Pre Gene Modal
BGIBMGA010289
Annotation
UDP-glucosyltransferase_precursor_[Bombyx_mori]
Location in the cell
PlasmaMembrane Reliability : 2.677
Sequence
CDS
ATGATGATACGAAGACTCACAATTGCAATAGCTGTTTGCTTCTGTTTGGGTGTCGATGCGTACAAGATACTGACAGTGTTTCCTGTGCCGGGAAGAAGCCATGGAATCCTTGGAGATGCTGTAGTCAGACATCTATTGGAAGCTGGCCACGAGGTGACACACATAACGCCGTTTCCGAAGAAGGAGCCGCCGCCGAATTTGGTTCAAATAGACGTTGCGGCAAACAAGGCGGCGTTTAATGAGGACTACATAGATATAAAGGCCTTGATGACAAAGGAATTTAACCTTAAAGACAAGAATGTACTATTTTCATTGATGAACAACATAAGCAGCTCCACTATTCTGAACGAGAATGTTCAGAGGCTACTGAGGGATCAGAGTCGTGAGCAGTTCGACGTGATCATAGCCGAATGGATGTTCAGTGATCTATACGCTTCATTCCACGCAGTTCTGGATTGCCCTTTGATTTGGTTCTCAACGATCGAACCTCATTGGATGGTTCTCAGATTGATCGATGAATATCCAAACCCAGCGTACACCTCACACTTTCAGGACAGCTTCGAAGTACCATTTACGTTCGTGGAAAGGATGTCTGTACTATCATCACAACTTACGTGGAGCTTAAGTTTAAATACGTGGGTATACGATCTTGAAAAGTATATATACGACAACAATATTGCTCCTATAATAAAAAAGAACGGTAAACCAGTACCTAACTATGACGAAGTGAGATACAATGGATCTTTACTGCTCGGTAATTCTCATGTCTCGTTGGGAGATGCGATTAAGGTACCGATTAATTACAAAGCCATTGGTGGTTATCATATTGATGGAAAAGTAAAAGAATTGCCACCGGATCTACAAAAAATAATGAACGAATCAAAACACGGCGTCATTTACTTCAGTATGGGCTCGAATTTGAAGAGCAAAGATCTACCGAAGGAGATTAAAGAGGGACTACTGAAAATGTTTAGCCAATTAAAACAGACGGTTTTATGGAAGTTTGAGGAAAACTTATCACCTTTACCGGAAAACGTTCATTTATTGAAATGGGCACCTCAACAGAGTATCCTGGCACACCCGAACTGCATTCTTTTCATAACTCACGGAGGTCTTCTATCGACAACAGAGGCTGTACATTTTGGTAAGCCCATCATCGGCATACCGGTCTTTGCCGATCAGTTTGGTAACGTCAACAGAGCAGTCCAGAAAGGTATTGCGAGGAGAGTGGATTTATCGTTCACGATGGTCCGTGATCTGGAGGAAGCTGTTGCGGAAATGATCAATAATTCTAGGTACATAGAAAAAATAAAGGAGTTATCTTTGATATATCACGATCGTCCGGTGTCTCCGGGTGCTGAGCTGGTCCACTGGGTGGAGCACGTCGTCAAGACTAAAGGAGCTCTCCATCTGCGATCACCAGCACTGCACGTGCCTTTCTACCAGAAGCTGTACTTAGATCTGCTCGCTATTGTTCTGGTCACTTTATTCTCGTTTGTATTAACATGTGATTGTTATAAAATTTTGATTGTATTTACGACTCCTATGAAAAGTCATAATATACTTGGAGAGGCTGCAGCGGAGTTATTGTTGAATGCTGGCCACGAGGTCACATACGTGACGCCCTTCCCAAAAGAATCTGTTCCCGATAAGATGCGCCAGGTAGATGTAACTTACATCGGTAAAATAGGTTTGTTTGATTTAAAGGGGTATCTTAAAAATGACACTGTTAAACCAATGTCTCTACGTAAAATATCATACTTCATGCATGACGTGAACGTAAAAGCCATACAGAATGAGAATCTACAGCAACTTTTAAACGACCCTAGTCAACGATTTGACGCAGTCATCGTCGATTGGTTGTTCAGTGAAATTTTCGTAGGGCTAGCATCATTGTATGATTGTCCATTGATTTGGATGTCGACTATGGATCCGCACTGGCAAATCTTGAGACTGGTAGACGAGATGCCCAATCCTGTATTTCTGGGCCGTTGTTTCCTTGATAGAATTGTGCCATTTCGTTTTTGGGAACGGACCCAAGAGTTACTTTATCAGATCTCATCGTTGTTCTTTAAGGATATAGAGTTCTTTAGTGAAGAGGATGCTGCGTTTAAAAGATTGCTCGGTCCCGTATTCGCTAGAAAGAAGAAACCCCTACCTTCATTTAATGCAGTGAGATACAATGCTTCGTTAGTGCTGAGTAATTCACACCATTCAATAGGTTACCCTGTAAAACTGCCGCCCAATTTTATATCAATCGGAGGTTTTTTTATCGATGACAAGAAACAGCGTCTGTCTTTGGATTTACAAACAATAATGGACAATGCCAAGCACGGTGTCATATTATTTAGCTTAGGGAGCAATTTGAAAAGTAAGGATATGCCGGAACATCTCGTTCGAAGCCTTTTAAATGTATTCAGCGAACTGAAACAAATAGTTATCTGGAAGGTCGAAGAACAAATAGCTGATCTACCTCAAAATGTTCATGTTTTGAAATGGTTGCCGCAGCAGAGTATTTTAGCTCATTCAAACTGTATTCTCTTCATTACCCACGGAGGATTGTTGTCGATTACCGAAGCATATCATCACGGGGTACCTTTAATAGGAATTCCGGTGTTCGCTGATCAATTTAAGAACGTTAATCTGGTTTCAAAGAAGGGCTTCGCGAAGAAAGTTGATCTCACCTACAATTTGCCGGGTGACTTAAAACATGCAATCAATGAAATTTTACATAACAAACGGTATCTTGAACAAGCCAAGCTGTGGTCAGAAATATTCCACCATAGATCAGTAAATCCTAGAAAAGAACTGGTACACTGGGTCGAACATGTTATTCACACTCGTGGAGCCACTTATCTACGATCTCCGGCATTGGACGTGCCTTTATACCAAAAGATGTATTTGGACCTTTTGGGCCTAGTTATGCTAGTTTTGATGGCATTAACATTCTTAATTAAGAATGTGATAAAATTATTTATGGTTCGGGGTATAGTACACGAGAAGATGGAATGA
Protein
MMIRRLTIAIAVCFCLGVDAYKILTVFPVPGRSHGILGDAVVRHLLEAGHEVTHITPFPKKEPPPNLVQIDVAANKAAFNEDYIDIKALMTKEFNLKDKNVLFSLMNNISSSTILNENVQRLLRDQSREQFDVIIAEWMFSDLYASFHAVLDCPLIWFSTIEPHWMVLRLIDEYPNPAYTSHFQDSFEVPFTFVERMSVLSSQLTWSLSLNTWVYDLEKYIYDNNIAPIIKKNGKPVPNYDEVRYNGSLLLGNSHVSLGDAIKVPINYKAIGGYHIDGKVKELPPDLQKIMNESKHGVIYFSMGSNLKSKDLPKEIKEGLLKMFSQLKQTVLWKFEENLSPLPENVHLLKWAPQQSILAHPNCILFITHGGLLSTTEAVHFGKPIIGIPVFADQFGNVNRAVQKGIARRVDLSFTMVRDLEEAVAEMINNSRYIEKIKELSLIYHDRPVSPGAELVHWVEHVVKTKGALHLRSPALHVPFYQKLYLDLLAIVLVTLFSFVLTCDCYKILIVFTTPMKSHNILGEAAAELLLNAGHEVTYVTPFPKESVPDKMRQVDVTYIGKIGLFDLKGYLKNDTVKPMSLRKISYFMHDVNVKAIQNENLQQLLNDPSQRFDAVIVDWLFSEIFVGLASLYDCPLIWMSTMDPHWQILRLVDEMPNPVFLGRCFLDRIVPFRFWERTQELLYQISSLFFKDIEFFSEEDAAFKRLLGPVFARKKKPLPSFNAVRYNASLVLSNSHHSIGYPVKLPPNFISIGGFFIDDKKQRLSLDLQTIMDNAKHGVILFSLGSNLKSKDMPEHLVRSLLNVFSELKQIVIWKVEEQIADLPQNVHVLKWLPQQSILAHSNCILFITHGGLLSITEAYHHGVPLIGIPVFADQFKNVNLVSKKGFAKKVDLTYNLPGDLKHAINEILHNKRYLEQAKLWSEIFHHRSVNPRKELVHWVEHVIHTRGATYLRSPALDVPLYQKMYLDLLGLVMLVLMALTFLIKNVIKLFMVRGIVHEKME
Summary
Similarity
Belongs to the UDP-glycosyltransferase family.
Uniprot
EMBL
BABH01011785
BABH01011786
BABH01011787
NWSH01000282
PCG77781.1
KQ460597
+ More
KPJ13897.1 AGBW02009118 OWR51610.1 ODYU01006413 SOQ48233.1 KQ460075 KPJ17950.1 KQ459582 KPI98794.1 JTDY01003050 KOB70242.1 KQ971371 EFA10103.2 EFA10106.1 APGK01058731 KB741291 ENN70577.1 KB631579 ERL84451.1 JRES01001197 KNC24662.1 KYB25474.1 KNC24650.1 KYB25566.1
KPJ13897.1 AGBW02009118 OWR51610.1 ODYU01006413 SOQ48233.1 KQ460075 KPJ17950.1 KQ459582 KPI98794.1 JTDY01003050 KOB70242.1 KQ971371 EFA10103.2 EFA10106.1 APGK01058731 KB741291 ENN70577.1 KB631579 ERL84451.1 JRES01001197 KNC24662.1 KYB25474.1 KNC24650.1 KYB25566.1
Proteomes
PRIDE
Pfam
PF00201 UDPGT
ProteinModelPortal
PDB
2O6L
E-value=2.32639e-24,
Score=282
Ontologies
PATHWAY
00040
Pentose and glucuronate interconversions - Bombyx mori (domestic silkworm)
00053 Ascorbate and aldarate metabolism - Bombyx mori (domestic silkworm)
00830 Retinol metabolism - Bombyx mori (domestic silkworm)
00860 Porphyrin and chlorophyll metabolism - Bombyx mori (domestic silkworm)
00980 Metabolism of xenobiotics by cytochrome P450 - Bombyx mori (domestic silkworm)
00982 Drug metabolism - cytochrome P450 - Bombyx mori (domestic silkworm)
00983 Drug metabolism - other enzymes - Bombyx mori (domestic silkworm)
01100 Metabolic pathways - Bombyx mori (domestic silkworm)
00053 Ascorbate and aldarate metabolism - Bombyx mori (domestic silkworm)
00830 Retinol metabolism - Bombyx mori (domestic silkworm)
00860 Porphyrin and chlorophyll metabolism - Bombyx mori (domestic silkworm)
00980 Metabolism of xenobiotics by cytochrome P450 - Bombyx mori (domestic silkworm)
00982 Drug metabolism - cytochrome P450 - Bombyx mori (domestic silkworm)
00983 Drug metabolism - other enzymes - Bombyx mori (domestic silkworm)
01100 Metabolic pathways - Bombyx mori (domestic silkworm)
GO
Topology
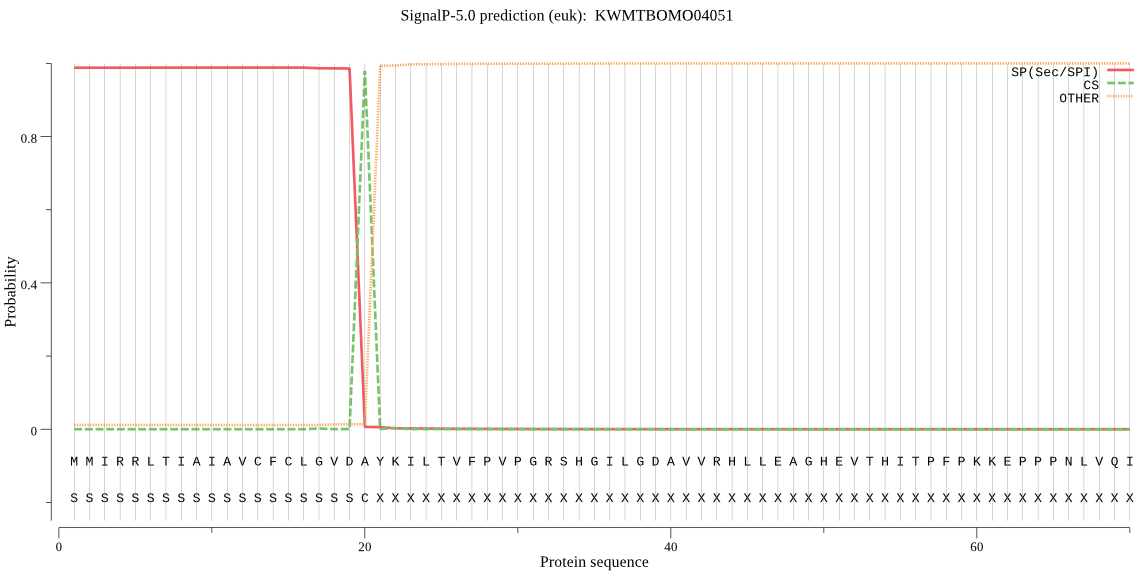
SignalP
Position: 1 - 20,
Likelihood: 0.987682
Length:
1003
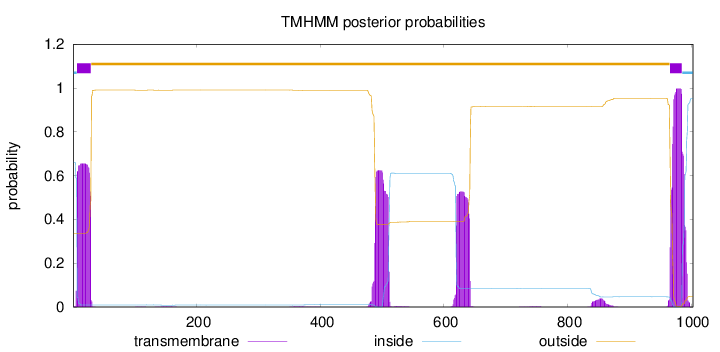
Number of predicted TMHs:
2
Exp number of AAs in TMHs:
63.29305
Exp number, first 60 AAs:
14.80966
Total prob of N-in:
0.66300
POSSIBLE N-term signal
sequence
inside
1 - 6
TMhelix
7 - 29
outside
30 - 965
TMhelix
966 - 985
inside
986 - 1003
Population Genetic Test Statistics
Pi
200.354769
Theta
156.257387
Tajima's D
1.151052
CLR
0.197144
CSRT
0.700064996750163
Interpretation
Uncertain
