Gene
KWMTBOMO03032
Annotation
AF461149_1_transposase_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 1.671
Sequence
CDS
ATGAATGCAGTTGTGTATCAAAATACAGTCCTGACGAACCTCGTGGAACCTGTTTCTCATACCATGTTCAATAACAGGCACTGGGTATTCCAACAAGATTCGGCGCCAGCTCATAGAGCAAAAAGCACACAAGACTGGCTGGCGGCGCGTAAAATCGACTTCATCCGGCACGAAGACTGGCCCTCCTCCAGTCCAGATTTGAATCCGTTAGATTATAAGATATGGCAACACTTGGAGGAAAAGGCGTGCTCAAAGCCTCATCCCAATTTGGAGTCACTCAAGACATCCTTGATTAAGGCAGCCGCTGATATTGACATGGACCTCGTTCGTGCTGCGATAGACGACTGGCCGCGCAGATTGAAGGCCTGTATTCAAAATCACGGAGGTCATTTTGAATAA
Protein
MNAVVYQNTVLTNLVEPVSHTMFNNRHWVFQQDSAPAHRAKSTQDWLAARKIDFIRHEDWPSSSPDLNPLDYKIWQHLEEKACSKPHPNLESLKTSLIKAAADIDMDLVRAAIDDWPRRLKACIQNHGGHFE
Summary
Uniprot
Q8ITJ9
B1Q3L9
B9A8E5
B1Q3L5
B1Q3L8
P90693
+ More
G3G3K1 A0A226D9B6 A0A1I7SJQ8 A0A1I7T5D4 A0A1I7RIM5 A0A2H2JHK7 A0A2G5T9N6 E3M065 A0A368H7I1 A0A0N4YHH1 E3NJL6 E3MBG9 A0A368FVD9 A0A0N4XI93 A0A0N4XXA5 A0A016TV39 A0A1I7RU52 W6NE86 A0A016TQR4 A0A183GAB9 A0A3P8EW48 E3NDP6 A0A0N4XQJ3 A0A016S9F8 A0A2G5TSG2 A0A016S0M1 A0A2G5SZY0 E3N9D7 A0A183FPD3 A0A3P7ZA30 A0A2G5VIT1 A0A016W3L5 A0A2G5SF52 A0A0C2CZV2 A0A0N4Y729 E3MXK8 A0A183F5L1 A0A3P7TF39 H3EUR4 A0A183C618 E3ND45 E3N1H1 E3N6E8 E3M897 E3MXK7 E3M1J2 E3NHD7 E3MQK2 A0A016T5R9 E3NHK7 E3N4H8 A0A016WY15 A0A183CKH9 A0A1I7SRX5 A0A016VJ32 A0A2H2I0K4 W2STE5 A0A0N4YD61 A0A0C2DMD3 A0A0N4YLD6 A0A016WN34 A0A2G5TZM3 A0A1I8CC25 H3EXA9 H3E4E9 A0A2H2IVX6 E3N749 H3DZR2 H3FB13 H3E1D2 A0A0C2FMR5 A0A0N4XTP7 B6IKQ6 H3FES1 A0A2H2J0T6 H3ESY7 A0A183FIQ4 A0A3P7YAH0 A0A016WKB6 A0A0N4XUY5 W6NUW0 A0A1I7TL85 A0A0N4Y722 A0A183F3N6 A0A3P7TDY7 A0A0N4Y0X0 H3DS69 A0A0C2GWR6 A0A0C2H248 A0A0C2GH80 A0A2G5TN79 A0A0N4YNS9 A0A0N4Y9Q8
G3G3K1 A0A226D9B6 A0A1I7SJQ8 A0A1I7T5D4 A0A1I7RIM5 A0A2H2JHK7 A0A2G5T9N6 E3M065 A0A368H7I1 A0A0N4YHH1 E3NJL6 E3MBG9 A0A368FVD9 A0A0N4XI93 A0A0N4XXA5 A0A016TV39 A0A1I7RU52 W6NE86 A0A016TQR4 A0A183GAB9 A0A3P8EW48 E3NDP6 A0A0N4XQJ3 A0A016S9F8 A0A2G5TSG2 A0A016S0M1 A0A2G5SZY0 E3N9D7 A0A183FPD3 A0A3P7ZA30 A0A2G5VIT1 A0A016W3L5 A0A2G5SF52 A0A0C2CZV2 A0A0N4Y729 E3MXK8 A0A183F5L1 A0A3P7TF39 H3EUR4 A0A183C618 E3ND45 E3N1H1 E3N6E8 E3M897 E3MXK7 E3M1J2 E3NHD7 E3MQK2 A0A016T5R9 E3NHK7 E3N4H8 A0A016WY15 A0A183CKH9 A0A1I7SRX5 A0A016VJ32 A0A2H2I0K4 W2STE5 A0A0N4YD61 A0A0C2DMD3 A0A0N4YLD6 A0A016WN34 A0A2G5TZM3 A0A1I8CC25 H3EXA9 H3E4E9 A0A2H2IVX6 E3N749 H3DZR2 H3FB13 H3E1D2 A0A0C2FMR5 A0A0N4XTP7 B6IKQ6 H3FES1 A0A2H2J0T6 H3ESY7 A0A183FIQ4 A0A3P7YAH0 A0A016WKB6 A0A0N4XUY5 W6NUW0 A0A1I7TL85 A0A0N4Y722 A0A183F3N6 A0A3P7TDY7 A0A0N4Y0X0 H3DS69 A0A0C2GWR6 A0A0C2H248 A0A0C2GH80 A0A2G5TN79 A0A0N4YNS9 A0A0N4Y9Q8
EMBL
AF461149
AAN06610.1
AB363028
BAG15927.1
AB473770
BAH20555.1
+ More
AB363006 AB363010 AB363014 BAG15923.1 BAG15924.1 AB363018 BAG15926.1 U47917 AAB47739.1 JF779677 AEO90418.1 LNIX01000030 OXA41211.1 PDUG01000005 PIC23776.1 DS268420 EFO87874.1 JOJR01000015 RCN51257.1 UYSL01022144 VDL79905.1 DS268747 EFP00909.1 DS268433 EFO97645.1 JOJR01000583 RCN36166.1 UYSL01002425 VDL65835.1 UYSL01019905 VDL71189.1 JARK01001411 EYC06502.1 CAVP010058755 CDL95145.1 JARK01001418 EYC05389.1 UZAH01031030 VDP13493.1 DS268612 EFO94085.1 UYSL01009772 VDL68385.1 JARK01001601 EYB87313.1 PIC29926.1 JARK01001658 EYB84158.1 PDUG01000006 PIC20538.1 DS268565 EFO90304.1 UZAH01026445 VDO80847.1 PDUG01000001 PIC51673.1 JARK01001337 EYC34439.1 PDUG01000012 PIC13506.1 KN729202 KIH62598.1 UYSL01020634 VDL75527.1 DS268492 EFP11674.1 UZAH01001595 VDO19836.1 DS268606 EFO93488.1 DS268508 EFO83264.1 DS268539 EFO88027.1 DS268428 EFO94379.1 EFP11689.1 DS268421 EFO88618.1 DS268673 EFO98001.1 DS268466 EFP06959.1 JARK01001471 EYB97976.1 DS268679 EFO98285.1 DS268525 EFO85497.1 JARK01000059 EYC44525.1 JARK01001345 EYC27589.1 KI662043 ETN73029.1 UYSL01021399 VDL78127.1 KN728435 KIH63782.1 UYSL01023059 VDL81638.1 JARK01000176 EYC41234.1 PDUG01000004 PIC32571.1 DS268545 EFO88382.1 KN769991 KIH46151.1 UYSL01019770 VDL69590.1 HE601413 CAS00486.1 UZAH01025744 VDO69727.1 JARK01000255 EYC39473.1 UYSL01019810 VDL70150.1 CAVP010059404 CDL95787.1 UYSL01020631 VDL75518.1 UZAH01000669 VDO19161.1 UYSL01020109 VDL72812.1 KN728607 KIH63514.1 KN726559 KIH67820.1 KN733367 KIH58229.1 PIC28734.1 UYSL01023743 VDL82626.1 UYSL01020934 VDL76658.1
AB363006 AB363010 AB363014 BAG15923.1 BAG15924.1 AB363018 BAG15926.1 U47917 AAB47739.1 JF779677 AEO90418.1 LNIX01000030 OXA41211.1 PDUG01000005 PIC23776.1 DS268420 EFO87874.1 JOJR01000015 RCN51257.1 UYSL01022144 VDL79905.1 DS268747 EFP00909.1 DS268433 EFO97645.1 JOJR01000583 RCN36166.1 UYSL01002425 VDL65835.1 UYSL01019905 VDL71189.1 JARK01001411 EYC06502.1 CAVP010058755 CDL95145.1 JARK01001418 EYC05389.1 UZAH01031030 VDP13493.1 DS268612 EFO94085.1 UYSL01009772 VDL68385.1 JARK01001601 EYB87313.1 PIC29926.1 JARK01001658 EYB84158.1 PDUG01000006 PIC20538.1 DS268565 EFO90304.1 UZAH01026445 VDO80847.1 PDUG01000001 PIC51673.1 JARK01001337 EYC34439.1 PDUG01000012 PIC13506.1 KN729202 KIH62598.1 UYSL01020634 VDL75527.1 DS268492 EFP11674.1 UZAH01001595 VDO19836.1 DS268606 EFO93488.1 DS268508 EFO83264.1 DS268539 EFO88027.1 DS268428 EFO94379.1 EFP11689.1 DS268421 EFO88618.1 DS268673 EFO98001.1 DS268466 EFP06959.1 JARK01001471 EYB97976.1 DS268679 EFO98285.1 DS268525 EFO85497.1 JARK01000059 EYC44525.1 JARK01001345 EYC27589.1 KI662043 ETN73029.1 UYSL01021399 VDL78127.1 KN728435 KIH63782.1 UYSL01023059 VDL81638.1 JARK01000176 EYC41234.1 PDUG01000004 PIC32571.1 DS268545 EFO88382.1 KN769991 KIH46151.1 UYSL01019770 VDL69590.1 HE601413 CAS00486.1 UZAH01025744 VDO69727.1 JARK01000255 EYC39473.1 UYSL01019810 VDL70150.1 CAVP010059404 CDL95787.1 UYSL01020631 VDL75518.1 UZAH01000669 VDO19161.1 UYSL01020109 VDL72812.1 KN728607 KIH63514.1 KN726559 KIH67820.1 KN733367 KIH58229.1 PIC28734.1 UYSL01023743 VDL82626.1 UYSL01020934 VDL76658.1
Proteomes
Interpro
SUPFAM
SSF46689
SSF46689
Gene 3D
ProteinModelPortal
Q8ITJ9
B1Q3L9
B9A8E5
B1Q3L5
B1Q3L8
P90693
+ More
G3G3K1 A0A226D9B6 A0A1I7SJQ8 A0A1I7T5D4 A0A1I7RIM5 A0A2H2JHK7 A0A2G5T9N6 E3M065 A0A368H7I1 A0A0N4YHH1 E3NJL6 E3MBG9 A0A368FVD9 A0A0N4XI93 A0A0N4XXA5 A0A016TV39 A0A1I7RU52 W6NE86 A0A016TQR4 A0A183GAB9 A0A3P8EW48 E3NDP6 A0A0N4XQJ3 A0A016S9F8 A0A2G5TSG2 A0A016S0M1 A0A2G5SZY0 E3N9D7 A0A183FPD3 A0A3P7ZA30 A0A2G5VIT1 A0A016W3L5 A0A2G5SF52 A0A0C2CZV2 A0A0N4Y729 E3MXK8 A0A183F5L1 A0A3P7TF39 H3EUR4 A0A183C618 E3ND45 E3N1H1 E3N6E8 E3M897 E3MXK7 E3M1J2 E3NHD7 E3MQK2 A0A016T5R9 E3NHK7 E3N4H8 A0A016WY15 A0A183CKH9 A0A1I7SRX5 A0A016VJ32 A0A2H2I0K4 W2STE5 A0A0N4YD61 A0A0C2DMD3 A0A0N4YLD6 A0A016WN34 A0A2G5TZM3 A0A1I8CC25 H3EXA9 H3E4E9 A0A2H2IVX6 E3N749 H3DZR2 H3FB13 H3E1D2 A0A0C2FMR5 A0A0N4XTP7 B6IKQ6 H3FES1 A0A2H2J0T6 H3ESY7 A0A183FIQ4 A0A3P7YAH0 A0A016WKB6 A0A0N4XUY5 W6NUW0 A0A1I7TL85 A0A0N4Y722 A0A183F3N6 A0A3P7TDY7 A0A0N4Y0X0 H3DS69 A0A0C2GWR6 A0A0C2H248 A0A0C2GH80 A0A2G5TN79 A0A0N4YNS9 A0A0N4Y9Q8
G3G3K1 A0A226D9B6 A0A1I7SJQ8 A0A1I7T5D4 A0A1I7RIM5 A0A2H2JHK7 A0A2G5T9N6 E3M065 A0A368H7I1 A0A0N4YHH1 E3NJL6 E3MBG9 A0A368FVD9 A0A0N4XI93 A0A0N4XXA5 A0A016TV39 A0A1I7RU52 W6NE86 A0A016TQR4 A0A183GAB9 A0A3P8EW48 E3NDP6 A0A0N4XQJ3 A0A016S9F8 A0A2G5TSG2 A0A016S0M1 A0A2G5SZY0 E3N9D7 A0A183FPD3 A0A3P7ZA30 A0A2G5VIT1 A0A016W3L5 A0A2G5SF52 A0A0C2CZV2 A0A0N4Y729 E3MXK8 A0A183F5L1 A0A3P7TF39 H3EUR4 A0A183C618 E3ND45 E3N1H1 E3N6E8 E3M897 E3MXK7 E3M1J2 E3NHD7 E3MQK2 A0A016T5R9 E3NHK7 E3N4H8 A0A016WY15 A0A183CKH9 A0A1I7SRX5 A0A016VJ32 A0A2H2I0K4 W2STE5 A0A0N4YD61 A0A0C2DMD3 A0A0N4YLD6 A0A016WN34 A0A2G5TZM3 A0A1I8CC25 H3EXA9 H3E4E9 A0A2H2IVX6 E3N749 H3DZR2 H3FB13 H3E1D2 A0A0C2FMR5 A0A0N4XTP7 B6IKQ6 H3FES1 A0A2H2J0T6 H3ESY7 A0A183FIQ4 A0A3P7YAH0 A0A016WKB6 A0A0N4XUY5 W6NUW0 A0A1I7TL85 A0A0N4Y722 A0A183F3N6 A0A3P7TDY7 A0A0N4Y0X0 H3DS69 A0A0C2GWR6 A0A0C2H248 A0A0C2GH80 A0A2G5TN79 A0A0N4YNS9 A0A0N4Y9Q8
PDB
5CR4
E-value=1.1831e-09,
Score=144
Ontologies
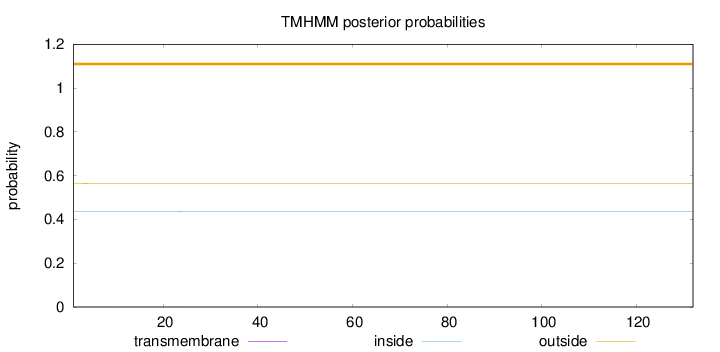
Topology
Length:
132
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00505
Exp number, first 60 AAs:
0.00505
Total prob of N-in:
0.43671
outside
1 - 132
Population Genetic Test Statistics
Pi
65.709776
Theta
72.736868
Tajima's D
-0.302022
CLR
0.000065
CSRT
0.290685465726714
Interpretation
Uncertain
