Gene
KWMTBOMO03564
Pre Gene Modal
BGIBMGA002389
Annotation
PREDICTED:_mucin-2-like_[Amyelois_transitella]
Location in the cell
Nuclear Reliability : 2.495
Sequence
CDS
ATGGCTGGAAGTTACATTGTCAACGTGCCTAAATTGAAAGGTCGCGAAAACTTCGACGAGTGGGCGTTTGCAGCTGAACATTTTTTGATGTTGGAAGGTGTCGATATTCATAAGTTAGAATCATCAACCGATGCTAGTTCTATTGATGAAAGAAAAGCTAAAGCTAAACTTATTATGACAATTGACTCTTCAATTTATGTTCACATTAAAAACGAGCAAACAGTACAAGGTGTGTGGAAGCGTTTAAAAGCCCTGTACGATGACTGTGGCTTTACGCGTCGCATCAGTCTACTTCGTCAGCTAATATCAATTCGATTAGAAGATAGCGAATCAATGGAAGCATATGTAACGCAAATGGTCGACACTGCACAAAAACTTAAAGGTACGGGTTTTCAGATCACTGATGAGTGGATCGGTTCACTATTATTAGCGGGCCTAACAGATACATTTGCTCCTATGATAATGGCCATTGAACATTCTGGATTAGCCATCAGTACGGATGCCATCAAGTCAAAATTACTGGACATGAGTTCAGAAGTTGGCAACACTGCAACTAACAATGCCTTTTTGTCGAAAGTAAGTCAGATGATTCGTAATGGAAACAAAGTGAGCTTCGAAGAAGACGTTTGTTACATACGCAATCGACAGAATGTGCTGGTAGGAAAAGCAAACCTGGTGAATGGAGTCTATAAATTGGACATTAAGTCAGACAGTTTGTTCGCGTGCTCAGCCACAACTGTGACTAGTGAGATATGGCATCGAAGATTAGGACACATTAACAGTAGAAGCTTGAATATTATGAAGAATGGTGCTGTGGATGGGATATCATACATGGATAAAGCTGAAATTGATATGTCCAAGTGCATCATATGCTGTGAAGGAAAACAGTCAAGGTTGCCATTTTCACATAAAGGTACGAGAAGTGAAAATGTGCTTGAACTTGTTCATGCAGATGTCTGTGGACCCATGGAACAACCTTCTATTGGACTATCAAAGTATTTTTTGCTTTTTGTAGATGACTACAGTAGAATGGCATTTGTGTTTTTTCTCAAAGCCAAGAACCAAGTTTTCAAATATTTTAAAGAGTTCAAAACTCTGGTTGAGACACAGACAGGCAGAAAACTAAAAGCCTTAAGGACAGATAATGGTGGAGAGTTTTGTTCGCTAGAATTTGATGAGTTTTTGAAGAAAAATGGCATAGTACATCAAACAACAAACCCTCATACACCAGAACAGAATGGCCTGTGTGAAAGACTTAATAGAACTGTCGTGGAGAGAGCTCGGTGTCTTCTGTTTGAAGCAGGCCTAGATAAAAAGTTTTGGGCCGAAGCAATCAACACAGCAGTCTATTTACGAAATAGAACATTGGCAGCAGGACTTCAGAAAACACCTTATGAAATATGGAGTGGTAAAAAGCCAAATCTAAGTCATATACGGATATTTGGTAGTCCTGCAATGGTCCATGTCCCCAAAGCAAAGCGGACTAAATGGAACAAGAAGGCCAATGAACACATCCTGATTGGATACTCAGAGAGTACCAAAGGATACAGGCTGTACAATGTAAAAACAAAAAATATTACCACTAGCAGGGATGTTGTTATCATGGAGAAAATCAACGACACAGATATGATACAAGCAGTAGTGGAAGAAAAAGAAACTATTCCAAGTGAAGAAGGAATACAAAAATGTGCTGTGACAAGTGAAGATTCCACTTGTGACTCACTAAATAGCACATACATGACAGATGATGCTACATATGTACCAGATGATACCAGTACCACCAGCTCAGATGAATTCTGTGATAGTAAATCGAATCACAAGTCAGAAAATCTGCAGCCAATAATTAAGGTGGAAAAGAGGACTAGGAAGAAACCAGACTGGTATGGGTACAATAACATATGTCAGAGCAGTGAGGAGGTGGACGACAATCTTCTCACTTATGAGGATGTAATGAGAGGATCAGAACAAGAAGAATGGTCAAAAGCCATGAAAGAGGAAATACAATCTTTCCATCAGAATGATGCTTGGGAGTTAGTAGACAGACCCAGTGGTGAAAATATTGTGGCATGCAAATGGGTGTATAAAAAGAAACTTAACAGTGACAACTCAGTGAGATATAGGGCACGGTTGGTTGCGAAGGGTTTCAGCCAGAGACAGGGTATAGACTTTAAAGAAACTTTTTCCCCAGTGCTAAAATATTCCACTTTAAAACTGTTATTTGCTATTAGTGTTAACTTGAATCTTAGCATAACCCATTTGGATGTAACGACGGCCTTTTTGAATGGTCACCTAGATGAAACTGTTTTTATGCAATTGCCTCAGAGTTTTGAATGTAAACCTAATGCTGATAAAGTCTTGAAATTGAAAAAGGCCATTTATGGACTAAAACAGTCAGCGAGGGCCTGGCGAATTTCGACTACCGGGGGACCACTAGCCCTTCTACAGGCGCATGATGTGGTCGCCAGAGAAGTGTACGGTGAGGAGTCATTGCGTGCCTCTCCGCCGCCGGTCAACGGTCGCACGACGGCACCGCCGCCCGAAGAGGAAGACGACGCCGAACCCGAATTCGAGAACGTGACCAGGGTGCGCCTCGTGCAGTTCCAGAAAAACACCGACGAGCCAATGGGAATTACTCTGAAACTAGCGGACGACGGACGTTGCATCGTTGCTCGTATTATGCACGGCGGTATGATACATCGCCAGGCGACGCTGCATGTCGGTGACGAGATTAAAGAGATTAACGGAACGCCTGTCGCAAATCAATCTGTCGCTCAACTACAAAGAATGCTGCGCGAGGCTCGCGGCTCCGTCACTTTCAAGATCGTACCTTCCTACCGCTCGGCACCACCACCCTGCGAGCTTTTCCGGATAAAGCCTTCGCCCGTGCTAATATTCGTTCGAGCGCAGTTTGATTACGATCCCTTGGAAGACGAACTAATCCCGTGTGCTCAAGCAGGCATCGCCTTCAACACCGGCGACATATTGCAGGTTAGTTGTGTATTATGGGTATAA
Protein
MAGSYIVNVPKLKGRENFDEWAFAAEHFLMLEGVDIHKLESSTDASSIDERKAKAKLIMTIDSSIYVHIKNEQTVQGVWKRLKALYDDCGFTRRISLLRQLISIRLEDSESMEAYVTQMVDTAQKLKGTGFQITDEWIGSLLLAGLTDTFAPMIMAIEHSGLAISTDAIKSKLLDMSSEVGNTATNNAFLSKVSQMIRNGNKVSFEEDVCYIRNRQNVLVGKANLVNGVYKLDIKSDSLFACSATTVTSEIWHRRLGHINSRSLNIMKNGAVDGISYMDKAEIDMSKCIICCEGKQSRLPFSHKGTRSENVLELVHADVCGPMEQPSIGLSKYFLLFVDDYSRMAFVFFLKAKNQVFKYFKEFKTLVETQTGRKLKALRTDNGGEFCSLEFDEFLKKNGIVHQTTNPHTPEQNGLCERLNRTVVERARCLLFEAGLDKKFWAEAINTAVYLRNRTLAAGLQKTPYEIWSGKKPNLSHIRIFGSPAMVHVPKAKRTKWNKKANEHILIGYSESTKGYRLYNVKTKNITTSRDVVIMEKINDTDMIQAVVEEKETIPSEEGIQKCAVTSEDSTCDSLNSTYMTDDATYVPDDTSTTSSDEFCDSKSNHKSENLQPIIKVEKRTRKKPDWYGYNNICQSSEEVDDNLLTYEDVMRGSEQEEWSKAMKEEIQSFHQNDAWELVDRPSGENIVACKWVYKKKLNSDNSVRYRARLVAKGFSQRQGIDFKETFSPVLKYSTLKLLFAISVNLNLSITHLDVTTAFLNGHLDETVFMQLPQSFECKPNADKVLKLKKAIYGLKQSARAWRISTTGGPLALLQAHDVVAREVYGEESLRASPPPVNGRTTAPPPEEEDDAEPEFENVTRVRLVQFQKNTDEPMGITLKLADDGRCIVARIMHGGMIHRQATLHVGDEIKEINGTPVANQSVAQLQRMLREARGSVTFKIVPSYRSAPPPCELFRIKPSPVLIFVRAQFDYDPLEDELIPCAQAGIAFNTGDILQVSCVLWV
Summary
Uniprot
EMBL
Proteomes
Interpro
SUPFAM
SSF53098
SSF53098
Gene 3D
ProteinModelPortal
PDB
1KWA
E-value=4.4075e-36,
Score=383
Ontologies
GO
PANTHER
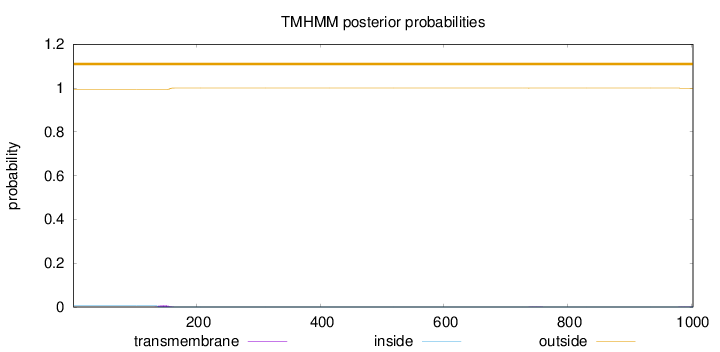
Topology
Length:
1003
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.13334
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00599
outside
1 - 1003
Population Genetic Test Statistics
Pi
230.645231
Theta
165.69263
Tajima's D
0.830504
CLR
0.525304
CSRT
0.608269586520674
Interpretation
Uncertain
