Gene
KWMTBOMO03522 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA006246
Annotation
PREDICTED:_NADP-dependent_malic_enzyme_[Bombyx_mori]
Full name
Malic enzyme
Location in the cell
Mitochondrial Reliability : 1.299
Sequence
CDS
ATGGAGGCCTGTGTACAGAGATACGGACAGAACACTCTGCTCCAATTTGAGGATTTCGCCCTTCCGAATGCCGGTCGGCTCCTGAAGAAATACCGCAATAAATACTGCACGTTCAACGACGACATCCAAGGTACGGCCAGCGTTGCCGTCGCCGGTCTGATGGCGGCCGTCAGGGTCACGAGGAGGAAGCTCAGTGAGAACATTTACCTTTTCCTCGGTGCTGGATCTGCAGCGAACGGCATCGCGAACTTGACTGTGGCCGCCATGATGGCGGACGGACTCTCCGAGAGGCAGGCCCGCGAGAGAGTGTACATGTTCGACGTCGACGGACTATTGTCCACTCGACGTGAAGGAGGGGTTCCGGAGCACGCCAAGGCCTTCGGGAAGGATATCGAACCGGAGAAAGACTTCGAGGCCTGCGTTGCCAAGATCAAGCCTAGCTGTCTTATTGGTTGTTCAACGGTGGGCGGCGCTTTCACGCCCAATGTATTGAAGCAAATGGCGATGAACACCGAGCGACCCGTCATATTCGCACTCTCGAATCCGACCAGTAAAGCTGAGTGCACAGCCCAAGACGCCTACGACCATACGGAAGGTCGTTGCATATTCGCGTCCGGCTCCCCGTTCCCGCCGGTGATGTACAGCGGCCGGGAGTACCGCCCGGGACAGGGCAACAACGCCTACATCTTCCCCGGTGTGGCGCTCGGCGTCATCGCCACGGCCACGCATCACATCTCCGAGAATATGTTCCTGACGGCGGCCAGGACTTTGGCGAACTACGTAAGCGAGTCGGATCTATCCATCGGTCGCATTTATCCGTCGCTGGCTGAAGTGAAAGAAGTGTCCGTCGCTATCGCCATCGAAGTCGCTAAGACCGCATACGACGAAGGTCTTGCCTCAGTGTATCCAGAGCCTCGCGATCTACCGAAGCACGTACAAAACCAAATGTACAGCTATTATTACGAATCTGCAATGCCGGTGACCTGGGATTGGAAGCCCGAGGAGAAGTTCAACGTTAGACCCATCCAACCTGTACCGAAGAACATTTAA
Protein
MEACVQRYGQNTLLQFEDFALPNAGRLLKKYRNKYCTFNDDIQGTASVAVAGLMAAVRVTRRKLSENIYLFLGAGSAANGIANLTVAAMMADGLSERQARERVYMFDVDGLLSTRREGGVPEHAKAFGKDIEPEKDFEACVAKIKPSCLIGCSTVGGAFTPNVLKQMAMNTERPVIFALSNPTSKAECTAQDAYDHTEGRCIFASGSPFPPVMYSGREYRPGQGNNAYIFPGVALGVIATATHHISENMFLTAARTLANYVSESDLSIGRIYPSLAEVKEVSVAIAIEVAKTAYDEGLASVYPEPRDLPKHVQNQMYSYYYESAMPVTWDWKPEEKFNVRPIQPVPKNI
Summary
Similarity
Belongs to the malic enzymes family.
Uniprot
H9J9Q3
A0A194QVQ3
A0A212F922
A0A2H1V9X6
A0A2A4JIT3
A0A194QBR2
+ More
A0A2W1BWB1 A0A194QQ65 A0A194QAH0 A0A182GYQ8 A0A182H086 A0A023EXF3 A0A1S4EXR8 Q17M99 A0A0K8WJK7 B0WQC9 A0A2M4ABQ9 W5JPA9 A0A2M4ABS9 A0A2M4BHA3 A0A182RYM9 A0A1B6C7V3 A0A2M4BHC5 A0A182W1I7 A0A182XJ28 A0A182V0I8 A0A182I7T1 A0A1Q3FI86 A0A0K8WI45 A0A0K8UER4 A0A182TL51 A0A2M3Z470 A0A182R0A2 A0A2M3Z419 W8BCJ7 A0A182K1D7 W8ASH1 A0A182FPU6 A0A182YGM6 A0A182LYI4 A0A0A1WP91 A0A0A1X9J1 Q7QB64 A0A182L7C6 F5HL44 T1PDU2 T1PDX1 A0A1I8NSM9 A0A2H1V9T4 A0A182N5S8 A0A1B6DG24 A0A1B6JHS1 A0A182IQ03 A0A1B6FNU9 A0A224XNH8 A0A0P4VPB6 A0A023F4G0 U5EUA8 A0A0L0BYF3 A0A0C9R9Z4 U4UQ11 B3P850 A0A1W4UH31 A0A1W4UIU7 T1I5N5 A0A0R1E5P7 A0A0V0GBL2 A0A0P8XU46 B4PRU5 E1JIZ5 E1JIZ4 B3MT32 A0A0M5J924 A0A1A9YSV7 B4IC50 A0A034V722 Q9VB69 A0A1A9ZZ14 A0A1B0FEH5 A0A0Q9WG72 A0A0Q9WYS2 B4KAX8 A0A0K8TU30 B4LXG6 D6WC30 A0A1A9VFY1 A0A182P1Q3 B4JU20 A0A3B0JE23 A0A3B0KAI4 A0A3B0K3M4 A0A1A9W7B7 A0A0J7KN64 A0A2J7Q043 V9IAE8 V9I857 A0A2W1B8P7 A0A0A9YR05 A0A195D5H8 A0A0Q9WT09
A0A2W1BWB1 A0A194QQ65 A0A194QAH0 A0A182GYQ8 A0A182H086 A0A023EXF3 A0A1S4EXR8 Q17M99 A0A0K8WJK7 B0WQC9 A0A2M4ABQ9 W5JPA9 A0A2M4ABS9 A0A2M4BHA3 A0A182RYM9 A0A1B6C7V3 A0A2M4BHC5 A0A182W1I7 A0A182XJ28 A0A182V0I8 A0A182I7T1 A0A1Q3FI86 A0A0K8WI45 A0A0K8UER4 A0A182TL51 A0A2M3Z470 A0A182R0A2 A0A2M3Z419 W8BCJ7 A0A182K1D7 W8ASH1 A0A182FPU6 A0A182YGM6 A0A182LYI4 A0A0A1WP91 A0A0A1X9J1 Q7QB64 A0A182L7C6 F5HL44 T1PDU2 T1PDX1 A0A1I8NSM9 A0A2H1V9T4 A0A182N5S8 A0A1B6DG24 A0A1B6JHS1 A0A182IQ03 A0A1B6FNU9 A0A224XNH8 A0A0P4VPB6 A0A023F4G0 U5EUA8 A0A0L0BYF3 A0A0C9R9Z4 U4UQ11 B3P850 A0A1W4UH31 A0A1W4UIU7 T1I5N5 A0A0R1E5P7 A0A0V0GBL2 A0A0P8XU46 B4PRU5 E1JIZ5 E1JIZ4 B3MT32 A0A0M5J924 A0A1A9YSV7 B4IC50 A0A034V722 Q9VB69 A0A1A9ZZ14 A0A1B0FEH5 A0A0Q9WG72 A0A0Q9WYS2 B4KAX8 A0A0K8TU30 B4LXG6 D6WC30 A0A1A9VFY1 A0A182P1Q3 B4JU20 A0A3B0JE23 A0A3B0KAI4 A0A3B0K3M4 A0A1A9W7B7 A0A0J7KN64 A0A2J7Q043 V9IAE8 V9I857 A0A2W1B8P7 A0A0A9YR05 A0A195D5H8 A0A0Q9WT09
Pubmed
19121390
26354079
22118469
28756777
26483478
24945155
+ More
17510324 20920257 23761445 24495485 25244985 25830018 12364791 14747013 17210077 20966253 25315136 27129103 25474469 26108605 23537049 17994087 17550304 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 25348373 26109357 26109356 26369729 18362917 19820115 25401762 26823975
17510324 20920257 23761445 24495485 25244985 25830018 12364791 14747013 17210077 20966253 25315136 27129103 25474469 26108605 23537049 17994087 17550304 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 25348373 26109357 26109356 26369729 18362917 19820115 25401762 26823975
EMBL
BABH01034014
BABH01034015
BABH01034016
BABH01034017
KQ461181
KPJ07636.1
+ More
AGBW02009653 OWR50228.1 ODYU01001426 SOQ37596.1 NWSH01001337 PCG71618.1 KQ459249 KPJ02430.1 KZ149899 PZC78631.1 KPJ07637.1 KPJ02429.1 JXUM01021496 JXUM01021497 JXUM01021498 KQ560602 KXJ81680.1 JXUM01100888 JXUM01100889 KQ564606 KXJ72035.1 GAPW01000494 JAC13104.1 CH477207 EAT47850.1 GDHF01000971 JAI51343.1 DS232039 EDS32800.1 GGFK01004902 MBW38223.1 ADMH02000944 ETN64604.1 GGFK01004916 MBW38237.1 GGFJ01003263 MBW52404.1 GEDC01027794 JAS09504.1 GGFJ01003314 MBW52455.1 APCN01002552 GFDL01007813 JAV27232.1 GDHF01001483 JAI50831.1 GDHF01027238 JAI25076.1 GGFM01002542 MBW23293.1 AXCN02001157 GGFM01002520 MBW23271.1 GAMC01019021 JAB87534.1 GAMC01019022 JAB87533.1 AXCM01000270 GBXI01013791 JAD00501.1 GBXI01006333 JAD07959.1 AAAB01008880 EAA08510.5 EGK97005.1 KA646942 AFP61571.1 KA646941 AFP61570.1 SOQ37595.1 GEDC01012664 JAS24634.1 GECU01008999 JAS98707.1 GECZ01017897 JAS51872.1 GFTR01006845 JAW09581.1 GDKW01003589 JAI53006.1 GBBI01002297 JAC16415.1 GANO01001567 JAB58304.1 JRES01001160 KNC25050.1 GBYB01009772 JAG79539.1 KB632397 ERL94598.1 CH954182 EDV53454.1 ACPB03014452 CM000160 KRK04335.1 GECL01000725 JAP05399.1 CH902623 KPU72841.1 EDW98538.1 AE014297 ACZ95043.1 ACZ95044.1 EDV30422.1 CP012526 ALC46036.1 CH480828 EDW45208.1 GAKP01020653 JAC38299.1 AY051465 AAF56674.1 AAK92889.1 CCAG010001986 CH940650 KRF83517.1 CH933806 KRG01135.1 EDW14656.1 GDAI01000138 JAI17465.1 EDW67844.1 KQ971310 EEZ99125.1 CH916374 EDV91599.1 OUUW01000005 SPP80577.1 SPP80578.1 SPP80579.1 LBMM01005231 KMQ91684.1 NEVH01020327 PNF21946.1 JR036673 AEY57309.1 JR036672 AEY57308.1 KZ150227 PZC72042.1 GBHO01011634 GBHO01009599 GDHC01004108 JAG31970.1 JAG34005.1 JAQ14521.1 KQ976870 KYN07694.1 CH964232 KRF99390.1
AGBW02009653 OWR50228.1 ODYU01001426 SOQ37596.1 NWSH01001337 PCG71618.1 KQ459249 KPJ02430.1 KZ149899 PZC78631.1 KPJ07637.1 KPJ02429.1 JXUM01021496 JXUM01021497 JXUM01021498 KQ560602 KXJ81680.1 JXUM01100888 JXUM01100889 KQ564606 KXJ72035.1 GAPW01000494 JAC13104.1 CH477207 EAT47850.1 GDHF01000971 JAI51343.1 DS232039 EDS32800.1 GGFK01004902 MBW38223.1 ADMH02000944 ETN64604.1 GGFK01004916 MBW38237.1 GGFJ01003263 MBW52404.1 GEDC01027794 JAS09504.1 GGFJ01003314 MBW52455.1 APCN01002552 GFDL01007813 JAV27232.1 GDHF01001483 JAI50831.1 GDHF01027238 JAI25076.1 GGFM01002542 MBW23293.1 AXCN02001157 GGFM01002520 MBW23271.1 GAMC01019021 JAB87534.1 GAMC01019022 JAB87533.1 AXCM01000270 GBXI01013791 JAD00501.1 GBXI01006333 JAD07959.1 AAAB01008880 EAA08510.5 EGK97005.1 KA646942 AFP61571.1 KA646941 AFP61570.1 SOQ37595.1 GEDC01012664 JAS24634.1 GECU01008999 JAS98707.1 GECZ01017897 JAS51872.1 GFTR01006845 JAW09581.1 GDKW01003589 JAI53006.1 GBBI01002297 JAC16415.1 GANO01001567 JAB58304.1 JRES01001160 KNC25050.1 GBYB01009772 JAG79539.1 KB632397 ERL94598.1 CH954182 EDV53454.1 ACPB03014452 CM000160 KRK04335.1 GECL01000725 JAP05399.1 CH902623 KPU72841.1 EDW98538.1 AE014297 ACZ95043.1 ACZ95044.1 EDV30422.1 CP012526 ALC46036.1 CH480828 EDW45208.1 GAKP01020653 JAC38299.1 AY051465 AAF56674.1 AAK92889.1 CCAG010001986 CH940650 KRF83517.1 CH933806 KRG01135.1 EDW14656.1 GDAI01000138 JAI17465.1 EDW67844.1 KQ971310 EEZ99125.1 CH916374 EDV91599.1 OUUW01000005 SPP80577.1 SPP80578.1 SPP80579.1 LBMM01005231 KMQ91684.1 NEVH01020327 PNF21946.1 JR036673 AEY57309.1 JR036672 AEY57308.1 KZ150227 PZC72042.1 GBHO01011634 GBHO01009599 GDHC01004108 JAG31970.1 JAG34005.1 JAQ14521.1 KQ976870 KYN07694.1 CH964232 KRF99390.1
Proteomes
UP000005204
UP000053240
UP000007151
UP000218220
UP000053268
UP000069940
+ More
UP000249989 UP000008820 UP000002320 UP000000673 UP000075900 UP000075920 UP000076407 UP000075903 UP000075840 UP000075902 UP000075886 UP000075881 UP000069272 UP000076408 UP000075883 UP000007062 UP000075882 UP000095301 UP000095300 UP000075884 UP000075880 UP000037069 UP000030742 UP000008711 UP000192221 UP000015103 UP000002282 UP000007801 UP000000803 UP000092553 UP000092443 UP000001292 UP000092445 UP000092444 UP000008792 UP000009192 UP000007266 UP000078200 UP000075885 UP000001070 UP000268350 UP000091820 UP000036403 UP000235965 UP000078542 UP000007798
UP000249989 UP000008820 UP000002320 UP000000673 UP000075900 UP000075920 UP000076407 UP000075903 UP000075840 UP000075902 UP000075886 UP000075881 UP000069272 UP000076408 UP000075883 UP000007062 UP000075882 UP000095301 UP000095300 UP000075884 UP000075880 UP000037069 UP000030742 UP000008711 UP000192221 UP000015103 UP000002282 UP000007801 UP000000803 UP000092553 UP000092443 UP000001292 UP000092445 UP000092444 UP000008792 UP000009192 UP000007266 UP000078200 UP000075885 UP000001070 UP000268350 UP000091820 UP000036403 UP000235965 UP000078542 UP000007798
Interpro
Gene 3D
CDD
ProteinModelPortal
H9J9Q3
A0A194QVQ3
A0A212F922
A0A2H1V9X6
A0A2A4JIT3
A0A194QBR2
+ More
A0A2W1BWB1 A0A194QQ65 A0A194QAH0 A0A182GYQ8 A0A182H086 A0A023EXF3 A0A1S4EXR8 Q17M99 A0A0K8WJK7 B0WQC9 A0A2M4ABQ9 W5JPA9 A0A2M4ABS9 A0A2M4BHA3 A0A182RYM9 A0A1B6C7V3 A0A2M4BHC5 A0A182W1I7 A0A182XJ28 A0A182V0I8 A0A182I7T1 A0A1Q3FI86 A0A0K8WI45 A0A0K8UER4 A0A182TL51 A0A2M3Z470 A0A182R0A2 A0A2M3Z419 W8BCJ7 A0A182K1D7 W8ASH1 A0A182FPU6 A0A182YGM6 A0A182LYI4 A0A0A1WP91 A0A0A1X9J1 Q7QB64 A0A182L7C6 F5HL44 T1PDU2 T1PDX1 A0A1I8NSM9 A0A2H1V9T4 A0A182N5S8 A0A1B6DG24 A0A1B6JHS1 A0A182IQ03 A0A1B6FNU9 A0A224XNH8 A0A0P4VPB6 A0A023F4G0 U5EUA8 A0A0L0BYF3 A0A0C9R9Z4 U4UQ11 B3P850 A0A1W4UH31 A0A1W4UIU7 T1I5N5 A0A0R1E5P7 A0A0V0GBL2 A0A0P8XU46 B4PRU5 E1JIZ5 E1JIZ4 B3MT32 A0A0M5J924 A0A1A9YSV7 B4IC50 A0A034V722 Q9VB69 A0A1A9ZZ14 A0A1B0FEH5 A0A0Q9WG72 A0A0Q9WYS2 B4KAX8 A0A0K8TU30 B4LXG6 D6WC30 A0A1A9VFY1 A0A182P1Q3 B4JU20 A0A3B0JE23 A0A3B0KAI4 A0A3B0K3M4 A0A1A9W7B7 A0A0J7KN64 A0A2J7Q043 V9IAE8 V9I857 A0A2W1B8P7 A0A0A9YR05 A0A195D5H8 A0A0Q9WT09
A0A2W1BWB1 A0A194QQ65 A0A194QAH0 A0A182GYQ8 A0A182H086 A0A023EXF3 A0A1S4EXR8 Q17M99 A0A0K8WJK7 B0WQC9 A0A2M4ABQ9 W5JPA9 A0A2M4ABS9 A0A2M4BHA3 A0A182RYM9 A0A1B6C7V3 A0A2M4BHC5 A0A182W1I7 A0A182XJ28 A0A182V0I8 A0A182I7T1 A0A1Q3FI86 A0A0K8WI45 A0A0K8UER4 A0A182TL51 A0A2M3Z470 A0A182R0A2 A0A2M3Z419 W8BCJ7 A0A182K1D7 W8ASH1 A0A182FPU6 A0A182YGM6 A0A182LYI4 A0A0A1WP91 A0A0A1X9J1 Q7QB64 A0A182L7C6 F5HL44 T1PDU2 T1PDX1 A0A1I8NSM9 A0A2H1V9T4 A0A182N5S8 A0A1B6DG24 A0A1B6JHS1 A0A182IQ03 A0A1B6FNU9 A0A224XNH8 A0A0P4VPB6 A0A023F4G0 U5EUA8 A0A0L0BYF3 A0A0C9R9Z4 U4UQ11 B3P850 A0A1W4UH31 A0A1W4UIU7 T1I5N5 A0A0R1E5P7 A0A0V0GBL2 A0A0P8XU46 B4PRU5 E1JIZ5 E1JIZ4 B3MT32 A0A0M5J924 A0A1A9YSV7 B4IC50 A0A034V722 Q9VB69 A0A1A9ZZ14 A0A1B0FEH5 A0A0Q9WG72 A0A0Q9WYS2 B4KAX8 A0A0K8TU30 B4LXG6 D6WC30 A0A1A9VFY1 A0A182P1Q3 B4JU20 A0A3B0JE23 A0A3B0KAI4 A0A3B0K3M4 A0A1A9W7B7 A0A0J7KN64 A0A2J7Q043 V9IAE8 V9I857 A0A2W1B8P7 A0A0A9YR05 A0A195D5H8 A0A0Q9WT09
PDB
1O0S
E-value=1.40776e-100,
Score=935
Ontologies
PATHWAY
GO
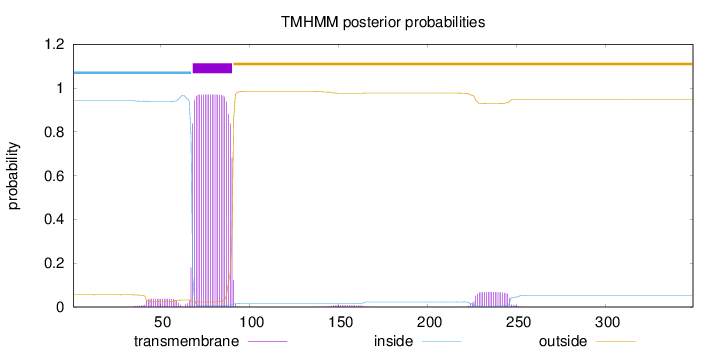
Topology
Length:
349
Number of predicted TMHs:
1
Exp number of AAs in TMHs:
24.22431
Exp number, first 60 AAs:
0.70531
Total prob of N-in:
0.94437
inside
1 - 67
TMhelix
68 - 90
outside
91 - 349
Population Genetic Test Statistics
Pi
199.659644
Theta
175.473868
Tajima's D
1.19624
CLR
0.909081
CSRT
0.710114494275286
Interpretation
Uncertain
