Gene
KWMTBOMO02956 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA003567
Annotation
alkaliphilic_serine_protease_[Bombyx_mori]
Full name
Trypsin, alkaline A
+ More
Trypsin, alkaline C
Trypsin, alkaline B
Trypsin CFT-1
Trypsin, alkaline C
Trypsin, alkaline B
Trypsin CFT-1
Location in the cell
Extracellular Reliability : 4.75
Sequence
CDS
ATGCGGTCTACCATTATTTTGTTGGCCCTTGGGCTTGCTGCTGTGGCGGCTGTTCCTACCAACCCACAAAGGATTATTGGTGGCTCCACGACCAACATTAACCAGTATCCCGGTATAGCTGCTTTGTTGTATACATGGAATTGGAATCAGTGGTGGCAGTCTTGTGGTGGAAACATCCTTAACCAGAGATCTATCCTTAGTGCCGCTCACTGTCCATACGGTGACGCTACAGGCCGTTGGCGTATTCGTGTTGGTTCCACCTTTGCCAACAGTGGGGGTGTTGTGCATAACGTGAACAGAATTATCATTCATCCCAACTATAACAGACGTACTGCTGACAGTGACCTCTGCATCTTACGCAGTAATTCCAACATAGCTTATAACAACAACGTTCGTCCTATTAATATTGCTGGTGCCAATTACAATCTTGGTGATAACCAAGTTGTTTGGGCTGCTGGTTGGGGTGCTACATCGCTCGGAGGGTCTAATTCGGAGCAACTCCGTCACGTCCAGGTCTGGACCATCAATCAGAATGCCTGCGTCCAACGTTACAGACCCATTAACCGTGCTATCACCGCTAACATGTTGTGCTCTGGTGTTTTGGACGTCGGTGGTCGCGACCAGTGCCAGGGTGACTCCGGTGGTCCTCTCCTCCACAACCGCGTCCTCGTTGGTGTCTGCTCTTGGGGACAGTACTGCGCTGACCGTCGCTTCCCTGGTGTTAACGTTCGCGTTTCCCGTTTCACCTCTTGGATCCAATCCAATTCGTAA
Protein
MRSTIILLALGLAAVAAVPTNPQRIIGGSTTNINQYPGIAALLYTWNWNQWWQSCGGNILNQRSILSAAHCPYGDATGRWRIRVGSTFANSGGVVHNVNRIIIHPNYNRRTADSDLCILRSNSNIAYNNNVRPINIAGANYNLGDNQVVWAAGWGATSLGGSNSEQLRHVQVWTINQNACVQRYRPINRAITANMLCSGVLDVGGRDQCQGDSGGPLLHNRVLVGVCSWGQYCADRRFPGVNVRVSRFTSWIQSNS
Summary
Catalytic Activity
Preferential cleavage: Arg-|-Xaa, Lys-|-Xaa.
Similarity
Belongs to the peptidase S1 family.
Keywords
Disulfide bond
Hydrolase
Protease
Secreted
Serine protease
Signal
Zymogen
Direct protein sequencing
Feature
propeptide Activation peptide
chain Trypsin, alkaline A
chain Trypsin, alkaline A
Uniprot
H9J230
J7JYV8
H9J229
Q9TXE6
P35045
P35047
+ More
A0A1L6UW50 A0A194PNR3 P35046 Q27540 P35042 A0A2U8NFD7 A0A194PNR6 A0A194PP33 A0A0N1I947 Q961Y0 B4X995 A0A2A4K5G7 B4X996 A0A2A4K6E9 B4X9A1 O18447 A0A2A4K648 C9W8F7 V9M0R7 A0A2A4K6E3 J7IJ83 Q4L1K4 O76954 A0A194PQQ5 A0A2H1VJ41 A0A2A4K747 A0A0N1PHJ0 S5PVP1 A0A2W1BA14 A0A2A4K5F8 A0A2A4K5I4 A0A2W1BQW9 A0A194PQ27 A0A2W1B606 H9J231 A0A2H1VPG9 H9J243 A0A194PVB6 Q4L1K6 J7IF09 A0A1B0RHN7 O18434 B4X9A0 Q4L1M0 A0A1C9M3A1 O18436 A0A2W1BL45 A0A2W1BKF9 A0A2W1BF78 A0A2W1BKV8 A0A2A4K769 B3F884 J7IFG7 I7E449 A0A2W1BP46 A0A2A4K6U2 A0A0K8SII3 A0A2W1BGQ7 A0A2W1B1G1 E7D004 A0A2A4K6W3 A0A2A4K6F3 C4PFX2 A0A2W1BP39 W5X3N9 I7DEI4 A0A194PQQ0 O62598 V9LL27 Q56IB6 A0A2H1W4N7 I7DHL6 A0A2W1BSA6 D7S002 A0A2A4K636 Q56IB8 Q4L1K8 Q56IB4 O18435 Q6R561 A0A2A4K756 D7S000 A0A2H1WNB6 A0A2W1BE44 Q56IB5 Q4L1L9 Q56IB7 Q4L1L8 A0A2W1BB90 D7S001 A0A2A4K5I0 C9W8J5 J7IN23 A0A2H1W9X8
A0A1L6UW50 A0A194PNR3 P35046 Q27540 P35042 A0A2U8NFD7 A0A194PNR6 A0A194PP33 A0A0N1I947 Q961Y0 B4X995 A0A2A4K5G7 B4X996 A0A2A4K6E9 B4X9A1 O18447 A0A2A4K648 C9W8F7 V9M0R7 A0A2A4K6E3 J7IJ83 Q4L1K4 O76954 A0A194PQQ5 A0A2H1VJ41 A0A2A4K747 A0A0N1PHJ0 S5PVP1 A0A2W1BA14 A0A2A4K5F8 A0A2A4K5I4 A0A2W1BQW9 A0A194PQ27 A0A2W1B606 H9J231 A0A2H1VPG9 H9J243 A0A194PVB6 Q4L1K6 J7IF09 A0A1B0RHN7 O18434 B4X9A0 Q4L1M0 A0A1C9M3A1 O18436 A0A2W1BL45 A0A2W1BKF9 A0A2W1BF78 A0A2W1BKV8 A0A2A4K769 B3F884 J7IFG7 I7E449 A0A2W1BP46 A0A2A4K6U2 A0A0K8SII3 A0A2W1BGQ7 A0A2W1B1G1 E7D004 A0A2A4K6W3 A0A2A4K6F3 C4PFX2 A0A2W1BP39 W5X3N9 I7DEI4 A0A194PQQ0 O62598 V9LL27 Q56IB6 A0A2H1W4N7 I7DHL6 A0A2W1BSA6 D7S002 A0A2A4K636 Q56IB8 Q4L1K8 Q56IB4 O18435 Q6R561 A0A2A4K756 D7S000 A0A2H1WNB6 A0A2W1BE44 Q56IB5 Q4L1L9 Q56IB7 Q4L1L8 A0A2W1BB90 D7S001 A0A2A4K5I0 C9W8J5 J7IN23 A0A2H1W9X8
EC Number
3.4.21.4
Pubmed
EMBL
BABH01007517
JX312360
AFQ59994.1
BABH01007520
L16805
L16807
+ More
KY379952 APS85762.1 KQ459597 KPI95066.1 L16806 U12917 AAA84423.1 L04749 MF407317 AWL83213.1 KPI95071.1 KPI95067.1 KQ460226 KPJ16474.1 AY040819 AAK81696.1 EF531631 EF531633 ABR88242.1 ABR88244.1 NWSH01000106 PCG79485.1 EF531632 ABR88243.1 PCG79488.1 EF531637 ABR88248.1 HM209426 Y12283 KZ150104 ADI32887.1 CAA72962.1 PZC73444.1 PCG79486.1 FJ205402 ACR15970.1 JX866720 AGB93875.1 PCG79474.1 JX195651 AFQ37934.1 AY587161 AAT95353.1 AJ007706 CAA07611.1 KPI95074.1 ODYU01002655 SOQ40442.1 PCG79472.1 KPJ16473.1 KC175562 AGR92345.1 KZ150747 PZC70337.1 PCG79475.1 PCG79487.1 KZ149992 PZC75527.1 KPI95068.1 PZC70335.1 BABH01007514 BABH01007515 ODYU01003655 SOQ42698.1 BABH01007473 KPI95070.1 AY587159 AAT95351.1 JX195649 AFQ37932.1 KM360188 AKH49604.1 Y12269 CAA72948.1 PZC73449.1 EF531636 ABR88247.1 PCG79483.1 AY587145 AAT95360.1 KU302250 AOQ30448.1 EF104916 Y12271 ABP96918.1 CAA72950.1 PZC73450.1 PZC75529.1 PZC73448.1 PZC75528.1 PCG79490.1 EF635223 ABU96714.1 JX195652 AFQ37935.1 JX046916 AFO83992.1 PZC73453.1 PCG79473.1 GBRD01013231 JAG52595.1 PZC73441.1 PZC70338.1 HM990174 ADT80822.1 PCG79493.1 PCG79484.1 FJ940726 ODYU01006287 ACR25157.1 SOQ47996.1 PZC73443.1 KF779933 AHI07442.1 JN793539 AFO68319.1 KPI95069.1 AF064525 AF064526 AAC36247.1 AAC36248.1 JQ340915 AFP74111.1 AY953061 AAX62034.1 ODYU01007611 SOQ47997.1 SOQ50538.1 JN793542 AFO68322.1 PZC75526.1 HM209425 ADI32886.1 PCG79476.1 AY953059 AAX62032.1 AY587157 AAT95349.1 AY953063 AAX62036.1 Y12270 CAA72949.1 AY513649 AAR98918.1 PCG79482.1 HM209423 ADI32884.1 ODYU01009856 SOQ54559.1 PZC73442.1 AY953062 AAX62035.1 AY587146 AAT95361.1 AY953060 AAX62033.1 AY587147 AAT95362.1 PZC70336.1 HM209424 ADI32885.1 PCG79481.1 FJ205445 ACR16008.2 JX195648 AFQ37931.1 ODYU01007266 SOQ49895.1
KY379952 APS85762.1 KQ459597 KPI95066.1 L16806 U12917 AAA84423.1 L04749 MF407317 AWL83213.1 KPI95071.1 KPI95067.1 KQ460226 KPJ16474.1 AY040819 AAK81696.1 EF531631 EF531633 ABR88242.1 ABR88244.1 NWSH01000106 PCG79485.1 EF531632 ABR88243.1 PCG79488.1 EF531637 ABR88248.1 HM209426 Y12283 KZ150104 ADI32887.1 CAA72962.1 PZC73444.1 PCG79486.1 FJ205402 ACR15970.1 JX866720 AGB93875.1 PCG79474.1 JX195651 AFQ37934.1 AY587161 AAT95353.1 AJ007706 CAA07611.1 KPI95074.1 ODYU01002655 SOQ40442.1 PCG79472.1 KPJ16473.1 KC175562 AGR92345.1 KZ150747 PZC70337.1 PCG79475.1 PCG79487.1 KZ149992 PZC75527.1 KPI95068.1 PZC70335.1 BABH01007514 BABH01007515 ODYU01003655 SOQ42698.1 BABH01007473 KPI95070.1 AY587159 AAT95351.1 JX195649 AFQ37932.1 KM360188 AKH49604.1 Y12269 CAA72948.1 PZC73449.1 EF531636 ABR88247.1 PCG79483.1 AY587145 AAT95360.1 KU302250 AOQ30448.1 EF104916 Y12271 ABP96918.1 CAA72950.1 PZC73450.1 PZC75529.1 PZC73448.1 PZC75528.1 PCG79490.1 EF635223 ABU96714.1 JX195652 AFQ37935.1 JX046916 AFO83992.1 PZC73453.1 PCG79473.1 GBRD01013231 JAG52595.1 PZC73441.1 PZC70338.1 HM990174 ADT80822.1 PCG79493.1 PCG79484.1 FJ940726 ODYU01006287 ACR25157.1 SOQ47996.1 PZC73443.1 KF779933 AHI07442.1 JN793539 AFO68319.1 KPI95069.1 AF064525 AF064526 AAC36247.1 AAC36248.1 JQ340915 AFP74111.1 AY953061 AAX62034.1 ODYU01007611 SOQ47997.1 SOQ50538.1 JN793542 AFO68322.1 PZC75526.1 HM209425 ADI32886.1 PCG79476.1 AY953059 AAX62032.1 AY587157 AAT95349.1 AY953063 AAX62036.1 Y12270 CAA72949.1 AY513649 AAR98918.1 PCG79482.1 HM209423 ADI32884.1 ODYU01009856 SOQ54559.1 PZC73442.1 AY953062 AAX62035.1 AY587146 AAT95361.1 AY953060 AAX62033.1 AY587147 AAT95362.1 PZC70336.1 HM209424 ADI32885.1 PCG79481.1 FJ205445 ACR16008.2 JX195648 AFQ37931.1 ODYU01007266 SOQ49895.1
Proteomes
Pfam
PF00089 Trypsin
Interpro
SUPFAM
SSF50494
SSF50494
CDD
ProteinModelPortal
H9J230
J7JYV8
H9J229
Q9TXE6
P35045
P35047
+ More
A0A1L6UW50 A0A194PNR3 P35046 Q27540 P35042 A0A2U8NFD7 A0A194PNR6 A0A194PP33 A0A0N1I947 Q961Y0 B4X995 A0A2A4K5G7 B4X996 A0A2A4K6E9 B4X9A1 O18447 A0A2A4K648 C9W8F7 V9M0R7 A0A2A4K6E3 J7IJ83 Q4L1K4 O76954 A0A194PQQ5 A0A2H1VJ41 A0A2A4K747 A0A0N1PHJ0 S5PVP1 A0A2W1BA14 A0A2A4K5F8 A0A2A4K5I4 A0A2W1BQW9 A0A194PQ27 A0A2W1B606 H9J231 A0A2H1VPG9 H9J243 A0A194PVB6 Q4L1K6 J7IF09 A0A1B0RHN7 O18434 B4X9A0 Q4L1M0 A0A1C9M3A1 O18436 A0A2W1BL45 A0A2W1BKF9 A0A2W1BF78 A0A2W1BKV8 A0A2A4K769 B3F884 J7IFG7 I7E449 A0A2W1BP46 A0A2A4K6U2 A0A0K8SII3 A0A2W1BGQ7 A0A2W1B1G1 E7D004 A0A2A4K6W3 A0A2A4K6F3 C4PFX2 A0A2W1BP39 W5X3N9 I7DEI4 A0A194PQQ0 O62598 V9LL27 Q56IB6 A0A2H1W4N7 I7DHL6 A0A2W1BSA6 D7S002 A0A2A4K636 Q56IB8 Q4L1K8 Q56IB4 O18435 Q6R561 A0A2A4K756 D7S000 A0A2H1WNB6 A0A2W1BE44 Q56IB5 Q4L1L9 Q56IB7 Q4L1L8 A0A2W1BB90 D7S001 A0A2A4K5I0 C9W8J5 J7IN23 A0A2H1W9X8
A0A1L6UW50 A0A194PNR3 P35046 Q27540 P35042 A0A2U8NFD7 A0A194PNR6 A0A194PP33 A0A0N1I947 Q961Y0 B4X995 A0A2A4K5G7 B4X996 A0A2A4K6E9 B4X9A1 O18447 A0A2A4K648 C9W8F7 V9M0R7 A0A2A4K6E3 J7IJ83 Q4L1K4 O76954 A0A194PQQ5 A0A2H1VJ41 A0A2A4K747 A0A0N1PHJ0 S5PVP1 A0A2W1BA14 A0A2A4K5F8 A0A2A4K5I4 A0A2W1BQW9 A0A194PQ27 A0A2W1B606 H9J231 A0A2H1VPG9 H9J243 A0A194PVB6 Q4L1K6 J7IF09 A0A1B0RHN7 O18434 B4X9A0 Q4L1M0 A0A1C9M3A1 O18436 A0A2W1BL45 A0A2W1BKF9 A0A2W1BF78 A0A2W1BKV8 A0A2A4K769 B3F884 J7IFG7 I7E449 A0A2W1BP46 A0A2A4K6U2 A0A0K8SII3 A0A2W1BGQ7 A0A2W1B1G1 E7D004 A0A2A4K6W3 A0A2A4K6F3 C4PFX2 A0A2W1BP39 W5X3N9 I7DEI4 A0A194PQQ0 O62598 V9LL27 Q56IB6 A0A2H1W4N7 I7DHL6 A0A2W1BSA6 D7S002 A0A2A4K636 Q56IB8 Q4L1K8 Q56IB4 O18435 Q6R561 A0A2A4K756 D7S000 A0A2H1WNB6 A0A2W1BE44 Q56IB5 Q4L1L9 Q56IB7 Q4L1L8 A0A2W1BB90 D7S001 A0A2A4K5I0 C9W8J5 J7IN23 A0A2H1W9X8
PDB
1EKB
E-value=2.92483e-29,
Score=318
Ontologies
GO
Topology
Subcellular location
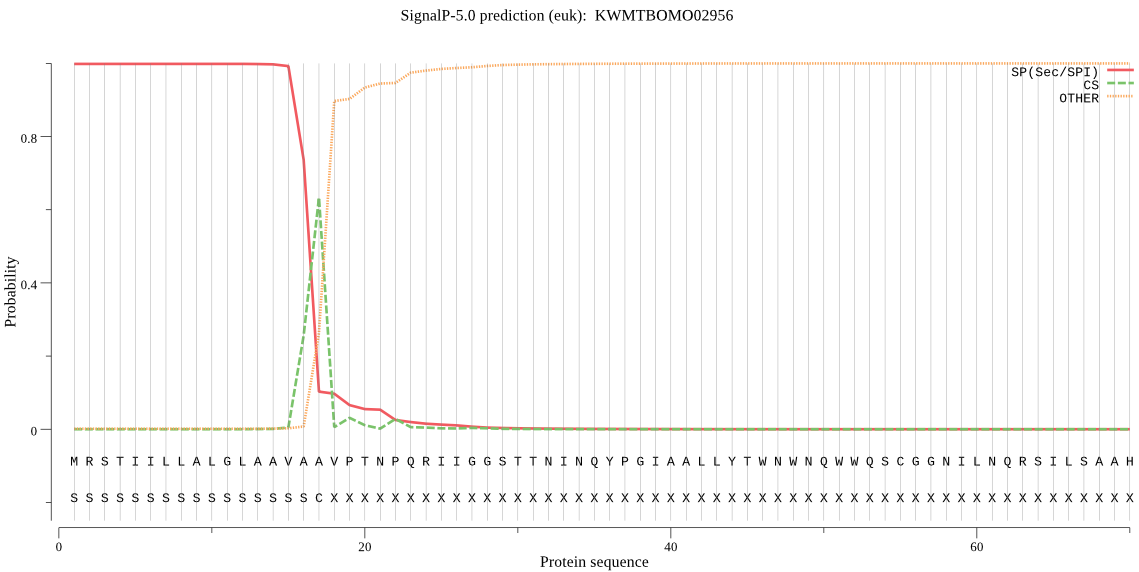
SignalP
Position: 1 - 17,
Likelihood: 0.998356
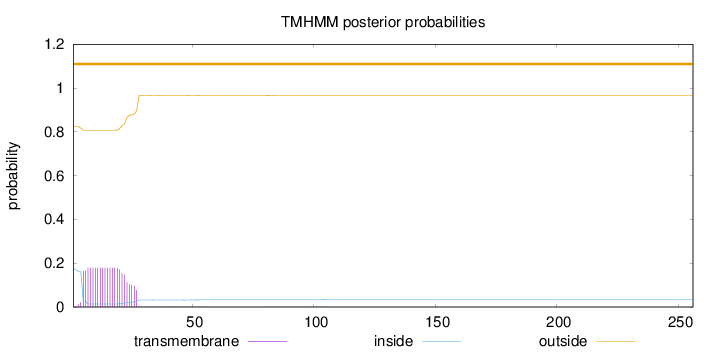
Length:
256
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
3.7294
Exp number, first 60 AAs:
3.70564
Total prob of N-in:
0.17413
outside
1 - 256
Population Genetic Test Statistics
Pi
208.423842
Theta
3.552175
Tajima's D
-1.004291
CLR
31.204866
CSRT
0.130393480325984
Interpretation
Uncertain
