Gene
KWMTBOMO03923
Annotation
PREDICTED:_uncharacterized_protein_LOC105388498_[Plutella_xylostella]
Location in the cell
Nuclear Reliability : 4.096
Sequence
CDS
ATGGACAAACTTGAACTTCAATCAACTTGTTTCGAGAATATCAAGACTTTGTGTACAAACTACCGCAAAGATTCTTCGTCTCGAAAAACTGAAGAGTACCTCAACATACGTTTGTCATCTTTGGAAGAACAGTGGAGAGACTTCGAGGCGAGAAATGCCTTATTACTACTGGAACTGGAAGACAAAACTATACATTATTTCTCTGAAGACGTTTACAACAAAACTAAAGAAATGTATCTAGCAACAAAAACAGACATAAACAACAGACTTAAATACTTGAAACAGCAAAAACATAGTGTGAATTTTGATTTAAGTGACATTTCTGTTGACTACACTCAAGACTGCGAGGACGGAAAATCTGAAAAAATACAACAACTTTTAAATAAACAAGAATGCAATTTTAATGCGATTTATCATGCATTGTCTAAAATTGACATCATTAGAACTAGTGAAAAGTGGGAGCTGGAAGATAATTTTAACATTTTAAAATCGAAGTGGGAGTCCATCGACAAACTGCACTGGGAAATTGACTTTCAGCTGAAGGGGTCTAACGATCACTATTACCGCATGTACTCAGATATCGAAGAAAAATATGATGAATTGAGAAAACAACTTCAATCTAAAATTTGGTCGAATGCACACTATCAGCAGTCGACACCGAGAATGGAATTACCTGATTTTTACGGAAACTACAGTAAGTGGATCTCTTTTAAGGATCTGTTTCTCGAAACCATCCATAACAACCCAACAATTAACAAAGCGCAAAAAATGAAACACCTGAAAACCAAATTGAAGGGGGAAGCAGAAAGTCTGGTACAGCATTTATCCGTCTCTGCTGAAAACTACAACTCCTGCTGGGATATTTTAGTGCAACGGTATGATAATCGCCGCCTACAATTTTCATCGTTTTTCAACACAATGATGAGTCTCCCTGTTATACATCAACCCGACGCTAACAATCTCAAGAAAATGCATGACGTCATTAAGGAATGCATGAACGGCATAGCTAACATCGGGCTGGATACGACAACGTGGGATCCAATGATCGTGCATTTAATGTTGCCGAAATTAGATTCAACAACTTTAAATGATTATATGAACGAAATACGAGATCACAGAGAATTACAAAACTTGCAAGATTTTTTATATTTTTTGGAGTGCAAGTTTATGGCATATGAAACTGCAAAATCAACAAAATCTGTAAAAGATACCACTTCCGAAAAAACATCTCATTGGAAACCGGCTAACAACAAACAACCTGTAAAAAAATTTACATATGACTCTAAGAAAACTTATAACTATCACGCTACTTATGGAAAATGCCCGAATTGTAATGGAAATCATGTTTTAATGCAATGTGCTAAATTTATTAACATGGATACTGCACAGCGAAACCACACTGTTGCGAAATTGAAACTCTGCAAAAATTGTCTCTATAGCCACAATAACGAAGAATGTAAATCAGCGAAAACGTGTAAGGAGTGTAACATGAAACATCATACCATGTTGCACAACGTGAACTGGAAAATTCCAGCGAATGGTAAAAATAACTTTGGACAACAAGCCTCCTCTTCTCAGCAACAAGCTTCGAATCACTTATCATCGGATAAAACTGAAATTTTACTAACTACAACTCTACTGAAAATAAAGGCTTATGATGGCACTTATGTAAAACTTAGGGCCCTCTTGGATCAGGGCTCGCAAGTAAACTTAATAACAGAAAATGCTGCTCAGGTATTACGCCTGCCGAGGAAAAGGTTCGCTGCTACCATCTCGGGGGTTGGTTCAGTATCTGGTGATTGCAGGGGTCAACTAAACTTATCTTGTAAATCAATAAACTCTGAATACGACTTCGAAATACAAGCACTTATAATGAAAAAACTTACAAATAACTTACCGAATGCTACTTTCGAAAAATCGAATTGGCCTCATTTGGAAAACTTAAAGCTGGCTGATCCTGACTTCAACATCTCTCGCCCCATCGACTTACTACTGGGTGCCGATGTTTATTCAGACATTATACTCAATGGAGTTTTAAAATACAACGTGCTCCCGACGGAAAATATATTGTGA
Protein
MDKLELQSTCFENIKTLCTNYRKDSSSRKTEEYLNIRLSSLEEQWRDFEARNALLLLELEDKTIHYFSEDVYNKTKEMYLATKTDINNRLKYLKQQKHSVNFDLSDISVDYTQDCEDGKSEKIQQLLNKQECNFNAIYHALSKIDIIRTSEKWELEDNFNILKSKWESIDKLHWEIDFQLKGSNDHYYRMYSDIEEKYDELRKQLQSKIWSNAHYQQSTPRMELPDFYGNYSKWISFKDLFLETIHNNPTINKAQKMKHLKTKLKGEAESLVQHLSVSAENYNSCWDILVQRYDNRRLQFSSFFNTMMSLPVIHQPDANNLKKMHDVIKECMNGIANIGLDTTTWDPMIVHLMLPKLDSTTLNDYMNEIRDHRELQNLQDFLYFLECKFMAYETAKSTKSVKDTTSEKTSHWKPANNKQPVKKFTYDSKKTYNYHATYGKCPNCNGNHVLMQCAKFINMDTAQRNHTVAKLKLCKNCLYSHNNEECKSAKTCKECNMKHHTMLHNVNWKIPANGKNNFGQQASSSQQQASNHLSSDKTEILLTTTLLKIKAYDGTYVKLRALLDQGSQVNLITENAAQVLRLPRKRFAATISGVGSVSGDCRGQLNLSCKSINSEYDFEIQALIMKKLTNNLPNATFEKSNWPHLENLKLADPDFNISRPIDLLLGADVYSDIILNGVLKYNVLPTENIL
Summary
Uniprot
A0A2W1BPI5
A0A2W1AZ18
A0A0L7LHD6
A0A2A4JZ49
A0A2W1BMV4
A0A2A4K6T9
+ More
A0A2H1X238 A0A2W1BJC9 A0A2H1W714 A0A2H1VXK6 A0A0L7LGK2 A0A0L7KWQ7 A0A0J7N314 A0A1Y1LE43 A0A1Y1LBZ3 A0A0J7N890 A0A0J7K152 A0A0J7N5C9 A0A0J7N536 A0A0J7K907 A0A3L8DYA9 A0A1B0DAL2 A0A151WGR4 A0A0N1IHU1 A0A2H1WI74 A0A0J7KPJ2 A0A3L8D6F4 X1WYQ8 A0A0J7KB34 A0A0J7KJY9 A0A151IN79 J9LHB7 J9JNP4 A0A226DBN6 A0A0J7KCT8 A0A2S2PTX9 J9LYM7 J9K774 W8B8B3 A0A224XG39 A0A1B0CQ10 A0A151IDD4
A0A2H1X238 A0A2W1BJC9 A0A2H1W714 A0A2H1VXK6 A0A0L7LGK2 A0A0L7KWQ7 A0A0J7N314 A0A1Y1LE43 A0A1Y1LBZ3 A0A0J7N890 A0A0J7K152 A0A0J7N5C9 A0A0J7N536 A0A0J7K907 A0A3L8DYA9 A0A1B0DAL2 A0A151WGR4 A0A0N1IHU1 A0A2H1WI74 A0A0J7KPJ2 A0A3L8D6F4 X1WYQ8 A0A0J7KB34 A0A0J7KJY9 A0A151IN79 J9LHB7 J9JNP4 A0A226DBN6 A0A0J7KCT8 A0A2S2PTX9 J9LYM7 J9K774 W8B8B3 A0A224XG39 A0A1B0CQ10 A0A151IDD4
EMBL
KZ150005
PZC75227.1
KZ150641
PZC70422.1
JTDY01001118
KOB74820.1
+ More
NWSH01000402 PCG76662.1 KZ149989 PZC75621.1 NWSH01000070 PCG79985.1 ODYU01012875 SOQ59395.1 KZ150008 PZC75149.1 ODYU01006730 SOQ48837.1 ODYU01005059 SOQ45565.1 JTDY01001277 KOB74321.1 JTDY01005003 KOB67489.1 LBMM01011051 KMQ87060.1 GEZM01060253 JAV71158.1 GEZM01060254 JAV71154.1 LBMM01008464 KMQ88855.1 LBMM01017223 KMQ84153.1 LBMM01009746 KMQ87900.1 LBMM01009898 KMQ87805.1 LBMM01011516 KMQ86784.1 QOIP01000003 RLU24798.1 AJVK01029183 KQ983160 KYQ46995.1 KQ460036 KPJ18393.1 ODYU01008829 SOQ52773.1 LBMM01004572 KMQ92278.1 QOIP01000012 RLU15613.1 ABLF02028666 ABLF02066713 LBMM01010405 KMQ87487.1 LBMM01006490 KMQ90572.1 KQ976954 KYN06860.1 ABLF02022040 ABLF02056888 ABLF02005684 LNIX01000026 OXA42254.1 LBMM01009284 KMQ88248.1 GGMR01020322 MBY32941.1 ABLF02016109 ABLF02016113 ABLF02023723 GAMC01020546 JAB86009.1 GFTR01008874 JAW07552.1 AJWK01022907 KQ977963 KYM98425.1
NWSH01000402 PCG76662.1 KZ149989 PZC75621.1 NWSH01000070 PCG79985.1 ODYU01012875 SOQ59395.1 KZ150008 PZC75149.1 ODYU01006730 SOQ48837.1 ODYU01005059 SOQ45565.1 JTDY01001277 KOB74321.1 JTDY01005003 KOB67489.1 LBMM01011051 KMQ87060.1 GEZM01060253 JAV71158.1 GEZM01060254 JAV71154.1 LBMM01008464 KMQ88855.1 LBMM01017223 KMQ84153.1 LBMM01009746 KMQ87900.1 LBMM01009898 KMQ87805.1 LBMM01011516 KMQ86784.1 QOIP01000003 RLU24798.1 AJVK01029183 KQ983160 KYQ46995.1 KQ460036 KPJ18393.1 ODYU01008829 SOQ52773.1 LBMM01004572 KMQ92278.1 QOIP01000012 RLU15613.1 ABLF02028666 ABLF02066713 LBMM01010405 KMQ87487.1 LBMM01006490 KMQ90572.1 KQ976954 KYN06860.1 ABLF02022040 ABLF02056888 ABLF02005684 LNIX01000026 OXA42254.1 LBMM01009284 KMQ88248.1 GGMR01020322 MBY32941.1 ABLF02016109 ABLF02016113 ABLF02023723 GAMC01020546 JAB86009.1 GFTR01008874 JAW07552.1 AJWK01022907 KQ977963 KYM98425.1
Proteomes
Pfam
Interpro
IPR005312
DUF1759
+ More
IPR029060 PIN-like_dom_sf
IPR008737 Peptidase_asp_put
IPR001995 Peptidase_A2_cat
IPR021109 Peptidase_aspartic_dom_sf
IPR001584 Integrase_cat-core
IPR000425 MIP
IPR008042 Retrotrans_Pao
IPR012337 RNaseH-like_sf
IPR040676 DUF5641
IPR023271 Aquaporin-like
IPR041588 Integrase_H2C2
IPR036397 RNaseH_sf
IPR022048 Envelope_fusion-like
IPR000477 RT_dom
IPR029526 PGBD
IPR001969 Aspartic_peptidase_AS
IPR001878 Znf_CCHC
IPR029060 PIN-like_dom_sf
IPR008737 Peptidase_asp_put
IPR001995 Peptidase_A2_cat
IPR021109 Peptidase_aspartic_dom_sf
IPR001584 Integrase_cat-core
IPR000425 MIP
IPR008042 Retrotrans_Pao
IPR012337 RNaseH-like_sf
IPR040676 DUF5641
IPR023271 Aquaporin-like
IPR041588 Integrase_H2C2
IPR036397 RNaseH_sf
IPR022048 Envelope_fusion-like
IPR000477 RT_dom
IPR029526 PGBD
IPR001969 Aspartic_peptidase_AS
IPR001878 Znf_CCHC
Gene 3D
ProteinModelPortal
A0A2W1BPI5
A0A2W1AZ18
A0A0L7LHD6
A0A2A4JZ49
A0A2W1BMV4
A0A2A4K6T9
+ More
A0A2H1X238 A0A2W1BJC9 A0A2H1W714 A0A2H1VXK6 A0A0L7LGK2 A0A0L7KWQ7 A0A0J7N314 A0A1Y1LE43 A0A1Y1LBZ3 A0A0J7N890 A0A0J7K152 A0A0J7N5C9 A0A0J7N536 A0A0J7K907 A0A3L8DYA9 A0A1B0DAL2 A0A151WGR4 A0A0N1IHU1 A0A2H1WI74 A0A0J7KPJ2 A0A3L8D6F4 X1WYQ8 A0A0J7KB34 A0A0J7KJY9 A0A151IN79 J9LHB7 J9JNP4 A0A226DBN6 A0A0J7KCT8 A0A2S2PTX9 J9LYM7 J9K774 W8B8B3 A0A224XG39 A0A1B0CQ10 A0A151IDD4
A0A2H1X238 A0A2W1BJC9 A0A2H1W714 A0A2H1VXK6 A0A0L7LGK2 A0A0L7KWQ7 A0A0J7N314 A0A1Y1LE43 A0A1Y1LBZ3 A0A0J7N890 A0A0J7K152 A0A0J7N5C9 A0A0J7N536 A0A0J7K907 A0A3L8DYA9 A0A1B0DAL2 A0A151WGR4 A0A0N1IHU1 A0A2H1WI74 A0A0J7KPJ2 A0A3L8D6F4 X1WYQ8 A0A0J7KB34 A0A0J7KJY9 A0A151IN79 J9LHB7 J9JNP4 A0A226DBN6 A0A0J7KCT8 A0A2S2PTX9 J9LYM7 J9K774 W8B8B3 A0A224XG39 A0A1B0CQ10 A0A151IDD4
Ontologies
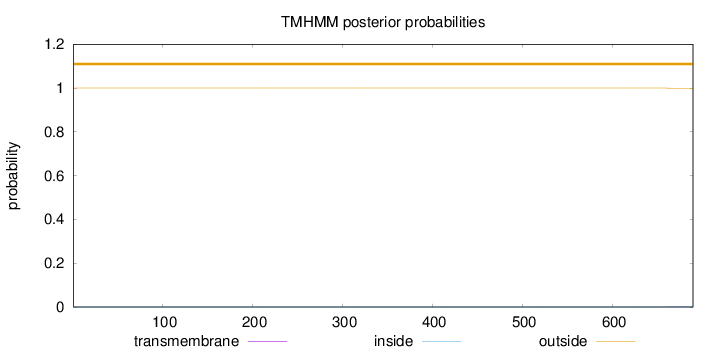
Topology
Length:
690
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.01124
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00033
outside
1 - 690
Population Genetic Test Statistics
Pi
331.250787
Theta
188.853094
Tajima's D
2.353785
CLR
1.889563
CSRT
0.933303334833258
Interpretation
Uncertain
