Gene
KWMTBOMO03641
Pre Gene Modal
BGIBMGA013783
Annotation
AF461149_1_transposase_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 3.253
Sequence
CDS
ATGACAGGTCAAGAAGTGGTCGACGAAGAAGTGGTGGAAGACTTAGGGCTTCGGGCATATCGAAGAAAAACAGGACATCGTTTGAATGCTCGTCTAATGGACCTGAGACTGAAGAGATGCCGCGCTTTGTTGAAGCGGTACGCGGGAAAAAGATGTCGGGAAATTCTTTTTTCGGATGAAAAAATTTTTACCTTAGAAGAGAGCTACAACAAACAAAATGATAAGGTGTACGCACACAGTAGTGAAGAAGCGAGCAACCGTATTCCGCGTGTTCAACGAGATCATTTTCCATCCTCGCTCATGGTATGGTTGGGAGTTTCTTATTGGGGCTTAACAGAGGTACATTTTTGTGAGAAAGGTGTAAAAACGAATGCAGTTGTGTATCAAAATACAGTCCTGACGAACCTTGTGGAACCTGTTTCTCATACTATGTTCAATAACAGGCACTGGGTATTCCAACAAGATTCGGCGCCAGCTCATAGAGCGAAGAGCACACAAGACTGGCTGGCGGCGCGTGAAATCGACTTCATCCGGCACGAAGACTGGCCCTCCTCCAGTCCAGATTTGAATCCGTTAGATTACAAGATATGGCAACACTTGGAGGAATAG
Protein
MTGQEVVDEEVVEDLGLRAYRRKTGHRLNARLMDLRLKRCRALLKRYAGKRCREILFSDEKIFTLEESYNKQNDKVYAHSSEEASNRIPRVQRDHFPSSLMVWLGVSYWGLTEVHFCEKGVKTNAVVYQNTVLTNLVEPVSHTMFNNRHWVFQQDSAPAHRAKSTQDWLAAREIDFIRHEDWPSSSPDLNPLDYKIWQHLEE
Summary
Similarity
Belongs to the TGF-beta family.
Uniprot
Q8ITJ9
B1Q3L9
B1Q3L5
B1Q3L8
B9A8E5
A0A0N0PF55
+ More
A0A3S2LTJ6 G3G3K1 A0A3S2M6Z0 P90693 A0A3S2LWW4 A0A3S2NF68 A0A3S2P5Q6 A0A3S2LXK8 A0A3S2M6X4 A0A3S2PAI6 A0A226D9B6 A0A3S2P4S1 A0A1I7SRX5 A0A1I7SJQ8 A0A1I7RU52 A0A0N4XXA5 W6NUW0 A0A368H7I1 A0A368FVD9 A0A016TV39 H3ETI8 H3EXA9 A0A2G5T9N6 H3E4E9 A0A016VJ32 A0A0C2DAI6 H3EE07 A0A2H1W6Q6 A0A1I8CC25 A0A016WN34 H3E1D2 H3DS69 A0A1I7T5D4 A0A2G5SF52 E3N6E8 H3FLR7 E3NJL6 A0A016TQR4 A0A016WY15 A0A016SXC9 E3ND45 E3MXK7 E3NHK7 E3N1H1 E3M897 E3N4H8 H3ESY7 A0A016WYI6 E3N749 E3M1J2 A0A1I7Z1I3 A0A183FUJ2 A0A3P8A0U6 A0A183G443 A0A3P8B6H1 E3NHD7 B6IKQ6 E3MQK2 A0A2G5TSG2 E3N9D7 A0A0N5BKM5 A0A016S9F8 A0A2G5VIT1 A0A016U6X0 A0A1L5BY18 E3MXK8 A0A183GP86 A0A3P8HGK6 A0A183F5L1 A0A3P7TF39 A0A016WKB6 E3NDP6 A0A2G5TN79 A0A0N4Y502 A0A016TQ99 A0A2G5TZM3 A0A016W3L5 A0A1I7XN71 A0A2G5TC57 T1PDL9
A0A3S2LTJ6 G3G3K1 A0A3S2M6Z0 P90693 A0A3S2LWW4 A0A3S2NF68 A0A3S2P5Q6 A0A3S2LXK8 A0A3S2M6X4 A0A3S2PAI6 A0A226D9B6 A0A3S2P4S1 A0A1I7SRX5 A0A1I7SJQ8 A0A1I7RU52 A0A0N4XXA5 W6NUW0 A0A368H7I1 A0A368FVD9 A0A016TV39 H3ETI8 H3EXA9 A0A2G5T9N6 H3E4E9 A0A016VJ32 A0A0C2DAI6 H3EE07 A0A2H1W6Q6 A0A1I8CC25 A0A016WN34 H3E1D2 H3DS69 A0A1I7T5D4 A0A2G5SF52 E3N6E8 H3FLR7 E3NJL6 A0A016TQR4 A0A016WY15 A0A016SXC9 E3ND45 E3MXK7 E3NHK7 E3N1H1 E3M897 E3N4H8 H3ESY7 A0A016WYI6 E3N749 E3M1J2 A0A1I7Z1I3 A0A183FUJ2 A0A3P8A0U6 A0A183G443 A0A3P8B6H1 E3NHD7 B6IKQ6 E3MQK2 A0A2G5TSG2 E3N9D7 A0A0N5BKM5 A0A016S9F8 A0A2G5VIT1 A0A016U6X0 A0A1L5BY18 E3MXK8 A0A183GP86 A0A3P8HGK6 A0A183F5L1 A0A3P7TF39 A0A016WKB6 E3NDP6 A0A2G5TN79 A0A0N4Y502 A0A016TQ99 A0A2G5TZM3 A0A016W3L5 A0A1I7XN71 A0A2G5TC57 T1PDL9
Pubmed
EMBL
AF461149
AAN06610.1
AB363028
BAG15927.1
AB363006
AB363010
+ More
AB363014 BAG15923.1 BAG15924.1 AB363018 BAG15926.1 AB473770 BAH20555.1 KQ459680 KPJ20472.1 RSAL01000012 RVE53510.1 JF779677 AEO90418.1 RSAL01000021 RVE52408.1 U47917 AAB47739.1 RSAL01000140 RVE46091.1 RSAL01000063 RVE49524.1 RSAL01000016 RVE53060.1 RSAL01000133 RVE46339.1 RSAL01000022 RVE52367.1 RSAL01000144 RVE45972.1 LNIX01000030 OXA41211.1 RSAL01000431 RVE41742.1 UYSL01019905 VDL71189.1 CAVP010059404 CDL95787.1 JOJR01000015 RCN51257.1 JOJR01000583 RCN36166.1 JARK01001411 EYC06502.1 PDUG01000005 PIC23776.1 JARK01001345 EYC27589.1 KN726990 KIH66733.1 ODYU01006378 SOQ48174.1 JARK01000176 EYC41234.1 PDUG01000012 PIC13506.1 DS268539 EFO88027.1 DS268747 EFP00909.1 JARK01001418 EYC05389.1 JARK01000059 EYC44525.1 JARK01001499 EYB95145.1 DS268606 EFO93488.1 DS268492 EFP11689.1 DS268679 EFO98285.1 DS268508 EFO83264.1 DS268428 EFO94379.1 DS268525 EFO85497.1 JARK01000073 EYC44053.1 DS268545 EFO88382.1 DS268421 EFO88618.1 UZAH01027257 VDO90094.1 UZAH01029309 VDP05368.1 DS268673 EFO98001.1 HE601413 CAS00486.1 DS268466 EFP06959.1 PIC29926.1 DS268565 EFO90304.1 JARK01001601 EYB87313.1 PDUG01000001 PIC51673.1 JARK01001390 EYC10681.1 KX931010 APL98300.1 EFP11674.1 UZAH01036445 VDP45493.1 UZAH01001595 VDO19836.1 JARK01000255 EYC39473.1 DS268612 EFO94085.1 PIC28734.1 UYSL01020440 VDL74616.1 JARK01001421 EYC04911.1 PDUG01000004 PIC32571.1 JARK01001337 EYC34439.1 PIC24955.1 KA646003 AFP60632.1
AB363014 BAG15923.1 BAG15924.1 AB363018 BAG15926.1 AB473770 BAH20555.1 KQ459680 KPJ20472.1 RSAL01000012 RVE53510.1 JF779677 AEO90418.1 RSAL01000021 RVE52408.1 U47917 AAB47739.1 RSAL01000140 RVE46091.1 RSAL01000063 RVE49524.1 RSAL01000016 RVE53060.1 RSAL01000133 RVE46339.1 RSAL01000022 RVE52367.1 RSAL01000144 RVE45972.1 LNIX01000030 OXA41211.1 RSAL01000431 RVE41742.1 UYSL01019905 VDL71189.1 CAVP010059404 CDL95787.1 JOJR01000015 RCN51257.1 JOJR01000583 RCN36166.1 JARK01001411 EYC06502.1 PDUG01000005 PIC23776.1 JARK01001345 EYC27589.1 KN726990 KIH66733.1 ODYU01006378 SOQ48174.1 JARK01000176 EYC41234.1 PDUG01000012 PIC13506.1 DS268539 EFO88027.1 DS268747 EFP00909.1 JARK01001418 EYC05389.1 JARK01000059 EYC44525.1 JARK01001499 EYB95145.1 DS268606 EFO93488.1 DS268492 EFP11689.1 DS268679 EFO98285.1 DS268508 EFO83264.1 DS268428 EFO94379.1 DS268525 EFO85497.1 JARK01000073 EYC44053.1 DS268545 EFO88382.1 DS268421 EFO88618.1 UZAH01027257 VDO90094.1 UZAH01029309 VDP05368.1 DS268673 EFO98001.1 HE601413 CAS00486.1 DS268466 EFP06959.1 PIC29926.1 DS268565 EFO90304.1 JARK01001601 EYB87313.1 PDUG01000001 PIC51673.1 JARK01001390 EYC10681.1 KX931010 APL98300.1 EFP11674.1 UZAH01036445 VDP45493.1 UZAH01001595 VDO19836.1 JARK01000255 EYC39473.1 DS268612 EFO94085.1 PIC28734.1 UYSL01020440 VDL74616.1 JARK01001421 EYC04911.1 PDUG01000004 PIC32571.1 JARK01001337 EYC34439.1 PIC24955.1 KA646003 AFP60632.1
Proteomes
Pfam
Interpro
IPR009057
Homeobox-like_sf
+ More
IPR038717 Tc1-like_DDE_dom
IPR001839 TGF-b_C
IPR029034 Cystine-knot_cytokine
IPR017948 TGFb_CS
IPR015615 TGF-beta-rel
IPR032171 COR
IPR020859 ROC_dom
IPR000719 Prot_kinase_dom
IPR011047 Quinoprotein_ADH-like_supfam
IPR011009 Kinase-like_dom_sf
IPR008271 Ser/Thr_kinase_AS
IPR027417 P-loop_NTPase
IPR000477 RT_dom
IPR036388 WH-like_DNA-bd_sf
IPR001523 Paired_dom
IPR025246 IS30-like_HTH
IPR038717 Tc1-like_DDE_dom
IPR001839 TGF-b_C
IPR029034 Cystine-knot_cytokine
IPR017948 TGFb_CS
IPR015615 TGF-beta-rel
IPR032171 COR
IPR020859 ROC_dom
IPR000719 Prot_kinase_dom
IPR011047 Quinoprotein_ADH-like_supfam
IPR011009 Kinase-like_dom_sf
IPR008271 Ser/Thr_kinase_AS
IPR027417 P-loop_NTPase
IPR000477 RT_dom
IPR036388 WH-like_DNA-bd_sf
IPR001523 Paired_dom
IPR025246 IS30-like_HTH
SUPFAM
Gene 3D
ProteinModelPortal
Q8ITJ9
B1Q3L9
B1Q3L5
B1Q3L8
B9A8E5
A0A0N0PF55
+ More
A0A3S2LTJ6 G3G3K1 A0A3S2M6Z0 P90693 A0A3S2LWW4 A0A3S2NF68 A0A3S2P5Q6 A0A3S2LXK8 A0A3S2M6X4 A0A3S2PAI6 A0A226D9B6 A0A3S2P4S1 A0A1I7SRX5 A0A1I7SJQ8 A0A1I7RU52 A0A0N4XXA5 W6NUW0 A0A368H7I1 A0A368FVD9 A0A016TV39 H3ETI8 H3EXA9 A0A2G5T9N6 H3E4E9 A0A016VJ32 A0A0C2DAI6 H3EE07 A0A2H1W6Q6 A0A1I8CC25 A0A016WN34 H3E1D2 H3DS69 A0A1I7T5D4 A0A2G5SF52 E3N6E8 H3FLR7 E3NJL6 A0A016TQR4 A0A016WY15 A0A016SXC9 E3ND45 E3MXK7 E3NHK7 E3N1H1 E3M897 E3N4H8 H3ESY7 A0A016WYI6 E3N749 E3M1J2 A0A1I7Z1I3 A0A183FUJ2 A0A3P8A0U6 A0A183G443 A0A3P8B6H1 E3NHD7 B6IKQ6 E3MQK2 A0A2G5TSG2 E3N9D7 A0A0N5BKM5 A0A016S9F8 A0A2G5VIT1 A0A016U6X0 A0A1L5BY18 E3MXK8 A0A183GP86 A0A3P8HGK6 A0A183F5L1 A0A3P7TF39 A0A016WKB6 E3NDP6 A0A2G5TN79 A0A0N4Y502 A0A016TQ99 A0A2G5TZM3 A0A016W3L5 A0A1I7XN71 A0A2G5TC57 T1PDL9
A0A3S2LTJ6 G3G3K1 A0A3S2M6Z0 P90693 A0A3S2LWW4 A0A3S2NF68 A0A3S2P5Q6 A0A3S2LXK8 A0A3S2M6X4 A0A3S2PAI6 A0A226D9B6 A0A3S2P4S1 A0A1I7SRX5 A0A1I7SJQ8 A0A1I7RU52 A0A0N4XXA5 W6NUW0 A0A368H7I1 A0A368FVD9 A0A016TV39 H3ETI8 H3EXA9 A0A2G5T9N6 H3E4E9 A0A016VJ32 A0A0C2DAI6 H3EE07 A0A2H1W6Q6 A0A1I8CC25 A0A016WN34 H3E1D2 H3DS69 A0A1I7T5D4 A0A2G5SF52 E3N6E8 H3FLR7 E3NJL6 A0A016TQR4 A0A016WY15 A0A016SXC9 E3ND45 E3MXK7 E3NHK7 E3N1H1 E3M897 E3N4H8 H3ESY7 A0A016WYI6 E3N749 E3M1J2 A0A1I7Z1I3 A0A183FUJ2 A0A3P8A0U6 A0A183G443 A0A3P8B6H1 E3NHD7 B6IKQ6 E3MQK2 A0A2G5TSG2 E3N9D7 A0A0N5BKM5 A0A016S9F8 A0A2G5VIT1 A0A016U6X0 A0A1L5BY18 E3MXK8 A0A183GP86 A0A3P8HGK6 A0A183F5L1 A0A3P7TF39 A0A016WKB6 E3NDP6 A0A2G5TN79 A0A0N4Y502 A0A016TQ99 A0A2G5TZM3 A0A016W3L5 A0A1I7XN71 A0A2G5TC57 T1PDL9
PDB
5CR4
E-value=7.65949e-08,
Score=132
Ontologies
KEGG
PANTHER
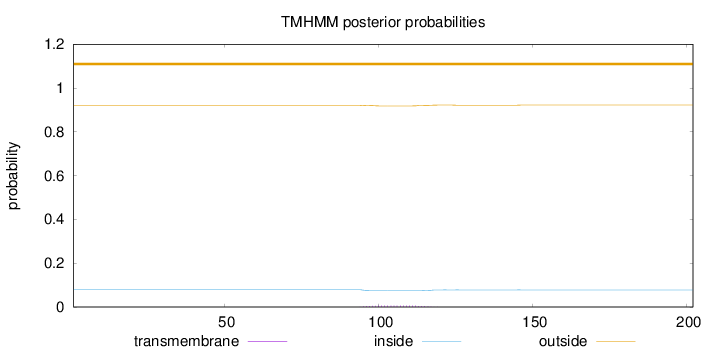
Topology
Subcellular location
Secreted
Length:
202
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.12676
Exp number, first 60 AAs:
0
Total prob of N-in:
0.07904
outside
1 - 202
Population Genetic Test Statistics
Pi
7.535108
Theta
11.554557
Tajima's D
-0.595975
CLR
22.64485
CSRT
0.216589170541473
Interpretation
Uncertain
