Gene
KWMTBOMO02969
Annotation
reverse_transcriptase_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 3.32
Sequence
CDS
ATGTCAGAGCTCGACTCAGACCACCGACCTGTCGTTATGCAGCTCGGTCGCCCTCACAACCCAGTCACTGTTACGAGGACCATGGTGGATTGGAATAAGCTGGGCACGTGCCTAGCCGACGCCGCTCCGCCAATCCTCCCTTACGGCCCGGATTCGAATCCATCCCCCGAGGACACCGTCGAATCCATAAACATCATTACCGATCACATCTCTTCCGCGATCATTAGATCTTCAAAAGAAGTCGATGTGGAGGATAGCTTCCACCGCATCAGACTGTCCCCCGATCTTAGGAATCTCTTAAGAGTTAGGAACGCGGCAATCCGGGCCTACGATCGTCTTCCCACGCATTCAAACCGGATTCAGATGCGTCGTCTACAACGCGAAGTCCACTCCCGCTTAAGCGACGCGCGTAACGATAATTGGCATAGTTATTTAGAACAACTCGCGCCCTCCCACCAAGCATACTGGCGACTAGCTAGGACTCTCAAATCCGAAACTACCGCTACTATGCCTCCCCTCATACGCCCTTCAGGCCAACCACCGGCATTCGATGACGATGACAAGGCTGAGCTGCTGCCCGATGCACTGCAAGAGCAGTGCACCACCAGCACTCAACACGCGGACCCCGAACACACCGAGTTAGTCGACAGGGAGGTCGAGCGCAGAGCTTCCCTGCCGCCCTCGGACGCGTTACCCCCCATTACCACTGACGAAGTTAGAGACGCGATCCACAACCTCCAACCTAGGAAGGCACCCGGCTCCAACGGCATCCACAACCGCGCGCTAAAACTCTTGCCAGTCCAACTGATAGCAATGTTGGCTACAATTTTAAATGCCGCTATGACGCACTGCATCTTTCCCGCGGTGTGGAAAGAAGCGAACGTTATCGGTATACATAAGCCGGGCAAACCGACAAACGAAACTTCTAGTTACCGTCCGATTAGTCTCCTCCCGACGATAGGAAAAATTTACGAACGGCTCCTTAGGAAACGCCTCTGGGATTTTGTTACCGCGAACAAAATTCTCATAGACGAACAGTTCGGATTCCGCTCTAAACACTCGTGCGTACAACAAGTGCACCGCCTCACGGAGCACATTCTGATAGGACTAAATAGGCGTAAACAAATCCCGACCGGCGCCCTCTTCTTCGATATCGCGAAGGCGTTCGACAAAGTCTGGCACAACGGTTTAATTTACAAACTGTACAACATGGGAGTGCCAGACAGACTCGTGCTCATCATACGAGACTTCTTGTCGAACCGTTCGTTTCGATATCGAGTAGAGGGAACTCGTTCTCGTCCCCGTCAACTGCCGGAGTCCCGCAAGGCTCCGCGCTCTCCCCGTTATTATTTAGTTTGTATATCAATGATATACCCCGGTCTCCGGAGACCCATCTAG
Protein
MSELDSDHRPVVMQLGRPHNPVTVTRTMVDWNKLGTCLADAAPPILPYGPDSNPSPEDTVESINIITDHISSAIIRSSKEVDVEDSFHRIRLSPDLRNLLRVRNAAIRAYDRLPTHSNRIQMRRLQREVHSRLSDARNDNWHSYLEQLAPSHQAYWRLARTLKSETTATMPPLIRPSGQPPAFDDDDKAELLPDALQEQCTTSTQHADPEHTELVDREVERRASLPPSDALPPITTDEVRDAIHNLQPRKAPGSNGIHNRALKLLPVQLIAMLATILNAAMTHCIFPAVWKEANVIGIHKPGKPTNETSSYRPISLLPTIGKIYERLLRKRLWDFVTANKILIDEQFGFRSKHSCVQQVHRLTEHILIGLNRRKQIPTGALFFDIAKAFDKVWHNGLIYKLYNMGVPDRLVLIIRDFLSNRSFRYRVEGTRSRPRQLPESRKAPRSPRYYLVCISMIYPGLRRPI
Summary
Uniprot
Q93137
Q9XXW0
Q9BPQ0
A0A0N1PFV1
O96546
A0A139W8G9
+ More
A0A139WA11 A0A1Y1NA23 V5GNJ2 A0A023EYS8 A0A2S2QEE0 A0A2S2PEZ9 A0A2S2Q352 Q07995 V9H092 A0A3N5Z7I3 K7IN67 A0A087UCW3 A0A2S2NPT6 A0A2H8U0H5 A0A2S2PPK8 A0A2S2PA10 A0A2S2PX53 A0A2J7R7D0 A0A087THU2 J9KLB1 A0A1B0CHA5 A0A087U3K8 J9LC21 A0A232ENU1 A0A2S2NCK2 A0A1Y1K0C8 J9KKL3 A0A023EW17 A0A087T3W0 X1WXE6 D7GYA5 A0A2S2R9D4 A0A2S2PWA8 A0A2S2P698 A0A2S2NGV8 J9M522 A0A224X6Q4 X1WZH0 X1WR00 X1WKS2 J9K7I4 A0A224X6U5 A0A2S2N629 A0A2M4BEF4 A0A2S2NJB5 A0A2M4BEZ9 A0A2S2N9P4 A0A2M4BDA5 A0A2M4BD66 A0A2M4BD62 A0A224XF24 A0A224XI99 A0A2S2P3X8 A0A224XJI0 A0A023F6S4 A0A1W7R655 A0A1Y1LKN6 A0A1Y1KFT3 A0A2S2Q2R8
A0A139WA11 A0A1Y1NA23 V5GNJ2 A0A023EYS8 A0A2S2QEE0 A0A2S2PEZ9 A0A2S2Q352 Q07995 V9H092 A0A3N5Z7I3 K7IN67 A0A087UCW3 A0A2S2NPT6 A0A2H8U0H5 A0A2S2PPK8 A0A2S2PA10 A0A2S2PX53 A0A2J7R7D0 A0A087THU2 J9KLB1 A0A1B0CHA5 A0A087U3K8 J9LC21 A0A232ENU1 A0A2S2NCK2 A0A1Y1K0C8 J9KKL3 A0A023EW17 A0A087T3W0 X1WXE6 D7GYA5 A0A2S2R9D4 A0A2S2PWA8 A0A2S2P698 A0A2S2NGV8 J9M522 A0A224X6Q4 X1WZH0 X1WR00 X1WKS2 J9K7I4 A0A224X6U5 A0A2S2N629 A0A2M4BEF4 A0A2S2NJB5 A0A2M4BEZ9 A0A2S2N9P4 A0A2M4BDA5 A0A2M4BD66 A0A2M4BD62 A0A224XF24 A0A224XI99 A0A2S2P3X8 A0A224XJI0 A0A023F6S4 A0A1W7R655 A0A1Y1LKN6 A0A1Y1KFT3 A0A2S2Q2R8
Pubmed
EMBL
U07847
AAA17752.1
AB018558
BAA76304.1
AB055391
BAB21761.1
+ More
KQ459265 KPJ02064.1 AF081103 AAC72921.1 KQ973342 KXZ75569.1 KQ971729 KYB24751.1 GEZM01008457 JAV94772.1 GALX01005299 JAB63167.1 GBBI01004395 JAC14317.1 GGMS01006359 MBY75562.1 GGMR01015412 MBY28031.1 GGMS01002955 MBY72158.1 S59870 AAB26437.2 L79944 AAB04627.1 RPOV01000134 RPJ78669.1 KK119261 KFM75202.1 GGMR01006581 MBY19200.1 GFXV01007093 MBW18898.1 GGMR01018716 MBY31335.1 GGMR01013409 MBY26028.1 GGMS01000737 MBY69940.1 NEVH01006736 PNF36716.1 KK115285 KFM64681.1 ABLF02040174 ABLF02040183 ABLF02055266 AJWK01012157 KK118005 KFM71947.1 ABLF02036316 ABLF02036319 ABLF02048663 ABLF02066977 NNAY01003095 OXU19992.1 GGMR01002322 MBY14941.1 GEZM01099921 GEZM01099920 GEZM01099918 GEZM01099906 JAV53125.1 ABLF02029306 ABLF02029314 ABLF02060138 GAPW01000276 JAC13322.1 KK113281 KFM59799.1 ABLF02016634 KQ973363 EFA13335.2 GGMS01017365 MBY86568.1 GGMS01000625 MBY69828.1 GGMR01012305 MBY24924.1 GGMR01003407 MBY16026.1 ABLF02024246 GFTR01008279 JAW08147.1 ABLF02034467 ABLF02003242 ABLF02006260 ABLF02042516 ABLF02005245 ABLF02013477 ABLF02047385 ABLF02066435 ABLF02017543 ABLF02017549 ABLF02017551 ABLF02021996 GFTR01008251 JAW08175.1 GGMR01000005 MBY12624.1 GGFJ01002256 MBW51397.1 GGMR01004661 MBY17280.1 GGFJ01002257 MBW51398.1 GGMR01000857 MBY13476.1 GGFJ01001842 MBW50983.1 GGFJ01001844 MBW50985.1 GGFJ01001843 MBW50984.1 GFTR01008038 JAW08388.1 GFTR01008256 JAW08170.1 GGMR01011511 MBY24130.1 GFTR01008257 JAW08169.1 GBBI01001646 JAC17066.1 GEHC01001034 JAV46611.1 GEZM01053022 JAV74219.1 GEZM01084970 JAV60359.1 GGMS01002786 MBY71989.1
KQ459265 KPJ02064.1 AF081103 AAC72921.1 KQ973342 KXZ75569.1 KQ971729 KYB24751.1 GEZM01008457 JAV94772.1 GALX01005299 JAB63167.1 GBBI01004395 JAC14317.1 GGMS01006359 MBY75562.1 GGMR01015412 MBY28031.1 GGMS01002955 MBY72158.1 S59870 AAB26437.2 L79944 AAB04627.1 RPOV01000134 RPJ78669.1 KK119261 KFM75202.1 GGMR01006581 MBY19200.1 GFXV01007093 MBW18898.1 GGMR01018716 MBY31335.1 GGMR01013409 MBY26028.1 GGMS01000737 MBY69940.1 NEVH01006736 PNF36716.1 KK115285 KFM64681.1 ABLF02040174 ABLF02040183 ABLF02055266 AJWK01012157 KK118005 KFM71947.1 ABLF02036316 ABLF02036319 ABLF02048663 ABLF02066977 NNAY01003095 OXU19992.1 GGMR01002322 MBY14941.1 GEZM01099921 GEZM01099920 GEZM01099918 GEZM01099906 JAV53125.1 ABLF02029306 ABLF02029314 ABLF02060138 GAPW01000276 JAC13322.1 KK113281 KFM59799.1 ABLF02016634 KQ973363 EFA13335.2 GGMS01017365 MBY86568.1 GGMS01000625 MBY69828.1 GGMR01012305 MBY24924.1 GGMR01003407 MBY16026.1 ABLF02024246 GFTR01008279 JAW08147.1 ABLF02034467 ABLF02003242 ABLF02006260 ABLF02042516 ABLF02005245 ABLF02013477 ABLF02047385 ABLF02066435 ABLF02017543 ABLF02017549 ABLF02017551 ABLF02021996 GFTR01008251 JAW08175.1 GGMR01000005 MBY12624.1 GGFJ01002256 MBW51397.1 GGMR01004661 MBY17280.1 GGFJ01002257 MBW51398.1 GGMR01000857 MBY13476.1 GGFJ01001842 MBW50983.1 GGFJ01001844 MBW50985.1 GGFJ01001843 MBW50984.1 GFTR01008038 JAW08388.1 GFTR01008256 JAW08170.1 GGMR01011511 MBY24130.1 GFTR01008257 JAW08169.1 GBBI01001646 JAC17066.1 GEHC01001034 JAV46611.1 GEZM01053022 JAV74219.1 GEZM01084970 JAV60359.1 GGMS01002786 MBY71989.1
Proteomes
Interpro
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
Q93137
Q9XXW0
Q9BPQ0
A0A0N1PFV1
O96546
A0A139W8G9
+ More
A0A139WA11 A0A1Y1NA23 V5GNJ2 A0A023EYS8 A0A2S2QEE0 A0A2S2PEZ9 A0A2S2Q352 Q07995 V9H092 A0A3N5Z7I3 K7IN67 A0A087UCW3 A0A2S2NPT6 A0A2H8U0H5 A0A2S2PPK8 A0A2S2PA10 A0A2S2PX53 A0A2J7R7D0 A0A087THU2 J9KLB1 A0A1B0CHA5 A0A087U3K8 J9LC21 A0A232ENU1 A0A2S2NCK2 A0A1Y1K0C8 J9KKL3 A0A023EW17 A0A087T3W0 X1WXE6 D7GYA5 A0A2S2R9D4 A0A2S2PWA8 A0A2S2P698 A0A2S2NGV8 J9M522 A0A224X6Q4 X1WZH0 X1WR00 X1WKS2 J9K7I4 A0A224X6U5 A0A2S2N629 A0A2M4BEF4 A0A2S2NJB5 A0A2M4BEZ9 A0A2S2N9P4 A0A2M4BDA5 A0A2M4BD66 A0A2M4BD62 A0A224XF24 A0A224XI99 A0A2S2P3X8 A0A224XJI0 A0A023F6S4 A0A1W7R655 A0A1Y1LKN6 A0A1Y1KFT3 A0A2S2Q2R8
A0A139WA11 A0A1Y1NA23 V5GNJ2 A0A023EYS8 A0A2S2QEE0 A0A2S2PEZ9 A0A2S2Q352 Q07995 V9H092 A0A3N5Z7I3 K7IN67 A0A087UCW3 A0A2S2NPT6 A0A2H8U0H5 A0A2S2PPK8 A0A2S2PA10 A0A2S2PX53 A0A2J7R7D0 A0A087THU2 J9KLB1 A0A1B0CHA5 A0A087U3K8 J9LC21 A0A232ENU1 A0A2S2NCK2 A0A1Y1K0C8 J9KKL3 A0A023EW17 A0A087T3W0 X1WXE6 D7GYA5 A0A2S2R9D4 A0A2S2PWA8 A0A2S2P698 A0A2S2NGV8 J9M522 A0A224X6Q4 X1WZH0 X1WR00 X1WKS2 J9K7I4 A0A224X6U5 A0A2S2N629 A0A2M4BEF4 A0A2S2NJB5 A0A2M4BEZ9 A0A2S2N9P4 A0A2M4BDA5 A0A2M4BD66 A0A2M4BD62 A0A224XF24 A0A224XI99 A0A2S2P3X8 A0A224XJI0 A0A023F6S4 A0A1W7R655 A0A1Y1LKN6 A0A1Y1KFT3 A0A2S2Q2R8
Ontologies
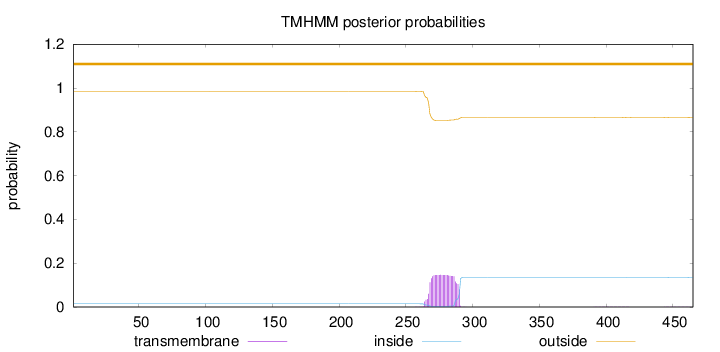
Topology
Length:
465
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
3.31173
Exp number, first 60 AAs:
0
Total prob of N-in:
0.01517
outside
1 - 465
Population Genetic Test Statistics
Pi
0
Theta
0
Tajima's D
0
CLR
5.117436
CSRT
0
Interpretation
Uncertain
