Gene
KWMTBOMO03395
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 1.852 Nuclear Reliability : 1.344
Sequence
CDS
ATGATTGCACACTGCAGAACCAGGCCCCTAGTCCCCGGCGGCCCCGCTATTTCTGGTTACGGTAGCGGCCATGGTAACGGCGGGGCAAGGGGTGCTAAGAATCCCCGGCAGAGATCCGGCTACCACCCCCCTTCACAACGACAACTGGCCCTGGTGACATACAACGTGCGCACTTTGAGGTTGGACGAGAAGCTAATAGAGCTGGAAGAAGAAATGAACAAGTTGCGCTGGGACGTTATTGGTTTATCGGAGATCCGAAGAGAGGGGGAGGATACGAAAACCTTACAGTCCGGCCACTTGTTCTACTACCGGGAGGGAGAACAGAAGTCCCAAGGTGGTGTTGGGTTTATCGTCCACAAATCTCTCATAAACAACATTGTGGCCATCGAAAGTGTGTCGAGTAGGGTAGTATACCTTGTCCTCAAAATCTCAAAGCGGTATTCGCTGAAGATTGTACAGGTATACGCCCCATCGACGTCACATCCAGACGATGAAGTGGAAGCTACGTATGAGGACATTAGTAGGGCCATACGTAACTCCCAATCACATTTTACCGTTGTTACGGGAGATTTCAATGCAAAACTGGGCCGTAGATGCGATGATGAGTTGAAAGTGGGGCCATTTGGATTTGGGCAGCGGAATCCCAGAGGACAGATGCTGGTGGACTTTATGGAGAAGGAAGGACTCTTTATGATGAATTCTTTCTTCAAGAAGCCGCCACAATGGAAATGGACCTGGTTGAGTCCCGACGGTGTAACAAGAAATGAAATTGACTTCATCATGAGCGCCAACAGACAGATATTCAACGATGTCTCTGTGATCAACAGTGTTAAAACAGGAAGTGATCACCGAATCGTCAGAGGCATGTTGAATATCAATCGTATCAAAGAGACGATTGAGCGAAACCAAGGCTCTAAAGTGTTCGCAAGAGACGCTTCTATTGGGCAAAGCCAGCTGACGAAGCTGAAGACCGAAGATGGACGTGTAACGTCGAATAAGCCAGAGGTCCTGCGGGAGATAGAGAGGTTCTACGGGCAGCTGTTTACCTCGGTCTCAGCTGAACCGCAAGGTTTAGCTGCAGACCCTAGAGCCAAGCTAACCCGACACTACACCGAAGACATCCCGGAAATCAGCCTGTACGAGATTAGGATGGCTCTCAAACAGCTTAAGAATAACAAGGCACCGGGCGAGGATGGAATTACTACGGAGCTTCTGAAAGCGGGCGGGACGCCGGTACTGAAAGTCCTTCAGAAGCTCTTCAATTCCTCTTTGCTCGATGGAAAACCGCCACAGGCATGGAACAGGGGTGTTGTGATCCTGTTCTTCAAAAAAGGTGACAAAACCTTATTGAAGAACTATAGGCCCATAGCACTGCTGAGCCATGTCTACAAGTTGTTCTCGAGAGTCATCACGAAGCGTCTCGAACGCAGGTTTGATGACTTCCAGCCTACCGAACAAGCCGGTTTCCGAAAAGGCTACAGTACCATAGACCACATACATACGCTGCGGCAGATTGTACAGAAGACCGAAGAGTACAATCGGCCATTATGCTTAGCGTTTGTGGACTATGAGAAAGCCTTTGATTCCATCGAAACATGGGCGGTCCTTAGATCTCTCCAGCGGTGCCGTATTGACCACCGGTACATCGAAGTATTGAAGTGTTTGTATAATAACGCCACCATGTCAATCCGAGTACAGGAGCTTTGCACGAAGGAAATCCCTGTAAAGAGAGGAGTGAGACAGGGAGATGTGATATCTCCGAAACTGTTTATCACTGCTCTGGAGGATTCCTTCAAGCTTCTGGAATGGCAAGGACTTGGCATCAATATTAACGGCGAATACATCACTCATCTTCGGTTTGCCGATGACATCGTGATCATGGCAGAGTCGCTGGAAGATTTAGGGCGTATGCTCGGTGACCTCAGTAGGGTTTCTCAACAAGTGGGTCTTACAATGAACATGGACAAAACAAAAATCATGTACAATGTCCATGTTGCACCCACCATAGTTACTGTTGGGAGCTCTACTCTTGAAGTTGTTGACGAATATATATACCTAGGACAGGCAGTCCGATTAGGTAGGTCCAACTTTGAGAGAGAGGTCAGTAGACGGATCCAACTCGGATGGGCAGCGTTCGGTAAACTGCACAACGTCTTCTCGTCCAAAATACCTCAGTGCCTTAAGACAAAGGCGTACAACCAATGTGTGTTACCAGTGATGACGTACGGCTCGGAGACGTGGTGCTTCACTAGGGATCTTTTTAGAAGGCTCAAAGTCGCGCAGCGAGCTATGGAGAGGGCTATGCTCGGCATTTCTCTACGTGATCGAATCCGAAATGAGGAGATCCGTAGAAGAACGAAGGTTGCTGACATAGCTCGTAGAATTAGTGAACTAAAGTGGCAATGGGCCGGCCACATAGCACGAAGAGAGGACGGTCGATGGGGCAGAAAGATCCTGGAGTGGCGACCACGTACTGGCAAGCGCAGTGTGGGACGGCCACCTACTAGATGGACCGACGACCTAGTAAAAAGCGCTGGGTATCGTTGGATGCAGGTGGCAATGGACCGGTCGCACTGGAAATCTATTGGAGAGGACTATGTTCAGCAGTGGACGGCGATAGGCTGA
Protein
MIAHCRTRPLVPGGPAISGYGSGHGNGGARGAKNPRQRSGYHPPSQRQLALVTYNVRTLRLDEKLIELEEEMNKLRWDVIGLSEIRREGEDTKTLQSGHLFYYREGEQKSQGGVGFIVHKSLINNIVAIESVSSRVVYLVLKISKRYSLKIVQVYAPSTSHPDDEVEATYEDISRAIRNSQSHFTVVTGDFNAKLGRRCDDELKVGPFGFGQRNPRGQMLVDFMEKEGLFMMNSFFKKPPQWKWTWLSPDGVTRNEIDFIMSANRQIFNDVSVINSVKTGSDHRIVRGMLNINRIKETIERNQGSKVFARDASIGQSQLTKLKTEDGRVTSNKPEVLREIERFYGQLFTSVSAEPQGLAADPRAKLTRHYTEDIPEISLYEIRMALKQLKNNKAPGEDGITTELLKAGGTPVLKVLQKLFNSSLLDGKPPQAWNRGVVILFFKKGDKTLLKNYRPIALLSHVYKLFSRVITKRLERRFDDFQPTEQAGFRKGYSTIDHIHTLRQIVQKTEEYNRPLCLAFVDYEKAFDSIETWAVLRSLQRCRIDHRYIEVLKCLYNNATMSIRVQELCTKEIPVKRGVRQGDVISPKLFITALEDSFKLLEWQGLGININGEYITHLRFADDIVIMAESLEDLGRMLGDLSRVSQQVGLTMNMDKTKIMYNVHVAPTIVTVGSSTLEVVDEYIYLGQAVRLGRSNFEREVSRRIQLGWAAFGKLHNVFSSKIPQCLKTKAYNQCVLPVMTYGSETWCFTRDLFRRLKVAQRAMERAMLGISLRDRIRNEEIRRRTKVADIARRISELKWQWAGHIARREDGRWGRKILEWRPRTGKRSVGRPPTRWTDDLVKSAGYRWMQVAMDRSHWKSIGEDYVQQWTAIG
Summary
Uniprot
Pubmed
Proteomes
Pfam
PF00078 RVT_1
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
PDB
6AR3
E-value=9.83654e-06,
Score=121
Ontologies
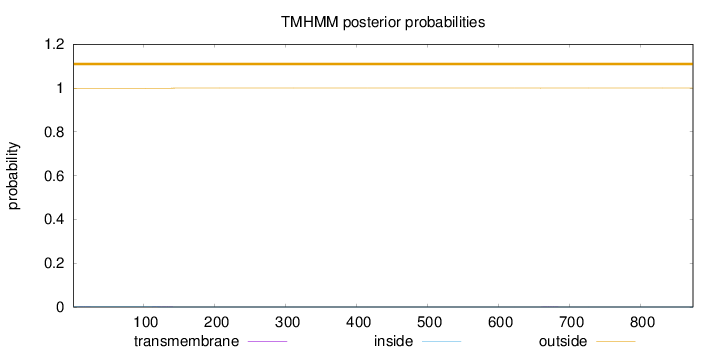
Topology
Length:
874
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.03043
Exp number, first 60 AAs:
0.00286
Total prob of N-in:
0.00124
outside
1 - 874
Population Genetic Test Statistics
Pi
541.246643
Theta
214.878984
Tajima's D
4.503548
CLR
0.526047
CSRT
0.99990000499975
Interpretation
Uncertain
