Gene
KWMTBOMO03669
Annotation
endonuclease_and_reverse_transcriptase-like_protein_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 1.39 PlasmaMembrane Reliability : 1.064
Sequence
CDS
ATGAAAAGGATACAAATCAATATTCTTTTGATTTGTTTGGGGGTTGGTGCTTCCCGCTTCCGTAGCAATGTCAGGGGCCTACATAGTAACCTTGACGCCGTACACCACCACCTTGAGACGGCGCAGCCTGCCCTGCTTTTTTTAACGGAGACGCAGATATCTGCTCCGGACGATACCTCATACCTTGAATACCCCGGCTACGTATTGGAGCACAACTTCCTACGTAAAGCCGGGGTGTGCGTATTCGTCCGGGCGGATGTCTGTTGTCGCCGTCTTCGGGGCCTCGAACAACGGGACCTGTCCCTATTGTGGCTGCGCGTGGATCACGGGGGCTGTATCCGAATCTACGCGTGCCTGTACAGGTCCCACAGCAGTGATGCAGGTTCAGCTCTGTTTGAGCATGTGCAAGAGGGGACTAACCGCGTGCTTGAGCAGTACCCATCTGCGGAGGTAGTGGTTCTTGGAGACTTTAACGCTCACCACCAAGAGTGGTTGGGGTCCAGAACCACTGACCTCCCGGGTCGGACTGCCTACGATTTCGCCTTGGCCTACGGCTTCTCCCAGCTGGTGACACAGCCCACCCGTGTCCCAGATATCGAGGGGCACGAGCCTTCTTTGTTGGACCTTCTGCTGACCACAGATCCGGCCGGATACAGTGTGGTGGTCGACGCTCCGCTAGGATCGTCTGATCACTGCCTTATTCGTGCTGCCGTACCACTCTCTCGTCCCGGTCGTCGTAGGACGACCGGGTATCGTAGAGTTTGGCGGTATATGTCAGCAGATTGGGATGGATTGCGTGAATTTTACGCATCCTACCCATGGGGGCGGTTCTGCTTTTCCTCTGCTGATCCTGACGTCTGTGCGGACCGTCTTAAAGACGTGGTGCTCCGGGGGATGGAATTGTTTATTCCCTCCTCTGAAGTGCCCGTTGGGGGTCGCAGCAGACCCTGGTATAACAATGCCAGTAGGGATGCTGCACACCTCAAGCGGTCCGCATACGTGGCATGGGATAATGCCAGGAGACGTCGGGATCCTAACATCTCAGGGGAAAGGCGGAAATATAACGCCGCTTCCAGGTCCTACAAGAAGGTAATTGCCAAGGCGAAGTCGGAGCACGTTGCTAGAGTTGGCGAGCGATTAAAGAGCTATCCCTCCGGGAGCCGTGCTTTTTGGTCGCTCGCCAAAGCTGCAGAAGGAAACTTTTGCAGGGTTCCGTCTTCATGGAAGACCGCCCACGTCCACCCTATCCCCAAGAAGGGTGACCGGTCGGACCCGTCGAGCTATAGGCCTATCGCGATAACTTCCTTGCTTTCCAAGGTGATGGAGCGAATAATAAATACACAACTCCTGAAGTATCTTGAGGATCGCCAGCTGATCAGTGACCGACAGTACGGTTTCCGTCACGGTCGCTCAGCTGGCGATCTTCTTGTATACCTTACTCACAGGTGGGCTGAAGCCTTGGAGAGCAAGGGCGAGGCTCTTGCTGTGAGCCTTGATATCGCGAAGGCCTTCGACAGGGTCTGGCATAGGGCACTTCTATCGAAGCTACCATCTTACGGAATCCCCGAGGGTCTCTGCAAGTGGATCGCTAGCTTTTTGGATGGGCGGAGCATCACGGTCGTTGTAGACGGTGACTGTTCTGATACCATGACCATTAACGCTGGCGTTCCACAAGGTTCGGTGCTCTCCCCCACGCTTTTCATCCTGTATATCAATGACATGCTGTCTATTGATGGCATGCATTGCTATGCGGATGACAGCACGGGGGATGCGCGATATATCGGCCATCAGAGTCTCTCTCGGAGCGCGGTGCAAGAGAGACGATCAAAACTTGTGTCTGAAGTGGAGAACTCTCTGGGGCGAGTCTCCGAATGGGGTGAATTGAACTTGGTTCAATTCAACCCGATAAAGACACAAGTTTGCGCGTTCACTGCGAAGAAGGACCCCTTTGTCATGGCGCCGCAATTCCAAGGAGTATCCCTGCAACCGTCCGAGAGTATTGGGATACTTGGGGTCGACATTTCGAGCGATGTCCAGTTTCGGAGTCATTTGGAAGGCAAAGCCAAGTTGGCGTCCAAAATGCTGGGAGTCCTCAACAGAGCGAAGCGGTACTTCACGCCTGGACAAAGGCTTTTGCTTTATAAAGCACAAGTCCGGCCTCGCGTGGAGTACTGCTCCCATCTCTGGGCCGGGGCTCCCAAATACCAGCTTCTTCCATTTGACTCCATACAGAGGAGGGCCGTTCGGATTGTCGATAATCCCATTCTCACGGATCGTTTGGAGCCTCTGGGTCTGCGGAGGGACTTCGGTTCCCTCTGTATTTTGTACCGTATGTTCCATGGGGAGTGCTCTGAGGAGTTGTTCGAGATGATACCGGCATCTCGTTTTTACCATCGCACCGCCCGCCACCGGAGTAGAGTTCATCCATACTACCTGGAGCCACTGCGGTCATCCACAGTGCGTTTCCAGAGATCTTTTTTGCCACGTACCATCCGGCTATGGAATGAGCTCCCCTCCACGGTGTTTCCTGAGCGCTATGACATGTCCTTCTTCAAACGAGGCTTGTGGAGAGTATTAAGCGGTAGGCAGCGGCTTGGCTCTGCCCCTGGCATTGCTGAAGTCCATGGGCGACGGTAA
Protein
MKRIQINILLICLGVGASRFRSNVRGLHSNLDAVHHHLETAQPALLFLTETQISAPDDTSYLEYPGYVLEHNFLRKAGVCVFVRADVCCRRLRGLEQRDLSLLWLRVDHGGCIRIYACLYRSHSSDAGSALFEHVQEGTNRVLEQYPSAEVVVLGDFNAHHQEWLGSRTTDLPGRTAYDFALAYGFSQLVTQPTRVPDIEGHEPSLLDLLLTTDPAGYSVVVDAPLGSSDHCLIRAAVPLSRPGRRRTTGYRRVWRYMSADWDGLREFYASYPWGRFCFSSADPDVCADRLKDVVLRGMELFIPSSEVPVGGRSRPWYNNASRDAAHLKRSAYVAWDNARRRRDPNISGERRKYNAASRSYKKVIAKAKSEHVARVGERLKSYPSGSRAFWSLAKAAEGNFCRVPSSWKTAHVHPIPKKGDRSDPSSYRPIAITSLLSKVMERIINTQLLKYLEDRQLISDRQYGFRHGRSAGDLLVYLTHRWAEALESKGEALAVSLDIAKAFDRVWHRALLSKLPSYGIPEGLCKWIASFLDGRSITVVVDGDCSDTMTINAGVPQGSVLSPTLFILYINDMLSIDGMHCYADDSTGDARYIGHQSLSRSAVQERRSKLVSEVENSLGRVSEWGELNLVQFNPIKTQVCAFTAKKDPFVMAPQFQGVSLQPSESIGILGVDISSDVQFRSHLEGKAKLASKMLGVLNRAKRYFTPGQRLLLYKAQVRPRVEYCSHLWAGAPKYQLLPFDSIQRRAVRIVDNPILTDRLEPLGLRRDFGSLCILYRMFHGECSEELFEMIPASRFYHRTARHRSRVHPYYLEPLRSSTVRFQRSFLPRTIRLWNELPSTVFPERYDMSFFKRGLWRVLSGRQRLGSAPGIAEVHGRR
Summary
Uniprot
ProteinModelPortal
PDB
6AR3
E-value=1.1491e-05,
Score=120
Ontologies
GO
Topology
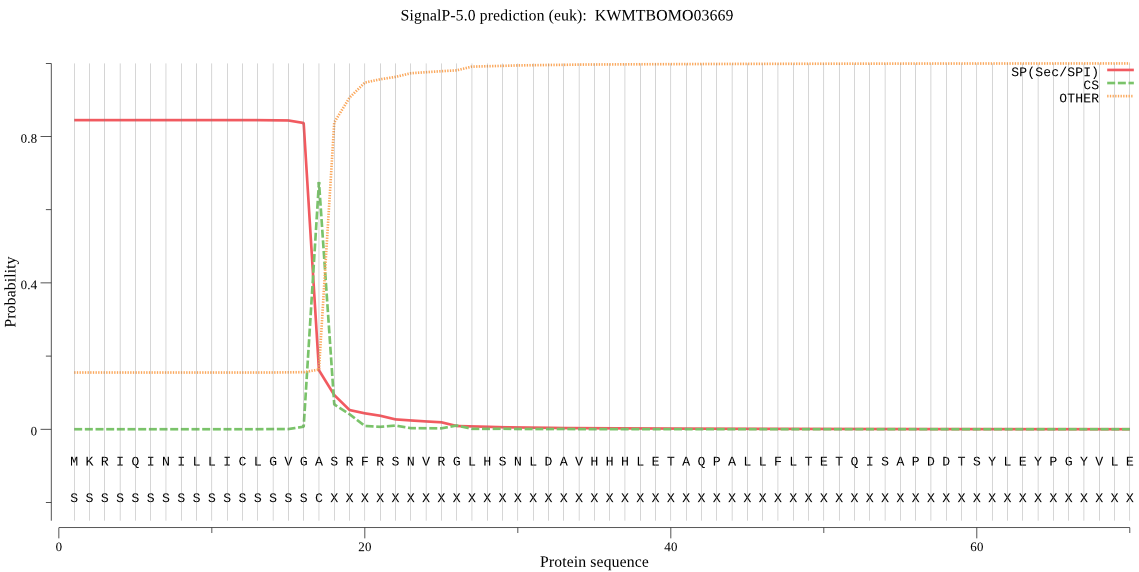
SignalP
Position: 1 - 17,
Likelihood: 0.843974
Length:
878
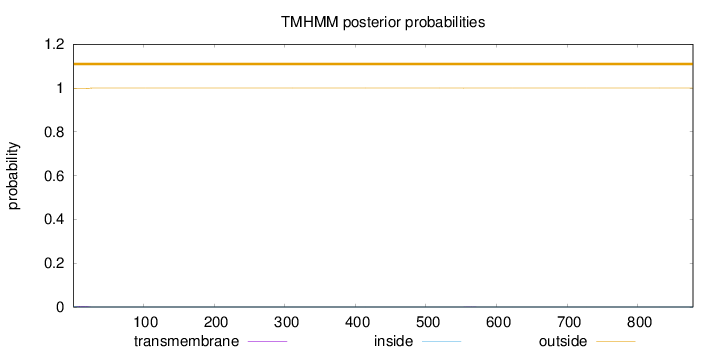
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.0507400000000001
Exp number, first 60 AAs:
0.04686
Total prob of N-in:
0.00276
outside
1 - 878
Population Genetic Test Statistics
Pi
3.592735
Theta
12.719479
Tajima's D
-1.662783
CLR
35.171225
CSRT
0.0394980250987451
Interpretation
Uncertain
