Gene
KWMTBOMO04047
Pre Gene Modal
BGIBMGA010286
Annotation
PREDICTED:_UDP-glucosyltransferase_isoform_X1_[Bombyx_mori]
Full name
UDP-glucuronosyltransferase
Location in the cell
Peroxisomal Reliability : 1.101 PlasmaMembrane Reliability : 1.288
Sequence
CDS
ATGTTCAAGCTCACATTCCTGGTCTGCTGTATCCTGGCTACTCAATCCGTTTCTGATGCCTATAAAATCCTGGTGGTGTTCCCTATGCCCGGCAAAAGCCACAGTATCCTGGGGTACAGTGTCGTGAAGCATCTCCTAAAAGCTGGTCATGAGGTGACATATGTAACTCCTTTCGTTGAGGACAATCACCACCCGAACCTGACTCAAGTTGATGTTAGCAGCAATATGCGTTTGATACCTAAGGGCGGATTAGACTTGAAGAGAGTTCTAGACAAAGAGGTAAACGTAATAGACAACGGGTTTATGTTTTATTTCATGAAACAAATACAGGAAGCTACTCTGGAGCATGAGCAAGTTAAAAAGTTGCTCGAGGACCCGAATAAGACCTTCGATATTGTTATCGTTGAGTGGATGTATTGCGAATTGGGTGCTAGTTATGCTGCGGTATTCGACGTTCCACTTATATGGTTGTCGACCATGGAGCCTCACTGGTTGGTGACGAGGCTCATCGATGGGAATCTAAATCCAGCCTACAACGGGGATTCTATGTCTTCCAGTATTCCTCCATTTACGTTCTTGCAAAGAGTCAAGGAACTCTGGATTCAAATTCACACTAGTTTTATATTGCTCAATGATGATCAAGAGAGATCGTACGACAGATTAGTGAGGCCACTCATTGAAAAAAAAGGTAGGAAGGCGCCTTCGTTTGAGGATTTGAAGTTCAATGCTTCTTTAGTACTCGGAAATTCGCATGTATCTTTGGGAGAAGCAACCGGAACTCCGCAGAGCTACAAACCCATCGCTGGATACCATATAGAAGAAGTTGTGAAACCTTTACCAGCTGACCTCAAAGAGATAATGGAAAACGCAAAGCACGGTGTCATCTATTTCAGTATGGGATCAAATTTGAAGAGTACAGAAATGCCAGATGAGATGAAGCAGAATCTTGTAAAAATGTTCGGTGAATTGAAACAGACTATTATCTGGAAGTTTGAAGAAGACTTCCCGAATCTCCCGAAAAACGTCCACATCGTAAATTGGGCACCACAACCAAGTATTTTATCTCATCCTAATTGCGTGTTATTCATCACACACGGAGGTCTACTTTCAACGACCGAAAGCGTACATTTTGGAGTGCCAATTGTGGGAATACCAGTTTTTGGTGATCAATTTATCAATGTCCAAAGGGCTGTTAAAAGGGGATTCGCGAAGAAAGTTGATTTCTCGTACTCAATGGTCGGAGAACTTAAGGTCGCTATTCAAGAAATTTTAAGTGACTCCAGCTACAGAACCAGAATTAAGGAGCTATCGTTGATATATCACGATCGTCCGGTGTCTCCTGGTGCCGAGCTGGTGCATTGGGTTGAGCACGTGGCCAGGACAAGAGGAGCTCTCCATCTGCGCTCACCAGCACTGCACGTGCCTTTCTACCAGAAGTTGTACTTAGATCTATTGGCCGTTGTATTAATAATTTCGCTAATCTTTTATAGAAAAATTTGTTTGATAAAAAATCTTTTACTGTCATTTTTTCAAACAAATGAAATCAAGAAGAAAAAGAAGAGGAATTAA
Protein
MFKLTFLVCCILATQSVSDAYKILVVFPMPGKSHSILGYSVVKHLLKAGHEVTYVTPFVEDNHHPNLTQVDVSSNMRLIPKGGLDLKRVLDKEVNVIDNGFMFYFMKQIQEATLEHEQVKKLLEDPNKTFDIVIVEWMYCELGASYAAVFDVPLIWLSTMEPHWLVTRLIDGNLNPAYNGDSMSSSIPPFTFLQRVKELWIQIHTSFILLNDDQERSYDRLVRPLIEKKGRKAPSFEDLKFNASLVLGNSHVSLGEATGTPQSYKPIAGYHIEEVVKPLPADLKEIMENAKHGVIYFSMGSNLKSTEMPDEMKQNLVKMFGELKQTIIWKFEEDFPNLPKNVHIVNWAPQPSILSHPNCVLFITHGGLLSTTESVHFGVPIVGIPVFGDQFINVQRAVKRGFAKKVDFSYSMVGELKVAIQEILSDSSYRTRIKELSLIYHDRPVSPGAELVHWVEHVARTRGALHLRSPALHVPFYQKLYLDLLAVVLIISLIFYRKICLIKNLLLSFFQTNEIKKKKKRN
Summary
Catalytic Activity
glucuronate acceptor + UDP-alpha-D-glucuronate = acceptor beta-D-glucuronoside + H(+) + UDP
Similarity
Belongs to the UDP-glycosyltransferase family.
Feature
chain UDP-glucuronosyltransferase
Uniprot
G9LPU8
D6RUU6
G9LPU6
D6RUU7
A0A2H1VVH6
A0A2W1BQS6
+ More
A0A2A4K0I8 A0A024AGG4 A0A2H1WNF9 G9LPQ7 A0A2A4K2B6 G9LPU7 D6RUV0 H9JL88 A0A191T2J8 A0A2W1BXG7 G9LPQ0 A0A191T2G6 A0A2W1BVZ0 Q8WPG4 A0A191T1M8 A0A140EA36 A0A3S2LD93 G9LPQ1 G9LPU0 A0A2H4WB58 A0A024AH17 A0A191T2K2 A0A2H4WB67 A0A0L7L1M9 A0A024AHN0 A0A191T1N5 A0A1L8D6H0 D6RUU8 A0A2H1X1Y8 G9LPQ3 A0A2H1W225 A0A3S2LTQ3 D0PWZ6 A0A1L8D6R9 A0A2W1BQJ5 A0A2A4J419 A0A191T1M6 A0A191T1M1 A0A194RKU4 A0A2H1V1W5 G9LPQ6 A0A140EA37 A0A194Q0G5 D6RUV2 A0A194Q5Q0 A0A194RKA9 G9LPV0 D6RUU9 A0A2A4J6T0 A0A191T1M3 G9LPU3 G9LPQ5 A0A2W1BX32 A0A2W1BSQ4 I4DN89 A0A3S2NJP8 A0A0N1IN21 A0A191T2G9 A0A194R8C7 A0A3S2P0K8 A0A2A4J325 G9LPU4 A0A212FKT5 A0A2H1WQL3 A0A191T1P1 I4DR17 A0A2H4WB78 H9JKP9 A0A3S2TLE9 A0A1L8D6E4 G9LPQ2 A0A2H4WB53 A0A024AG04 A0A191T2K0 D6RUV1 A0A2A4J0B4 A0A2H1VVH4 A0A194RDA8 A0A194Q1C7 G9LPU2 G9LPQ4 A0A212FD09 A0A2H1W5E5 G9LPU1 A0A2W1BQJ6 A0A0L7L448 A0A212ERW0 A0A1L8D6G3
A0A2A4K0I8 A0A024AGG4 A0A2H1WNF9 G9LPQ7 A0A2A4K2B6 G9LPU7 D6RUV0 H9JL88 A0A191T2J8 A0A2W1BXG7 G9LPQ0 A0A191T2G6 A0A2W1BVZ0 Q8WPG4 A0A191T1M8 A0A140EA36 A0A3S2LD93 G9LPQ1 G9LPU0 A0A2H4WB58 A0A024AH17 A0A191T2K2 A0A2H4WB67 A0A0L7L1M9 A0A024AHN0 A0A191T1N5 A0A1L8D6H0 D6RUU8 A0A2H1X1Y8 G9LPQ3 A0A2H1W225 A0A3S2LTQ3 D0PWZ6 A0A1L8D6R9 A0A2W1BQJ5 A0A2A4J419 A0A191T1M6 A0A191T1M1 A0A194RKU4 A0A2H1V1W5 G9LPQ6 A0A140EA37 A0A194Q0G5 D6RUV2 A0A194Q5Q0 A0A194RKA9 G9LPV0 D6RUU9 A0A2A4J6T0 A0A191T1M3 G9LPU3 G9LPQ5 A0A2W1BX32 A0A2W1BSQ4 I4DN89 A0A3S2NJP8 A0A0N1IN21 A0A191T2G9 A0A194R8C7 A0A3S2P0K8 A0A2A4J325 G9LPU4 A0A212FKT5 A0A2H1WQL3 A0A191T1P1 I4DR17 A0A2H4WB78 H9JKP9 A0A3S2TLE9 A0A1L8D6E4 G9LPQ2 A0A2H4WB53 A0A024AG04 A0A191T2K0 D6RUV1 A0A2A4J0B4 A0A2H1VVH4 A0A194RDA8 A0A194Q1C7 G9LPU2 G9LPQ4 A0A212FD09 A0A2H1W5E5 G9LPU1 A0A2W1BQJ6 A0A0L7L448 A0A212ERW0 A0A1L8D6G3
EC Number
2.4.1.17
Pubmed
EMBL
JQ070255
AEW43171.1
AB539963
BAJ08153.1
JQ070253
AEW43169.1
+ More
JQ070252 AB539964 AEW43168.1 BAJ08154.1 ODYU01004685 SOQ44830.1 KZ149930 PZC77368.1 NWSH01000282 PCG77781.1 KF777115 AHY99686.1 ODYU01009829 SOQ54516.1 JQ070214 AEW43130.1 PCG77782.1 JQ070254 AEW43170.1 AB539967 BAJ08157.1 BABH01011785 BABH01011786 BABH01011787 KU680294 ANI22000.1 PZC77366.1 JQ070207 AEW43123.1 KU680295 ANI22001.1 PZC77367.1 AF324465 AAL55994.1 KU680303 ANI22009.1 KU214507 AMK97473.1 RSAL01000210 RVE44216.1 JQ070208 AEW43124.1 JQ070247 AEW43163.1 KY202932 AUC64276.1 KF777116 AHY99687.1 KU680304 ANI22010.1 KY202937 AUC64281.1 JTDY01003683 KOB69154.1 KF777113 AHY99684.1 KU680296 ANI22002.1 GEYN01000076 JAV02053.1 AB539965 BAJ08155.1 ODYU01012744 SOQ59206.1 JQ070210 AEW43126.1 ODYU01005852 SOQ47145.1 RVE44215.1 FJ373022 ACQ66003.1 GEYN01000085 JAV02044.1 PZC77369.1 NWSH01003535 PCG66153.1 KU680302 ANI22008.1 KU680297 ANI22003.1 KQ460075 KPJ17950.1 ODYU01000116 SOQ34342.1 JQ070213 AEW43129.1 KU214508 AMK97474.1 KQ459582 KPI98793.1 JQ070256 AB539969 AEW43172.1 BAJ08159.1 KPI98725.1 KPJ17977.1 JQ070257 AEW43173.1 AB539966 BAJ08156.1 NWSH01002698 PCG67661.1 KU680298 ANI22004.1 JQ070250 AEW43166.1 JQ070212 AEW43128.1 PZC77370.1 PZC77371.1 AK402777 BAM19379.1 RSAL01000067 RVE49308.1 KQ459356 KPJ01283.1 KU680300 ANI22006.1 KQ460597 KPJ13897.1 RVE49310.1 NWSH01003818 PCG65793.1 JQ070251 AEW43167.1 AGBW02007992 OWR54346.1 ODYU01010264 SOQ55252.1 KU680301 ANI22007.1 AK404892 BAM20357.1 KY202946 AUC64290.1 RVE49311.1 GEYN01000100 JAV02029.1 JQ070209 AEW43125.1 KY202925 AUC64269.1 KF777114 AHY99685.1 KU680299 ANI22005.1 JQ070258 AB539968 AEW43174.1 BAJ08158.1 NWSH01003979 PCG65597.1 SOQ44829.1 KPJ13896.1 KPI98794.1 JQ070249 AEW43165.1 JQ070211 AEW43127.1 AGBW02009118 OWR51610.1 ODYU01006413 SOQ48233.1 JQ070248 AEW43164.1 PZC77372.1 JTDY01003050 KOB70242.1 AGBW02012922 OWR44215.1 GEYN01000087 JAV02042.1
JQ070252 AB539964 AEW43168.1 BAJ08154.1 ODYU01004685 SOQ44830.1 KZ149930 PZC77368.1 NWSH01000282 PCG77781.1 KF777115 AHY99686.1 ODYU01009829 SOQ54516.1 JQ070214 AEW43130.1 PCG77782.1 JQ070254 AEW43170.1 AB539967 BAJ08157.1 BABH01011785 BABH01011786 BABH01011787 KU680294 ANI22000.1 PZC77366.1 JQ070207 AEW43123.1 KU680295 ANI22001.1 PZC77367.1 AF324465 AAL55994.1 KU680303 ANI22009.1 KU214507 AMK97473.1 RSAL01000210 RVE44216.1 JQ070208 AEW43124.1 JQ070247 AEW43163.1 KY202932 AUC64276.1 KF777116 AHY99687.1 KU680304 ANI22010.1 KY202937 AUC64281.1 JTDY01003683 KOB69154.1 KF777113 AHY99684.1 KU680296 ANI22002.1 GEYN01000076 JAV02053.1 AB539965 BAJ08155.1 ODYU01012744 SOQ59206.1 JQ070210 AEW43126.1 ODYU01005852 SOQ47145.1 RVE44215.1 FJ373022 ACQ66003.1 GEYN01000085 JAV02044.1 PZC77369.1 NWSH01003535 PCG66153.1 KU680302 ANI22008.1 KU680297 ANI22003.1 KQ460075 KPJ17950.1 ODYU01000116 SOQ34342.1 JQ070213 AEW43129.1 KU214508 AMK97474.1 KQ459582 KPI98793.1 JQ070256 AB539969 AEW43172.1 BAJ08159.1 KPI98725.1 KPJ17977.1 JQ070257 AEW43173.1 AB539966 BAJ08156.1 NWSH01002698 PCG67661.1 KU680298 ANI22004.1 JQ070250 AEW43166.1 JQ070212 AEW43128.1 PZC77370.1 PZC77371.1 AK402777 BAM19379.1 RSAL01000067 RVE49308.1 KQ459356 KPJ01283.1 KU680300 ANI22006.1 KQ460597 KPJ13897.1 RVE49310.1 NWSH01003818 PCG65793.1 JQ070251 AEW43167.1 AGBW02007992 OWR54346.1 ODYU01010264 SOQ55252.1 KU680301 ANI22007.1 AK404892 BAM20357.1 KY202946 AUC64290.1 RVE49311.1 GEYN01000100 JAV02029.1 JQ070209 AEW43125.1 KY202925 AUC64269.1 KF777114 AHY99685.1 KU680299 ANI22005.1 JQ070258 AB539968 AEW43174.1 BAJ08158.1 NWSH01003979 PCG65597.1 SOQ44829.1 KPJ13896.1 KPI98794.1 JQ070249 AEW43165.1 JQ070211 AEW43127.1 AGBW02009118 OWR51610.1 ODYU01006413 SOQ48233.1 JQ070248 AEW43164.1 PZC77372.1 JTDY01003050 KOB70242.1 AGBW02012922 OWR44215.1 GEYN01000087 JAV02042.1
Proteomes
PRIDE
Pfam
PF00201 UDPGT
ProteinModelPortal
G9LPU8
D6RUU6
G9LPU6
D6RUU7
A0A2H1VVH6
A0A2W1BQS6
+ More
A0A2A4K0I8 A0A024AGG4 A0A2H1WNF9 G9LPQ7 A0A2A4K2B6 G9LPU7 D6RUV0 H9JL88 A0A191T2J8 A0A2W1BXG7 G9LPQ0 A0A191T2G6 A0A2W1BVZ0 Q8WPG4 A0A191T1M8 A0A140EA36 A0A3S2LD93 G9LPQ1 G9LPU0 A0A2H4WB58 A0A024AH17 A0A191T2K2 A0A2H4WB67 A0A0L7L1M9 A0A024AHN0 A0A191T1N5 A0A1L8D6H0 D6RUU8 A0A2H1X1Y8 G9LPQ3 A0A2H1W225 A0A3S2LTQ3 D0PWZ6 A0A1L8D6R9 A0A2W1BQJ5 A0A2A4J419 A0A191T1M6 A0A191T1M1 A0A194RKU4 A0A2H1V1W5 G9LPQ6 A0A140EA37 A0A194Q0G5 D6RUV2 A0A194Q5Q0 A0A194RKA9 G9LPV0 D6RUU9 A0A2A4J6T0 A0A191T1M3 G9LPU3 G9LPQ5 A0A2W1BX32 A0A2W1BSQ4 I4DN89 A0A3S2NJP8 A0A0N1IN21 A0A191T2G9 A0A194R8C7 A0A3S2P0K8 A0A2A4J325 G9LPU4 A0A212FKT5 A0A2H1WQL3 A0A191T1P1 I4DR17 A0A2H4WB78 H9JKP9 A0A3S2TLE9 A0A1L8D6E4 G9LPQ2 A0A2H4WB53 A0A024AG04 A0A191T2K0 D6RUV1 A0A2A4J0B4 A0A2H1VVH4 A0A194RDA8 A0A194Q1C7 G9LPU2 G9LPQ4 A0A212FD09 A0A2H1W5E5 G9LPU1 A0A2W1BQJ6 A0A0L7L448 A0A212ERW0 A0A1L8D6G3
A0A2A4K0I8 A0A024AGG4 A0A2H1WNF9 G9LPQ7 A0A2A4K2B6 G9LPU7 D6RUV0 H9JL88 A0A191T2J8 A0A2W1BXG7 G9LPQ0 A0A191T2G6 A0A2W1BVZ0 Q8WPG4 A0A191T1M8 A0A140EA36 A0A3S2LD93 G9LPQ1 G9LPU0 A0A2H4WB58 A0A024AH17 A0A191T2K2 A0A2H4WB67 A0A0L7L1M9 A0A024AHN0 A0A191T1N5 A0A1L8D6H0 D6RUU8 A0A2H1X1Y8 G9LPQ3 A0A2H1W225 A0A3S2LTQ3 D0PWZ6 A0A1L8D6R9 A0A2W1BQJ5 A0A2A4J419 A0A191T1M6 A0A191T1M1 A0A194RKU4 A0A2H1V1W5 G9LPQ6 A0A140EA37 A0A194Q0G5 D6RUV2 A0A194Q5Q0 A0A194RKA9 G9LPV0 D6RUU9 A0A2A4J6T0 A0A191T1M3 G9LPU3 G9LPQ5 A0A2W1BX32 A0A2W1BSQ4 I4DN89 A0A3S2NJP8 A0A0N1IN21 A0A191T2G9 A0A194R8C7 A0A3S2P0K8 A0A2A4J325 G9LPU4 A0A212FKT5 A0A2H1WQL3 A0A191T1P1 I4DR17 A0A2H4WB78 H9JKP9 A0A3S2TLE9 A0A1L8D6E4 G9LPQ2 A0A2H4WB53 A0A024AG04 A0A191T2K0 D6RUV1 A0A2A4J0B4 A0A2H1VVH4 A0A194RDA8 A0A194Q1C7 G9LPU2 G9LPQ4 A0A212FD09 A0A2H1W5E5 G9LPU1 A0A2W1BQJ6 A0A0L7L448 A0A212ERW0 A0A1L8D6G3
PDB
2O6L
E-value=1.2326e-25,
Score=290
Ontologies
PATHWAY
00040
Pentose and glucuronate interconversions - Bombyx mori (domestic silkworm)
00053 Ascorbate and aldarate metabolism - Bombyx mori (domestic silkworm)
00830 Retinol metabolism - Bombyx mori (domestic silkworm)
00860 Porphyrin and chlorophyll metabolism - Bombyx mori (domestic silkworm)
00980 Metabolism of xenobiotics by cytochrome P450 - Bombyx mori (domestic silkworm)
00982 Drug metabolism - cytochrome P450 - Bombyx mori (domestic silkworm)
00983 Drug metabolism - other enzymes - Bombyx mori (domestic silkworm)
01100 Metabolic pathways - Bombyx mori (domestic silkworm)
00053 Ascorbate and aldarate metabolism - Bombyx mori (domestic silkworm)
00830 Retinol metabolism - Bombyx mori (domestic silkworm)
00860 Porphyrin and chlorophyll metabolism - Bombyx mori (domestic silkworm)
00980 Metabolism of xenobiotics by cytochrome P450 - Bombyx mori (domestic silkworm)
00982 Drug metabolism - cytochrome P450 - Bombyx mori (domestic silkworm)
00983 Drug metabolism - other enzymes - Bombyx mori (domestic silkworm)
01100 Metabolic pathways - Bombyx mori (domestic silkworm)
GO
Topology
Subcellular location
Membrane
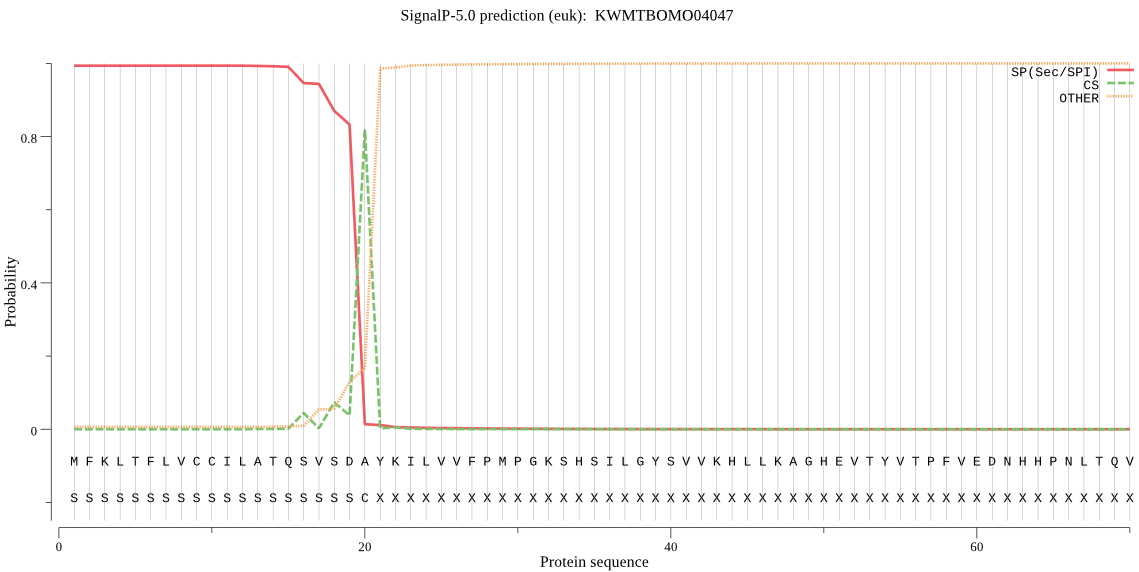
SignalP
Position: 1 - 20,
Likelihood: 0.993325
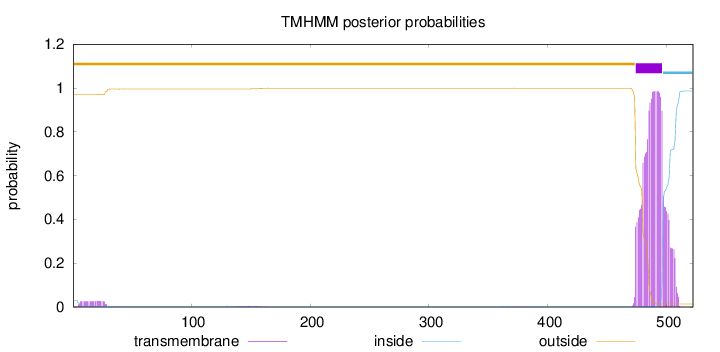
Length:
522
Number of predicted TMHs:
1
Exp number of AAs in TMHs:
22.52149
Exp number, first 60 AAs:
0.61798
Total prob of N-in:
0.03002
outside
1 - 473
TMhelix
474 - 496
inside
497 - 522
Population Genetic Test Statistics
Pi
25.273633
Theta
23.402756
Tajima's D
0.698552
CLR
120.424005
CSRT
0.581270936453177
Interpretation
Uncertain
