAAAGGTGTAGGGAAAAAACTAAAACACAAGATCAGAATATTGCTGAAAAAGAAAATAAACAAGAAAACGGACCAGACACTGAAAAAATACTACTTGACGCAATAA
Protein
MPRVYKPDPRGKRYRKYDENIINEAVAAYANGKFSLKAIAEKYDIDKSVLYRHSVKTMKKQGGQTILTNETEECMIKYINICAEWGCPLDALDLRYIVKTYLDQVGRTVLKFKNNLPGPDFVASFLKRHKNQISQRYSQNIKKKRAEVSPNLIKEYFKELEVSLAGINPSSIINYDETNLTDDPGRKKIITKRGTKYPERVMNSSKSSVSLMIAATADGTLLPPYVVYKAQNLYNTWTENGPPGARYNRTQSGWFDATIFEDWIKSIIIPHFKDKVGKKCLIGDNLSSHLSIETIKLCHEQNISFIFLPANSTHLTQPLDVAFFRPMKIAWRNVVLKWKKTDGKAQSTIPKGCFPRLLNKLMEELRNNSKANIIAGFRKTGIYPINEEEVLSRLPEVQNNEKKTEIEKSVLDLLREMRYGTMNITEPKRKKKLIVIPGRSVGSESEDYIETEQEESQQKPKKADLRSKITQNKENTTTNTYDGKENNKYKGKGVGKKLKHKIRILLKKKINKKTDQTLKKYYLTQ
Summary
Uniprot
H9JBQ2
A0A2S2NMJ8
A0A3Q0J1E8
J9JX79
A0A1S4EG94
J9LWC9
+ More
J9KAI8 X1WST4 A0A1B6GCB9 A0A1B6CLN0 J9K1E0 X1WTH7 J9JPC0 A0A1B6KLJ7 J9M6Q1 B0XDM0 J9KC80 A0A336M092 X1X0I1 X1XEK3 A0A1B6L0P8 X1WP22 A0A2S2R564 J9JNH9 A0A2P8Z814 J9KE52 X1WLE6 A0A1E1WU51 A0A2P8YWM9 A0A2W1BUJ3 J9K4L7 J9K6M6 A0A3S2TIP6 A0A2H8TJC7 A0A1B0GJE9 J9M3V3 J9LW80 A0A1Y1K537 A0A2A4IT23 A0A1Y1KZ51 X1XMI2 J9K5J2 J9KH76 J9L3Y1 X1WL18 J9LJP6 X1XT95 J9JJQ5 A0A1B6JFL2 B3RTS2 A0A1W7R625 A0A2H1V2L8 A0A0L7KQ98 A0A0J7KF01 A0A1B0DJ12 C3ZFC2
J9KAI8 X1WST4 A0A1B6GCB9 A0A1B6CLN0 J9K1E0 X1WTH7 J9JPC0 A0A1B6KLJ7 J9M6Q1 B0XDM0 J9KC80 A0A336M092 X1X0I1 X1XEK3 A0A1B6L0P8 X1WP22 A0A2S2R564 J9JNH9 A0A2P8Z814 J9KE52 X1WLE6 A0A1E1WU51 A0A2P8YWM9 A0A2W1BUJ3 J9K4L7 J9K6M6 A0A3S2TIP6 A0A2H8TJC7 A0A1B0GJE9 J9M3V3 J9LW80 A0A1Y1K537 A0A2A4IT23 A0A1Y1KZ51 X1XMI2 J9K5J2 J9KH76 J9L3Y1 X1WL18 J9LJP6 X1XT95 J9JJQ5 A0A1B6JFL2 B3RTS2 A0A1W7R625 A0A2H1V2L8 A0A0L7KQ98 A0A0J7KF01 A0A1B0DJ12 C3ZFC2
EMBL
BABH01024482
GGMR01005791
MBY18410.1
ABLF02024787
ABLF02036293
ABLF02036295
+ More
ABLF02036300 ABLF02036303 ABLF02036307 ABLF02020969 ABLF02020971 ABLF02006951 GECZ01009826 JAS59943.1 GEDC01023010 JAS14288.1 ABLF02018386 ABLF02041722 ABLF02062703 ABLF02021095 ABLF02021103 ABLF02021105 GEBQ01027923 JAT12054.1 ABLF02004276 ABLF02004277 ABLF02019754 ABLF02043119 DS232767 EDS45530.1 ABLF02031950 UFQS01000373 UFQT01000373 SSX03288.1 SSX23654.1 ABLF02003004 ABLF02053093 ABLF02064790 ABLF02014120 ABLF02031335 GEBQ01022752 JAT17225.1 ABLF02034490 GGMS01015966 MBY85169.1 ABLF02024890 PYGN01000900 PYGN01000153 PSN39546.1 PSN52644.1 ABLF02010048 ABLF02042417 GDQN01000733 JAT90321.1 PYGN01000316 PSN48625.1 KZ149957 PZC76450.1 ABLF02035413 ABLF02012638 RSAL01000111 RVE47096.1 GFXV01002448 MBW14253.1 AJWK01020237 AJWK01020238 ABLF02005267 ABLF02005269 ABLF02032408 ABLF02032409 ABLF02032413 GEZM01096654 GEZM01096605 JAV54775.1 NWSH01008222 PCG62646.1 GEZM01069098 JAV66644.1 ABLF02010036 ABLF02067222 ABLF02002783 ABLF02012639 ABLF02034252 ABLF02010037 ABLF02010039 ABLF02066755 ABLF02034492 ABLF02013132 ABLF02013134 ABLF02013135 ABLF02005217 GECU01010063 JAS97643.1 DS985244 EDV26187.1 GEHC01001021 JAV46624.1 ODYU01000374 SOQ35049.1 JTDY01007086 KOB65467.1 LBMM01008491 KMQ88832.1 AJVK01063592 GG666613 EEN48676.1
ABLF02036300 ABLF02036303 ABLF02036307 ABLF02020969 ABLF02020971 ABLF02006951 GECZ01009826 JAS59943.1 GEDC01023010 JAS14288.1 ABLF02018386 ABLF02041722 ABLF02062703 ABLF02021095 ABLF02021103 ABLF02021105 GEBQ01027923 JAT12054.1 ABLF02004276 ABLF02004277 ABLF02019754 ABLF02043119 DS232767 EDS45530.1 ABLF02031950 UFQS01000373 UFQT01000373 SSX03288.1 SSX23654.1 ABLF02003004 ABLF02053093 ABLF02064790 ABLF02014120 ABLF02031335 GEBQ01022752 JAT17225.1 ABLF02034490 GGMS01015966 MBY85169.1 ABLF02024890 PYGN01000900 PYGN01000153 PSN39546.1 PSN52644.1 ABLF02010048 ABLF02042417 GDQN01000733 JAT90321.1 PYGN01000316 PSN48625.1 KZ149957 PZC76450.1 ABLF02035413 ABLF02012638 RSAL01000111 RVE47096.1 GFXV01002448 MBW14253.1 AJWK01020237 AJWK01020238 ABLF02005267 ABLF02005269 ABLF02032408 ABLF02032409 ABLF02032413 GEZM01096654 GEZM01096605 JAV54775.1 NWSH01008222 PCG62646.1 GEZM01069098 JAV66644.1 ABLF02010036 ABLF02067222 ABLF02002783 ABLF02012639 ABLF02034252 ABLF02010037 ABLF02010039 ABLF02066755 ABLF02034492 ABLF02013132 ABLF02013134 ABLF02013135 ABLF02005217 GECU01010063 JAS97643.1 DS985244 EDV26187.1 GEHC01001021 JAV46624.1 ODYU01000374 SOQ35049.1 JTDY01007086 KOB65467.1 LBMM01008491 KMQ88832.1 AJVK01063592 GG666613 EEN48676.1
Proteomes
PRIDE
Pfam
Interpro
IPR004875
DDE_SF_endonuclease_dom
+ More
IPR007889 HTH_Psq
IPR009057 Homeobox-like_sf
IPR029526 PGBD
IPR012337 RNaseH-like_sf
IPR001584 Integrase_cat-core
IPR036397 RNaseH_sf
IPR011335 Restrct_endonuc-II-like
IPR011604 Exonuc_phg/RecB_C
IPR008042 Retrotrans_Pao
IPR006600 HTH_CenpB_DNA-bd_dom
IPR007021 DUF659
IPR008906 HATC_C_dom
IPR006579 Pre_C2HC_dom
IPR000477 RT_dom
IPR027124 Swc5/CFDP2
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR036875 Znf_CCHC_sf
IPR001878 Znf_CCHC
IPR007588 Znf_FLYWCH
IPR007889 HTH_Psq
IPR009057 Homeobox-like_sf
IPR029526 PGBD
IPR012337 RNaseH-like_sf
IPR001584 Integrase_cat-core
IPR036397 RNaseH_sf
IPR011335 Restrct_endonuc-II-like
IPR011604 Exonuc_phg/RecB_C
IPR008042 Retrotrans_Pao
IPR006600 HTH_CenpB_DNA-bd_dom
IPR007021 DUF659
IPR008906 HATC_C_dom
IPR006579 Pre_C2HC_dom
IPR000477 RT_dom
IPR027124 Swc5/CFDP2
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR036875 Znf_CCHC_sf
IPR001878 Znf_CCHC
IPR007588 Znf_FLYWCH
SUPFAM
Gene 3D
ProteinModelPortal
H9JBQ2
A0A2S2NMJ8
A0A3Q0J1E8
J9JX79
A0A1S4EG94
J9LWC9
+ More
J9KAI8 X1WST4 A0A1B6GCB9 A0A1B6CLN0 J9K1E0 X1WTH7 J9JPC0 A0A1B6KLJ7 J9M6Q1 B0XDM0 J9KC80 A0A336M092 X1X0I1 X1XEK3 A0A1B6L0P8 X1WP22 A0A2S2R564 J9JNH9 A0A2P8Z814 J9KE52 X1WLE6 A0A1E1WU51 A0A2P8YWM9 A0A2W1BUJ3 J9K4L7 J9K6M6 A0A3S2TIP6 A0A2H8TJC7 A0A1B0GJE9 J9M3V3 J9LW80 A0A1Y1K537 A0A2A4IT23 A0A1Y1KZ51 X1XMI2 J9K5J2 J9KH76 J9L3Y1 X1WL18 J9LJP6 X1XT95 J9JJQ5 A0A1B6JFL2 B3RTS2 A0A1W7R625 A0A2H1V2L8 A0A0L7KQ98 A0A0J7KF01 A0A1B0DJ12 C3ZFC2
J9KAI8 X1WST4 A0A1B6GCB9 A0A1B6CLN0 J9K1E0 X1WTH7 J9JPC0 A0A1B6KLJ7 J9M6Q1 B0XDM0 J9KC80 A0A336M092 X1X0I1 X1XEK3 A0A1B6L0P8 X1WP22 A0A2S2R564 J9JNH9 A0A2P8Z814 J9KE52 X1WLE6 A0A1E1WU51 A0A2P8YWM9 A0A2W1BUJ3 J9K4L7 J9K6M6 A0A3S2TIP6 A0A2H8TJC7 A0A1B0GJE9 J9M3V3 J9LW80 A0A1Y1K537 A0A2A4IT23 A0A1Y1KZ51 X1XMI2 J9K5J2 J9KH76 J9L3Y1 X1WL18 J9LJP6 X1XT95 J9JJQ5 A0A1B6JFL2 B3RTS2 A0A1W7R625 A0A2H1V2L8 A0A0L7KQ98 A0A0J7KF01 A0A1B0DJ12 C3ZFC2
Ontologies
GO
PANTHER
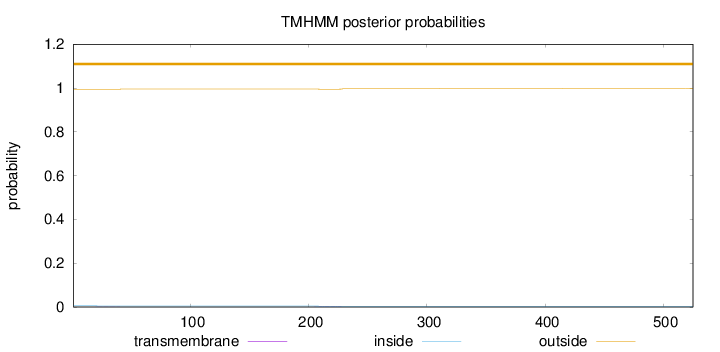
Topology
Subcellular location
Nucleus
Length:
525
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.05841
Exp number, first 60 AAs:
0.01874
Total prob of N-in:
0.00575
outside
1 - 525
Population Genetic Test Statistics
Pi
195.353978
Theta
180.262279
Tajima's D
0.287348
CLR
0.315766
CSRT
0.444477776111194
Interpretation
Uncertain
