Gene
KWMTBOMO04054
Annotation
retroelement_polyprotein_[Glyptapanteles_flavicoxis]
Location in the cell
Nuclear Reliability : 2.527
Sequence
CDS
ATGCCACATGTTAATCAAGTTCAGTGTATAAATCCAATTTTCGATAAACTGGATTGTGATCTCACAAATGAAGAAGATATCGCAAAATTACGCACCCTTCTAAACAAATATCAGCACCTGTTTATCAGAGGGTATCCAACAACTCGTGTCAAGACAGGTGAACTGGAAATCCGCCTGAAGAATCCAAACAAATATGTAGAACGAAGACCTTACAGGCTCAGTCCAATCGAACGAGAAAGGGTGCGAGCCATTGTGAAAGAGTTGATGGAGCATGGCATTATACGCGAAAGCAAGTCCCCATACTCCAGTCCCATCATCTTGGTGAAGAAGAAGAATGGTGACGATCGCATGTGCGTGGATTTCAGAGAACTTAACTCTAATACCTTACGGGACCATTACCCGCTGCCACTAATTTCAGACCAAATTGACCAATTGGCCAAGGGGCAATTCTTTACAACCTTTGACATGGCCGCAGGGTTTCACCAAATGCCTATTGCAGAATCATCAATTGAAAAGACAGCCTTTGTCACACCGGATGGGCTGTATGAATATACGACTATGCCATTTGGATTAAGTAATGCATGTTCAGTCTATCAACGATGCATCAACAGAGCTCTGAGCTCGTTAATAGGGAATGCAGCTCAAGTCTACGTCGACGACGTATTAAGCAGATGTAAAAATATCGCAGAAGGGCTGTCAAACATAGAACGTATTCTAATAGCACTACAAGATGCGGGCTTTTCCATTAATGCTGATAAATGCAGTTTTTTTAAGCGTTCAATCGAATATTTGGGTAACGTAGTATCTAATGGACAAGTCTGGCCAAGCCCACGGAAAATAGATGCTCTGGTAAAATCGCCTGTACCAACAACACCCAGACAAGTAAGACAATTTTGTGGATTGGCTGGCTACTTTAGAAAATTTATAGCAAACTTCTCACGCATAATGATACCACTATACGAGTTGACCAAATCAGGGGCTAAATGGGAGTGGGACGAGCGCCATGAGAAAGCAAGAGAGAATGTCATTCAGTGCTTAACATCAACTCCAGTACTAACACTATTCCAGGAGGAAGCACCTATTCAATTGTACACAGATGCAAGTAGTTTGGGATTCGGAGCAGTACTGGTACAAGTAATTGCGGGACGACAACATGCAGTTGCCTTTATGAGCATGAGAACTACTGAAACAGAAAGTCGCTATCACTCTTATGAATTAGAGACCTTGGCAGTAGTCCGAGCGATAAAACACTTCCGGCAGTATCTTTATGGACGTAAGTTTACAGTTATCACAGACTGTAATGCACTAAAAGCATCCAAACACAAAAAGGACTTACTCCCTAGAATCCACCGTTGGTGGGCATTTCTACAAAACTATAATTTTGAGGTTGAATACCGCAAGGGTCAACAGCTTCAACACGCAGACTACTTCAGTAGGAACCCAGTCGAACTCACAGTGAATATAATGACCAGAGATCTAGACTGGCTGAAAATTGAACAGCGACGTGATGAAACACTACGAGCTATAATGAATGGTATGAGTAATAACAATCCAAGCAACGGCTATATGCTAGAAAACAATGTCCTCAAGCGTATCGTACAAGATCCAACCTTCGGACACCGATACTGCACAGTGGTACCCAGGTCCTTTCAGTGGAGTTTAATAAACTCCTTCCACACCGCACTGAAACACCCTGGCTGGGAAAAGACGTTGCAGAAAATTAAGGACAGCCACTGGTTTGACAACATGAGCACGAAGGTACGAAAATTTGTTGACAATTGCGTGGTCTGCAGGACATCGAAAGGAGCATCTGGTGCGGTACAAGCTCAAATGCACCCGATTCAGAAACCGTCTGCTACTTTTCAAGTAATACACATGGATATAACAGGCAAGATGGGAACATCCAACGACCAACAGTATGTGATTATCACCATAGATGCTTTTACAAAATACGTGCTATTCTATTATGCAACTAACAAAAATCCACCCAGCACATTAGCTGCCCTAAAGCGTACGGTTCATCTATTCGGTACTCCTATCCAAATAATAGTAGACGGTGGAAGAGAGTTCCTCGGTGAATTCAAGACCTACTGCGATAGGGTAGGTATTGACATTCATGCTATAGCACCAGGAGTCAGCCGACCAAATGGACAAGTAGAACGCGTCGTAGCTACACTAAAAAATACTCTTACTATGATCAAGAATTATGAAACAGAAGAGTGGCACACAACGATTGCAGAACTTCAGCTAGCAATAAATTGCACTCCGCATCGTGTGACAGGCGTGGCTCCACTTACATTACTCACTCAGCGAAAGCATTGTGTTCCTCCAGAGCTACTCAATTTAGTAAATATTGACGAGCAGACTATTGATATTGAAGCACTCACTAACCATGTTCAACACAAGATGTCACAAAATGCAGAACAGGACAGGCAACGTTTCAATTCAAAAAAAGCCAGGATCCATCATTTCCAGCGTGGTGATTATGTGCTCATAAAAAATAATCCTCGTAATCAAACTTCGTTGGATCTCAAATTCAGCGAGCCATATGAGATAACAAGAATCCTGGACAACAACCGCTATTTAGTAAAAAAGGTGGTTGGTTCAGGGCGTTCAAGAAAAGTAGCTCATCATCAACTGCGTCGGGCACCGCAACCTGGAGACCAGATAGCCGTATCGGCGGAAGAAGACGACCCTCCAGCATCAGCCGATGAACCTATATGA
Protein
MPHVNQVQCINPIFDKLDCDLTNEEDIAKLRTLLNKYQHLFIRGYPTTRVKTGELEIRLKNPNKYVERRPYRLSPIERERVRAIVKELMEHGIIRESKSPYSSPIILVKKKNGDDRMCVDFRELNSNTLRDHYPLPLISDQIDQLAKGQFFTTFDMAAGFHQMPIAESSIEKTAFVTPDGLYEYTTMPFGLSNACSVYQRCINRALSSLIGNAAQVYVDDVLSRCKNIAEGLSNIERILIALQDAGFSINADKCSFFKRSIEYLGNVVSNGQVWPSPRKIDALVKSPVPTTPRQVRQFCGLAGYFRKFIANFSRIMIPLYELTKSGAKWEWDERHEKARENVIQCLTSTPVLTLFQEEAPIQLYTDASSLGFGAVLVQVIAGRQHAVAFMSMRTTETESRYHSYELETLAVVRAIKHFRQYLYGRKFTVITDCNALKASKHKKDLLPRIHRWWAFLQNYNFEVEYRKGQQLQHADYFSRNPVELTVNIMTRDLDWLKIEQRRDETLRAIMNGMSNNNPSNGYMLENNVLKRIVQDPTFGHRYCTVVPRSFQWSLINSFHTALKHPGWEKTLQKIKDSHWFDNMSTKVRKFVDNCVVCRTSKGASGAVQAQMHPIQKPSATFQVIHMDITGKMGTSNDQQYVIITIDAFTKYVLFYYATNKNPPSTLAALKRTVHLFGTPIQIIVDGGREFLGEFKTYCDRVGIDIHAIAPGVSRPNGQVERVVATLKNTLTMIKNYETEEWHTTIAELQLAINCTPHRVTGVAPLTLLTQRKHCVPPELLNLVNIDEQTIDIEALTNHVQHKMSQNAEQDRQRFNSKKARIHHFQRGDYVLIKNNPRNQTSLDLKFSEPYEITRILDNNRYLVKKVVGSGRSRKVAHHQLRRAPQPGDQIAVSAEEDDPPASADEPI
Summary
Uniprot
B7S8P8
A0A0A9Z954
A0A0A9WJJ8
A0A146KXF1
A0A146M2J2
A0A0A9X1Z8
+ More
A0A146LDG4 A0A0A9YM49 A0A0A9YEP8 Q24310 D2A5M1 A0A0A9X5S1 A0A0C9RXL6 A0A0J7KEC9 A0A1Y1NG42 A0A1Y1NAV9 A0A139WJA1 A0A139WIQ0 A0A194R1L3 A0A147BLC5 A0A3S2L135 A0A1Y1KFN9 A0A1Y1KFM9 A0A139WIB9 X1X5H2 A0A0A9XL77 A0A0A9YZ15 A0A2S2P032 A0A0A9ZDJ9 X1WPP0 A0A1Y1MFA9 A0A0J7K9X8 A0A0J7NF11 A0A1Y1NEJ5 A0A023EZG2 A0A023EYH9 A0A0J7KLK4 A0A0J7K890 X1XM84 A0A2S2QIS3 A0A139WCC5 A0A139WCG2 A0A2L2Y402 A0A2L2Y1I8 A0A2L2Y1Z9 A0A023EY26 A0A2S2PFS2 J9JJ45 J9LBS9 A0A2H8TRV6 A0A1Y1MFE0 A0A1W7R6L8 A0A2S2NFQ4 A0A0A1XB16 A0A2S2NKJ2 J9JZ77 A0A1W7R6G2 A0A034WRX5 T1PMR9 X1XU22 A0A224XHR1 A0A1Y1LN89 A0A034VAM6 X1XD76 A0A2A4J7U9 A0A034VKV9 X1XU14 Q94885 A0A1W7R6H1 A0A2S2PGQ9 W8APB2 A0A164P0I2 A0A034VPR8 A0A0A1WKC8 A0A2M3Z5R3 W8B0V9 W8C8Q1 A0A023EYF0 X1X261 A0A2S2QVZ0 J9JQ65 A0A2S2PD17 A0A023F013 A0A2S2N9B2 A0A1W7R6G1 A0A034W795 A0A2M4CKM2
A0A146LDG4 A0A0A9YM49 A0A0A9YEP8 Q24310 D2A5M1 A0A0A9X5S1 A0A0C9RXL6 A0A0J7KEC9 A0A1Y1NG42 A0A1Y1NAV9 A0A139WJA1 A0A139WIQ0 A0A194R1L3 A0A147BLC5 A0A3S2L135 A0A1Y1KFN9 A0A1Y1KFM9 A0A139WIB9 X1X5H2 A0A0A9XL77 A0A0A9YZ15 A0A2S2P032 A0A0A9ZDJ9 X1WPP0 A0A1Y1MFA9 A0A0J7K9X8 A0A0J7NF11 A0A1Y1NEJ5 A0A023EZG2 A0A023EYH9 A0A0J7KLK4 A0A0J7K890 X1XM84 A0A2S2QIS3 A0A139WCC5 A0A139WCG2 A0A2L2Y402 A0A2L2Y1I8 A0A2L2Y1Z9 A0A023EY26 A0A2S2PFS2 J9JJ45 J9LBS9 A0A2H8TRV6 A0A1Y1MFE0 A0A1W7R6L8 A0A2S2NFQ4 A0A0A1XB16 A0A2S2NKJ2 J9JZ77 A0A1W7R6G2 A0A034WRX5 T1PMR9 X1XU22 A0A224XHR1 A0A1Y1LN89 A0A034VAM6 X1XD76 A0A2A4J7U9 A0A034VKV9 X1XU14 Q94885 A0A1W7R6H1 A0A2S2PGQ9 W8APB2 A0A164P0I2 A0A034VPR8 A0A0A1WKC8 A0A2M3Z5R3 W8B0V9 W8C8Q1 A0A023EYF0 X1X261 A0A2S2QVZ0 J9JQ65 A0A2S2PD17 A0A023F013 A0A2S2N9B2 A0A1W7R6G1 A0A034W795 A0A2M4CKM2
Pubmed
EMBL
EF710649
ACE75273.1
GBHO01040010
GBHO01001812
JAG03594.1
JAG41792.1
+ More
GBHO01036023 GBHO01032007 JAG07581.1 JAG11597.1 GDHC01018907 JAP99721.1 GDHC01004980 JAQ13649.1 GBHO01030786 JAG12818.1 GDHC01012341 JAQ06288.1 GBHO01011083 GBHO01009497 JAG32521.1 JAG34107.1 GBHO01015604 JAG28000.1 X14037 CAA32198.1 KQ971345 EFA05054.2 GBHO01031154 JAG12450.1 GBYB01013920 JAG83687.1 LBMM01008871 KMQ88569.1 GEZM01007614 GEZM01007613 GEZM01007612 JAV95056.1 GEZM01007616 GEZM01007615 GEZM01007611 JAV95051.1 KQ971338 KYB27887.1 KYB27888.1 KQ460883 KPJ11409.1 GEGO01004299 JAR91105.1 RSAL01000274 RVE43148.1 GEZM01089268 GEZM01089261 JAV57577.1 GEZM01089269 GEZM01089265 JAV57567.1 KQ971339 KYB27733.1 ABLF02004401 ABLF02013544 ABLF02013550 GBHO01043544 GBHO01031946 GBHO01031931 GBHO01022910 GBHO01022909 GBHO01015498 GBHO01001135 GBHO01001106 JAG00060.1 JAG11658.1 JAG11673.1 JAG20694.1 JAG20695.1 JAG28106.1 JAG42469.1 JAG42498.1 GBHO01022918 GBHO01006165 GBHO01006163 GBHO01001110 JAG20686.1 JAG37439.1 JAG37441.1 JAG42494.1 GGMR01009949 MBY22568.1 GBHO01041008 GBHO01006162 GBHO01001134 GBHO01001105 JAG02596.1 JAG37442.1 JAG42470.1 JAG42499.1 ABLF02033943 ABLF02042737 GEZM01033083 JAV84424.1 LBMM01010976 KMQ87102.1 LBMM01005919 KMQ91095.1 GEZM01005427 JAV96179.1 GBBI01004551 JAC14161.1 GBBI01004550 JAC14162.1 KMQ91096.1 LBMM01011782 KMQ86633.1 ABLF02004526 ABLF02054969 GGMS01008442 MBY77645.1 KQ971370 KYB25609.1 KYB25610.1 IAAA01004057 LAA02040.1 IAAA01004058 LAA02042.1 IAAA01004056 LAA02037.1 GBBI01004477 JAC14235.1 GGMR01015682 MBY28301.1 ABLF02008401 ABLF02008404 ABLF02041876 GFXV01005120 MBW16925.1 GEZM01034557 JAV83728.1 GEHC01000930 JAV46715.1 GGMR01003388 MBY16007.1 GBXI01005773 JAD08519.1 GGMR01005091 MBY17710.1 ABLF02014660 ABLF02042774 GEHC01000888 JAV46757.1 GAKP01001563 JAC57389.1 KA649989 AFP64618.1 ABLF02017401 GFTR01008753 JAW07673.1 GEZM01051217 GEZM01051215 JAV75083.1 GAKP01019383 JAC39569.1 ABLF02022032 NWSH01002797 PCG67520.1 GAKP01016235 JAC42717.1 ABLF02003120 ABLF02004015 ABLF02057179 ABLF02066208 X95908 CAA65152.1 GEHC01000876 JAV46769.1 GGMR01016004 MBY28623.1 GAMC01020207 JAB86348.1 LRGB01002748 KZS06406.1 GAKP01014483 JAC44469.1 GBXI01015151 JAC99140.1 GGFM01003105 MBW23856.1 GAMC01015907 JAB90648.1 GAMC01007326 JAB99229.1 GBBI01004549 JAC14163.1 ABLF02005127 ABLF02005129 ABLF02007401 GGMS01012726 MBY81929.1 ABLF02005846 GGMR01014665 MBY27284.1 GBBI01004613 JAC14099.1 GGMR01000717 MBY13336.1 GEHC01000891 JAV46754.1 GAKP01008942 JAC50010.1 GGFL01001689 MBW65867.1
GBHO01036023 GBHO01032007 JAG07581.1 JAG11597.1 GDHC01018907 JAP99721.1 GDHC01004980 JAQ13649.1 GBHO01030786 JAG12818.1 GDHC01012341 JAQ06288.1 GBHO01011083 GBHO01009497 JAG32521.1 JAG34107.1 GBHO01015604 JAG28000.1 X14037 CAA32198.1 KQ971345 EFA05054.2 GBHO01031154 JAG12450.1 GBYB01013920 JAG83687.1 LBMM01008871 KMQ88569.1 GEZM01007614 GEZM01007613 GEZM01007612 JAV95056.1 GEZM01007616 GEZM01007615 GEZM01007611 JAV95051.1 KQ971338 KYB27887.1 KYB27888.1 KQ460883 KPJ11409.1 GEGO01004299 JAR91105.1 RSAL01000274 RVE43148.1 GEZM01089268 GEZM01089261 JAV57577.1 GEZM01089269 GEZM01089265 JAV57567.1 KQ971339 KYB27733.1 ABLF02004401 ABLF02013544 ABLF02013550 GBHO01043544 GBHO01031946 GBHO01031931 GBHO01022910 GBHO01022909 GBHO01015498 GBHO01001135 GBHO01001106 JAG00060.1 JAG11658.1 JAG11673.1 JAG20694.1 JAG20695.1 JAG28106.1 JAG42469.1 JAG42498.1 GBHO01022918 GBHO01006165 GBHO01006163 GBHO01001110 JAG20686.1 JAG37439.1 JAG37441.1 JAG42494.1 GGMR01009949 MBY22568.1 GBHO01041008 GBHO01006162 GBHO01001134 GBHO01001105 JAG02596.1 JAG37442.1 JAG42470.1 JAG42499.1 ABLF02033943 ABLF02042737 GEZM01033083 JAV84424.1 LBMM01010976 KMQ87102.1 LBMM01005919 KMQ91095.1 GEZM01005427 JAV96179.1 GBBI01004551 JAC14161.1 GBBI01004550 JAC14162.1 KMQ91096.1 LBMM01011782 KMQ86633.1 ABLF02004526 ABLF02054969 GGMS01008442 MBY77645.1 KQ971370 KYB25609.1 KYB25610.1 IAAA01004057 LAA02040.1 IAAA01004058 LAA02042.1 IAAA01004056 LAA02037.1 GBBI01004477 JAC14235.1 GGMR01015682 MBY28301.1 ABLF02008401 ABLF02008404 ABLF02041876 GFXV01005120 MBW16925.1 GEZM01034557 JAV83728.1 GEHC01000930 JAV46715.1 GGMR01003388 MBY16007.1 GBXI01005773 JAD08519.1 GGMR01005091 MBY17710.1 ABLF02014660 ABLF02042774 GEHC01000888 JAV46757.1 GAKP01001563 JAC57389.1 KA649989 AFP64618.1 ABLF02017401 GFTR01008753 JAW07673.1 GEZM01051217 GEZM01051215 JAV75083.1 GAKP01019383 JAC39569.1 ABLF02022032 NWSH01002797 PCG67520.1 GAKP01016235 JAC42717.1 ABLF02003120 ABLF02004015 ABLF02057179 ABLF02066208 X95908 CAA65152.1 GEHC01000876 JAV46769.1 GGMR01016004 MBY28623.1 GAMC01020207 JAB86348.1 LRGB01002748 KZS06406.1 GAKP01014483 JAC44469.1 GBXI01015151 JAC99140.1 GGFM01003105 MBW23856.1 GAMC01015907 JAB90648.1 GAMC01007326 JAB99229.1 GBBI01004549 JAC14163.1 ABLF02005127 ABLF02005129 ABLF02007401 GGMS01012726 MBY81929.1 ABLF02005846 GGMR01014665 MBY27284.1 GBBI01004613 JAC14099.1 GGMR01000717 MBY13336.1 GEHC01000891 JAV46754.1 GAKP01008942 JAC50010.1 GGFL01001689 MBW65867.1
Proteomes
Pfam
Interpro
IPR041577
RT_RNaseH_2
+ More
IPR001584 Integrase_cat-core
IPR036875 Znf_CCHC_sf
IPR012337 RNaseH-like_sf
IPR001878 Znf_CCHC
IPR021109 Peptidase_aspartic_dom_sf
IPR000477 RT_dom
IPR036397 RNaseH_sf
IPR041588 Integrase_H2C2
IPR041373 RT_RNaseH
IPR001969 Aspartic_peptidase_AS
IPR001995 Peptidase_A2_cat
IPR018061 Retropepsins
IPR034122 Retropepsin-like_bacterial
IPR025836 Zn_knuckle_CX2CX4HX4C
IPR005162 Retrotrans_gag_dom
IPR036361 SAP_dom_sf
IPR003034 SAP_dom
IPR004955 Baculovirus_Gp64
IPR006579 Pre_C2HC_dom
IPR028002 Myb_DNA-bind_5
IPR019103 Peptidase_aspartic_DDI1-type
IPR024650 Peptidase_A2B
IPR032071 DUF4806
IPR001584 Integrase_cat-core
IPR036875 Znf_CCHC_sf
IPR012337 RNaseH-like_sf
IPR001878 Znf_CCHC
IPR021109 Peptidase_aspartic_dom_sf
IPR000477 RT_dom
IPR036397 RNaseH_sf
IPR041588 Integrase_H2C2
IPR041373 RT_RNaseH
IPR001969 Aspartic_peptidase_AS
IPR001995 Peptidase_A2_cat
IPR018061 Retropepsins
IPR034122 Retropepsin-like_bacterial
IPR025836 Zn_knuckle_CX2CX4HX4C
IPR005162 Retrotrans_gag_dom
IPR036361 SAP_dom_sf
IPR003034 SAP_dom
IPR004955 Baculovirus_Gp64
IPR006579 Pre_C2HC_dom
IPR028002 Myb_DNA-bind_5
IPR019103 Peptidase_aspartic_DDI1-type
IPR024650 Peptidase_A2B
IPR032071 DUF4806
Gene 3D
ProteinModelPortal
B7S8P8
A0A0A9Z954
A0A0A9WJJ8
A0A146KXF1
A0A146M2J2
A0A0A9X1Z8
+ More
A0A146LDG4 A0A0A9YM49 A0A0A9YEP8 Q24310 D2A5M1 A0A0A9X5S1 A0A0C9RXL6 A0A0J7KEC9 A0A1Y1NG42 A0A1Y1NAV9 A0A139WJA1 A0A139WIQ0 A0A194R1L3 A0A147BLC5 A0A3S2L135 A0A1Y1KFN9 A0A1Y1KFM9 A0A139WIB9 X1X5H2 A0A0A9XL77 A0A0A9YZ15 A0A2S2P032 A0A0A9ZDJ9 X1WPP0 A0A1Y1MFA9 A0A0J7K9X8 A0A0J7NF11 A0A1Y1NEJ5 A0A023EZG2 A0A023EYH9 A0A0J7KLK4 A0A0J7K890 X1XM84 A0A2S2QIS3 A0A139WCC5 A0A139WCG2 A0A2L2Y402 A0A2L2Y1I8 A0A2L2Y1Z9 A0A023EY26 A0A2S2PFS2 J9JJ45 J9LBS9 A0A2H8TRV6 A0A1Y1MFE0 A0A1W7R6L8 A0A2S2NFQ4 A0A0A1XB16 A0A2S2NKJ2 J9JZ77 A0A1W7R6G2 A0A034WRX5 T1PMR9 X1XU22 A0A224XHR1 A0A1Y1LN89 A0A034VAM6 X1XD76 A0A2A4J7U9 A0A034VKV9 X1XU14 Q94885 A0A1W7R6H1 A0A2S2PGQ9 W8APB2 A0A164P0I2 A0A034VPR8 A0A0A1WKC8 A0A2M3Z5R3 W8B0V9 W8C8Q1 A0A023EYF0 X1X261 A0A2S2QVZ0 J9JQ65 A0A2S2PD17 A0A023F013 A0A2S2N9B2 A0A1W7R6G1 A0A034W795 A0A2M4CKM2
A0A146LDG4 A0A0A9YM49 A0A0A9YEP8 Q24310 D2A5M1 A0A0A9X5S1 A0A0C9RXL6 A0A0J7KEC9 A0A1Y1NG42 A0A1Y1NAV9 A0A139WJA1 A0A139WIQ0 A0A194R1L3 A0A147BLC5 A0A3S2L135 A0A1Y1KFN9 A0A1Y1KFM9 A0A139WIB9 X1X5H2 A0A0A9XL77 A0A0A9YZ15 A0A2S2P032 A0A0A9ZDJ9 X1WPP0 A0A1Y1MFA9 A0A0J7K9X8 A0A0J7NF11 A0A1Y1NEJ5 A0A023EZG2 A0A023EYH9 A0A0J7KLK4 A0A0J7K890 X1XM84 A0A2S2QIS3 A0A139WCC5 A0A139WCG2 A0A2L2Y402 A0A2L2Y1I8 A0A2L2Y1Z9 A0A023EY26 A0A2S2PFS2 J9JJ45 J9LBS9 A0A2H8TRV6 A0A1Y1MFE0 A0A1W7R6L8 A0A2S2NFQ4 A0A0A1XB16 A0A2S2NKJ2 J9JZ77 A0A1W7R6G2 A0A034WRX5 T1PMR9 X1XU22 A0A224XHR1 A0A1Y1LN89 A0A034VAM6 X1XD76 A0A2A4J7U9 A0A034VKV9 X1XU14 Q94885 A0A1W7R6H1 A0A2S2PGQ9 W8APB2 A0A164P0I2 A0A034VPR8 A0A0A1WKC8 A0A2M3Z5R3 W8B0V9 W8C8Q1 A0A023EYF0 X1X261 A0A2S2QVZ0 J9JQ65 A0A2S2PD17 A0A023F013 A0A2S2N9B2 A0A1W7R6G1 A0A034W795 A0A2M4CKM2
PDB
4OL8
E-value=2.54414e-69,
Score=669
Ontologies
KEGG
GO
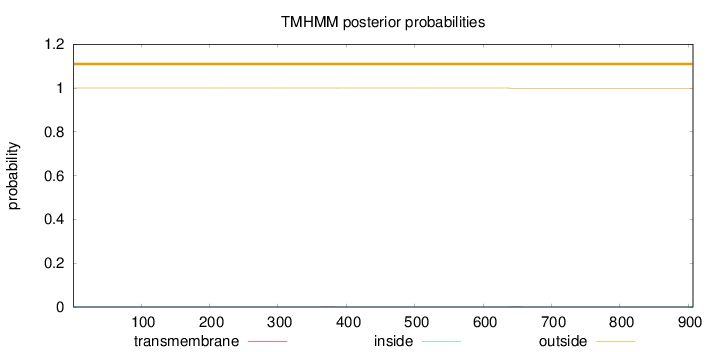
Topology
Length:
907
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.01823
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00034
outside
1 - 907
Population Genetic Test Statistics
Pi
377.617507
Theta
220.41816
Tajima's D
3.478683
CLR
28.534411
CSRT
0.993150342482876
Interpretation
Uncertain
