Gene
KWMTBOMO02798
Annotation
hypothetical_protein_[Piscirickettsia_salmonis]
Location in the cell
Mitochondrial Reliability : 1.419 PlasmaMembrane Reliability : 1.421
Sequence
CDS
ATGGACCGCACCTATCGCTTCCTCCAAGGAAACCTAAACCACTCCGCCAGAGCTGAGGACATGCTGTCTCAGAGCATGGCGGAGTGGTCCATAGATATGGCTGTGGTCGCCGAGCCGTACTTCGTTCCCCCAGCAAATGATTGCTGGTTTGGGGACGACGGCGGCCTGGCGGCCATAGTCTTAAGAAGAGACGCGGCGATCCCCCCCCTGGCGATGGTGACCAGGGGTCAAGGGTTCGTCGCGGTGCAGCTGGAAGGGACCGTCGTGATCGGGGCGTACTTCTCTCCGAATAGGCCTACCGTCGAGTTCGGGCATTTCCTGGGCGGGATCGGGGCGATTGCTCGCCGCCTCGCTCCTCGTCCAGTGATTCTCGCGGGGGACCTCAATGCGAAGTCTGTCGCATGGGGCTCCCCCCGCTCGGACGCTCGTGGTAGGCTATTGGAGAGGTGGGCGCACGCGGCCGGCCTCTGTTTGTTGAATAGAGGTTCGGTTGCGACGTGCGTGCGGTGGCAGGGCGAGTCTATTGTGGACGTTACGTTCGCAAGCCCGGCCATCGCACGCCGTATCCGCGACTGGAGAGTTTTGGAGGGGGCGGAGACGCTGTCAGATCACCGATATATTCGGTTTGATCTCTCCGCCCGCTCCGCAGTCGCGGACGTCCACGGGGATCACCCCCGTGGTGCATCCCGGTCATTTCCCAAGTGGGCACTGAAGCGCCTGAATAAGGAGGTCCTGATGGAAGCCTCTACGGTGGCGGCGTGGGCGCCCATGCGTCCACACCCGGTCGAGGTCGAGGGAGAGGCAGGGTGGTTCCGGGACACGATGCGTTGCATTTGTGATGCGTCGATGCCCCGGATCGGCCCTAGACTCCCTAATCGCCAGGCGTACTGGTGGTCGCCCGAAATTGCGCAACTCCGCGTGGAATGCGTTCGGGCGAGGCGCGAGTGCGCCAGGCATCGCCGCCGCCGCCTGCCGCGGCGCAACGATCCGGTTGCGTTCGCGGCGGCAGAGGCCCGGCTGCACGGCGCATTTCGCGTCGCGCAGAGGGCATTGCAGCTGGCCATCAGAAGAGCCAAGAACCAACACATGGAGGGTCTCTTGGAGACGCTGGATGAGGATCCGTGGGGGCGGCCCTATCATATGGTGCGAAATAAAATGCGCCCATGGGCCCCCCCGATCACGGAGCGTCTCCAGCCTCAGCAGCTGCGGGACATCGTCTCGGCGCTGTTCCCGCAGAAGCGGGAGGGATTTGTCGCTCCCGCTATGGACGCGCCGCCGGAATACGACGGCGAAGTCCCTGCTGAGGTGCCCCGTATAACGGGGGCGGAGCTCCGTGTGGCCGTGCGCAAAATGTGCGCGAAAGACACTGCACCCGGCCCTGACGGCGTGCATGGCCGGGTTTGGGCCTTGGCTCTTGGTGCCCTAGGGGACCGACTCTTGAGGCTTTACAACTCCTGCCTCGAGTCGGGACGGTTTCCTTCATCGTGGAGGACGGGCAGACTCGTGCTGTTGAGAAAGGAGGGGCGCCCGGCGGATACTGCCGCCGGGTACCGTCCCATCGTGTTGCTGGATGAGGTGGGCAAGCTGCTGGAACGCATTCTGGCGGCCCGCATCATCCAGTATCTGGTCAGGGTGGGACCCGATCTGTCGGCGGAGCAGTATGGCTTCCGAGAGGGCCGCTCAACGGTAGACGCGATCCTTCGCGTGCGGGCCCTCTCGGAGGAGGACGTCTCCAGGGGTGGGGTGGCTTTGGCGGTGTCTCTCGACATCGCCAATGCGTTTAACACCCTGCCCTGGTCCGTGATAGGGGGGGCGCTGGAACGACACGGAGTGCCCCCCTACCTTTGCCGGCTGGTGGGTTCCTACTTGGAGGACCGATCGGTCACGTGTACCGGACACGGTGGGATCGTGCACCGGTTCCCGGTCGTGCGCGGTGTTCCACAGGGGTCGGTGCTCGGCCCCCTTTTGTGGAATATCGGGTATGACTGGGTACTGAGGGGTGCCCTCCTCCCGGGCCTAAGCGTAATTTGTTACGCAGACGACACGTTGGTCGTGGCCCGGGGGGGGAGTTTTGCTGAGTCTGCCCGTCTTGCTACCGCTGGGGTGGCGCATGTCGTCGGCAAAATCAAGAGATTGGGCCTCGACGTGGCGCTCAGTAAATCCGAGGCCATGTGGTTCCACAGGCCCCGGAGAGTGCCACCTGTCGATGCCCATATCGTGGTTGGAGGCGTCCGGATTGGGGTCGGGGTGCAGTTGAAGTACCTCGGCCTCATTCTGGACAGTCGTTGGACCTTCCGTGCTCACTTTCAGCACCTGGTCCCTCGTTTGTTGGGGGTGGCCGGTGCGTTAAGCCGGCTTCTGCCCAACGTCGGGGGGCCTGACCGGGTGACGCGCCGTCTCTATACGGGGGTGGTGCGATCAATGGCCCTATACAAGGCGCCTGTGTGGGACCAGTCCCTGGCCGCGGGGGTGGCGAAGCTGCTGCAACGGCCGCAACGCACCATCTCGGTCAGGGTCATTCGTGGTTATCGCACCATCTCCTTTGAGGCGGCGTGTGTACTGGCTAGGACGCCGCCTTGGGCTCTGGAAGCGGAGGCGCTCGCCGCTGACTATCAGTGGCGGGCTGACCTTCGCGCCCGGGGCCCAGCCACAGTGTGGTCAGAGCGCGGAGGGCCCAATCTCGGCGGTCCGTGCTGGAGTCATGGTCCAGACGGCTGGCTGATCCTTCGGCCGGTCGTAGGACCGTCGAGGCGATTCGCCCGGTCCTTGTGGATTGGGTGA
Protein
MDRTYRFLQGNLNHSARAEDMLSQSMAEWSIDMAVVAEPYFVPPANDCWFGDDGGLAAIVLRRDAAIPPLAMVTRGQGFVAVQLEGTVVIGAYFSPNRPTVEFGHFLGGIGAIARRLAPRPVILAGDLNAKSVAWGSPRSDARGRLLERWAHAAGLCLLNRGSVATCVRWQGESIVDVTFASPAIARRIRDWRVLEGAETLSDHRYIRFDLSARSAVADVHGDHPRGASRSFPKWALKRLNKEVLMEASTVAAWAPMRPHPVEVEGEAGWFRDTMRCICDASMPRIGPRLPNRQAYWWSPEIAQLRVECVRARRECARHRRRRLPRRNDPVAFAAAEARLHGAFRVAQRALQLAIRRAKNQHMEGLLETLDEDPWGRPYHMVRNKMRPWAPPITERLQPQQLRDIVSALFPQKREGFVAPAMDAPPEYDGEVPAEVPRITGAELRVAVRKMCAKDTAPGPDGVHGRVWALALGALGDRLLRLYNSCLESGRFPSSWRTGRLVLLRKEGRPADTAAGYRPIVLLDEVGKLLERILAARIIQYLVRVGPDLSAEQYGFREGRSTVDAILRVRALSEEDVSRGGVALAVSLDIANAFNTLPWSVIGGALERHGVPPYLCRLVGSYLEDRSVTCTGHGGIVHRFPVVRGVPQGSVLGPLLWNIGYDWVLRGALLPGLSVICYADDTLVVARGGSFAESARLATAGVAHVVGKIKRLGLDVALSKSEAMWFHRPRRVPPVDAHIVVGGVRIGVGVQLKYLGLILDSRWTFRAHFQHLVPRLLGVAGALSRLLPNVGGPDRVTRRLYTGVVRSMALYKAPVWDQSLAAGVAKLLQRPQRTISVRVIRGYRTISFEAACVLARTPPWALEAEALAADYQWRADLRARGPATVWSERGGPNLGGPCWSHGPDGWLILRPVVGPSRRFARSLWIG
Summary
Uniprot
Q868Q4
Q8MY35
Q8MY31
Q8MY33
Q8MY27
Q8MY25
+ More
O01419 A0A0J7N1D9 A0A2W1C3J5 A0A212EYI2 A0A0J7KJG6 A0A0J7KIY5 A0A0J7KQK3 A0A0J7KQY4 A0A0J7K3F1 A0A0J7MWN5 A0A0J7KF37 A0A0J7KIM6 A0A0J7KJH4 A0A0J7KPR0 A0A0J7KYD2 A0A0J7KRB8 A0A0J7KF91 A0A0J7MY17 A0A0J7KF40 A0A0J7NFZ0 A0A2S2PEZ7 A0A0J7N0R3 A0A0J7KFH2 A0A0J7N8R4 A0A2S2NQQ6 J9LVU7 A0A0J7KCD9 A0A0J7KCE1 A0A0J7NAV8 A0A0J7NC28 X1WJY4 A0A0J7NJI3 T1DCM0 A0A0J7KIS5 J9JWJ3 J9KT61 A0A2M4BBV9 A0A2M4BC28 A0A2M4BC01 A0A2M4CS98 J9LQQ1 A0A2M4BCE1 T1DG89 T1DG59 A0A2S2PRL8 A0A0K8TR60 A0A2M4ADW9 A0A2S2P7D1 Q868S4 J9LA43 J9KM84 X1WZ64 A0A2H8TIE5 F7IYV5 J9KNN8 F7IYV7 A0A2M4AGH2 A0A2M4CL98 J9LM37 A0A0J7KWU5 J9L0J0 A0A2M4BC18 W8ADE4 W8ANK7 A0A2M4BC24 A0A0J7NKG1 J9KJG8 X1WXT5 A0A139WAH4 A0A142LX37 Q868S0 A0A2M4AHG6 A0A139WA31 F7IYU8 A0A2H8TR36 Q868R0 J9LB02 D6WC19 A0A142LX45 A0A2M4CJ33
O01419 A0A0J7N1D9 A0A2W1C3J5 A0A212EYI2 A0A0J7KJG6 A0A0J7KIY5 A0A0J7KQK3 A0A0J7KQY4 A0A0J7K3F1 A0A0J7MWN5 A0A0J7KF37 A0A0J7KIM6 A0A0J7KJH4 A0A0J7KPR0 A0A0J7KYD2 A0A0J7KRB8 A0A0J7KF91 A0A0J7MY17 A0A0J7KF40 A0A0J7NFZ0 A0A2S2PEZ7 A0A0J7N0R3 A0A0J7KFH2 A0A0J7N8R4 A0A2S2NQQ6 J9LVU7 A0A0J7KCD9 A0A0J7KCE1 A0A0J7NAV8 A0A0J7NC28 X1WJY4 A0A0J7NJI3 T1DCM0 A0A0J7KIS5 J9JWJ3 J9KT61 A0A2M4BBV9 A0A2M4BC28 A0A2M4BC01 A0A2M4CS98 J9LQQ1 A0A2M4BCE1 T1DG89 T1DG59 A0A2S2PRL8 A0A0K8TR60 A0A2M4ADW9 A0A2S2P7D1 Q868S4 J9LA43 J9KM84 X1WZ64 A0A2H8TIE5 F7IYV5 J9KNN8 F7IYV7 A0A2M4AGH2 A0A2M4CL98 J9LM37 A0A0J7KWU5 J9L0J0 A0A2M4BC18 W8ADE4 W8ANK7 A0A2M4BC24 A0A0J7NKG1 J9KJG8 X1WXT5 A0A139WAH4 A0A142LX37 Q868S0 A0A2M4AHG6 A0A139WA31 F7IYU8 A0A2H8TR36 Q868R0 J9LB02 D6WC19 A0A142LX45 A0A2M4CJ33
Pubmed
EMBL
AB090825
BAC57926.1
AB078929
BAC06452.1
AB078931
BAC06456.1
+ More
AB078930 BAC06454.1 AB078935 BAC06462.1 AB078936 BAC06464.1 D85594 BAA19776.1 LBMM01012027 KMQ86470.1 KZ149896 PZC78733.1 AGBW02011529 OWR46521.1 LBMM01006671 KMQ90407.1 LBMM01006954 KMQ90166.1 LBMM01004272 KMQ92618.1 LBMM01004262 KMQ92649.1 LBMM01015025 KMQ84953.1 LBMM01015285 KMQ84870.1 LBMM01008448 KMQ88867.1 LBMM01007036 KMQ90099.1 LBMM01006657 KMQ90417.1 LBMM01004634 KMQ92224.1 LBMM01002052 KMQ95336.1 LBMM01003988 KMQ92922.1 LBMM01008445 KMQ88871.1 LBMM01014122 KMQ85315.1 LBMM01008422 KMQ88894.1 LBMM01005474 KMQ91485.1 GGMR01015159 MBY27778.1 LBMM01012412 KMQ86235.1 LBMM01008278 KMQ89009.1 LBMM01008226 KMQ89040.1 GGMR01006838 MBY19457.1 ABLF02034925 LBMM01009610 KMQ87992.1 LBMM01009535 KMQ88053.1 LBMM01007425 KMQ89740.1 LBMM01006996 KMQ90130.1 ABLF02011238 LBMM01004261 KMQ92650.1 GALA01001777 JAA93075.1 LBMM01006977 KMQ90149.1 ABLF02013358 ABLF02013361 ABLF02054869 ABLF02009073 ABLF02030702 ABLF02042963 GGFJ01001369 MBW50510.1 GGFJ01001439 MBW50580.1 GGFJ01001370 MBW50511.1 GGFL01004044 MBW68222.1 ABLF02030663 ABLF02041312 GGFJ01001541 MBW50682.1 GALA01001776 JAA93076.1 GALA01000302 JAA94550.1 GGMR01019493 MBY32112.1 GDAI01000741 JAI16862.1 GGFK01005660 MBW38981.1 GGMR01012748 MBY25367.1 AB090815 BAC57906.1 ABLF02018098 ABLF02018942 ABLF02041885 ABLF02011183 ABLF02041884 GFXV01002090 MBW13895.1 AB593325 BAK38646.1 ABLF02023257 AB593327 BAK38648.1 GGFK01006563 MBW39884.1 GGFL01001916 MBW66094.1 ABLF02014650 LBMM01002480 KMQ94776.1 ABLF02027248 GGFJ01001453 MBW50594.1 GAMC01020519 GAMC01020515 JAB86040.1 GAMC01020517 JAB86038.1 GGFJ01001454 MBW50595.1 LBMM01003941 KMQ92970.1 ABLF02041886 ABLF02004854 ABLF02041317 KQ971388 KYB24913.1 KU543679 AMS38363.1 AB090817 BAC57910.1 GGFK01006866 MBW40187.1 KQ971675 KYB24771.1 AB593321 BAK38639.1 GFXV01003903 MBW15708.1 AB090822 BAC57920.1 ABLF02011136 ABLF02014426 KQ971309 EEZ99146.1 KU543683 AMS38371.1 GGFL01001192 MBW65370.1
AB078930 BAC06454.1 AB078935 BAC06462.1 AB078936 BAC06464.1 D85594 BAA19776.1 LBMM01012027 KMQ86470.1 KZ149896 PZC78733.1 AGBW02011529 OWR46521.1 LBMM01006671 KMQ90407.1 LBMM01006954 KMQ90166.1 LBMM01004272 KMQ92618.1 LBMM01004262 KMQ92649.1 LBMM01015025 KMQ84953.1 LBMM01015285 KMQ84870.1 LBMM01008448 KMQ88867.1 LBMM01007036 KMQ90099.1 LBMM01006657 KMQ90417.1 LBMM01004634 KMQ92224.1 LBMM01002052 KMQ95336.1 LBMM01003988 KMQ92922.1 LBMM01008445 KMQ88871.1 LBMM01014122 KMQ85315.1 LBMM01008422 KMQ88894.1 LBMM01005474 KMQ91485.1 GGMR01015159 MBY27778.1 LBMM01012412 KMQ86235.1 LBMM01008278 KMQ89009.1 LBMM01008226 KMQ89040.1 GGMR01006838 MBY19457.1 ABLF02034925 LBMM01009610 KMQ87992.1 LBMM01009535 KMQ88053.1 LBMM01007425 KMQ89740.1 LBMM01006996 KMQ90130.1 ABLF02011238 LBMM01004261 KMQ92650.1 GALA01001777 JAA93075.1 LBMM01006977 KMQ90149.1 ABLF02013358 ABLF02013361 ABLF02054869 ABLF02009073 ABLF02030702 ABLF02042963 GGFJ01001369 MBW50510.1 GGFJ01001439 MBW50580.1 GGFJ01001370 MBW50511.1 GGFL01004044 MBW68222.1 ABLF02030663 ABLF02041312 GGFJ01001541 MBW50682.1 GALA01001776 JAA93076.1 GALA01000302 JAA94550.1 GGMR01019493 MBY32112.1 GDAI01000741 JAI16862.1 GGFK01005660 MBW38981.1 GGMR01012748 MBY25367.1 AB090815 BAC57906.1 ABLF02018098 ABLF02018942 ABLF02041885 ABLF02011183 ABLF02041884 GFXV01002090 MBW13895.1 AB593325 BAK38646.1 ABLF02023257 AB593327 BAK38648.1 GGFK01006563 MBW39884.1 GGFL01001916 MBW66094.1 ABLF02014650 LBMM01002480 KMQ94776.1 ABLF02027248 GGFJ01001453 MBW50594.1 GAMC01020519 GAMC01020515 JAB86040.1 GAMC01020517 JAB86038.1 GGFJ01001454 MBW50595.1 LBMM01003941 KMQ92970.1 ABLF02041886 ABLF02004854 ABLF02041317 KQ971388 KYB24913.1 KU543679 AMS38363.1 AB090817 BAC57910.1 GGFK01006866 MBW40187.1 KQ971675 KYB24771.1 AB593321 BAK38639.1 GFXV01003903 MBW15708.1 AB090822 BAC57920.1 ABLF02011136 ABLF02014426 KQ971309 EEZ99146.1 KU543683 AMS38371.1 GGFL01001192 MBW65370.1
Proteomes
Pfam
Interpro
Gene 3D
ProteinModelPortal
Q868Q4
Q8MY35
Q8MY31
Q8MY33
Q8MY27
Q8MY25
+ More
O01419 A0A0J7N1D9 A0A2W1C3J5 A0A212EYI2 A0A0J7KJG6 A0A0J7KIY5 A0A0J7KQK3 A0A0J7KQY4 A0A0J7K3F1 A0A0J7MWN5 A0A0J7KF37 A0A0J7KIM6 A0A0J7KJH4 A0A0J7KPR0 A0A0J7KYD2 A0A0J7KRB8 A0A0J7KF91 A0A0J7MY17 A0A0J7KF40 A0A0J7NFZ0 A0A2S2PEZ7 A0A0J7N0R3 A0A0J7KFH2 A0A0J7N8R4 A0A2S2NQQ6 J9LVU7 A0A0J7KCD9 A0A0J7KCE1 A0A0J7NAV8 A0A0J7NC28 X1WJY4 A0A0J7NJI3 T1DCM0 A0A0J7KIS5 J9JWJ3 J9KT61 A0A2M4BBV9 A0A2M4BC28 A0A2M4BC01 A0A2M4CS98 J9LQQ1 A0A2M4BCE1 T1DG89 T1DG59 A0A2S2PRL8 A0A0K8TR60 A0A2M4ADW9 A0A2S2P7D1 Q868S4 J9LA43 J9KM84 X1WZ64 A0A2H8TIE5 F7IYV5 J9KNN8 F7IYV7 A0A2M4AGH2 A0A2M4CL98 J9LM37 A0A0J7KWU5 J9L0J0 A0A2M4BC18 W8ADE4 W8ANK7 A0A2M4BC24 A0A0J7NKG1 J9KJG8 X1WXT5 A0A139WAH4 A0A142LX37 Q868S0 A0A2M4AHG6 A0A139WA31 F7IYU8 A0A2H8TR36 Q868R0 J9LB02 D6WC19 A0A142LX45 A0A2M4CJ33
O01419 A0A0J7N1D9 A0A2W1C3J5 A0A212EYI2 A0A0J7KJG6 A0A0J7KIY5 A0A0J7KQK3 A0A0J7KQY4 A0A0J7K3F1 A0A0J7MWN5 A0A0J7KF37 A0A0J7KIM6 A0A0J7KJH4 A0A0J7KPR0 A0A0J7KYD2 A0A0J7KRB8 A0A0J7KF91 A0A0J7MY17 A0A0J7KF40 A0A0J7NFZ0 A0A2S2PEZ7 A0A0J7N0R3 A0A0J7KFH2 A0A0J7N8R4 A0A2S2NQQ6 J9LVU7 A0A0J7KCD9 A0A0J7KCE1 A0A0J7NAV8 A0A0J7NC28 X1WJY4 A0A0J7NJI3 T1DCM0 A0A0J7KIS5 J9JWJ3 J9KT61 A0A2M4BBV9 A0A2M4BC28 A0A2M4BC01 A0A2M4CS98 J9LQQ1 A0A2M4BCE1 T1DG89 T1DG59 A0A2S2PRL8 A0A0K8TR60 A0A2M4ADW9 A0A2S2P7D1 Q868S4 J9LA43 J9KM84 X1WZ64 A0A2H8TIE5 F7IYV5 J9KNN8 F7IYV7 A0A2M4AGH2 A0A2M4CL98 J9LM37 A0A0J7KWU5 J9L0J0 A0A2M4BC18 W8ADE4 W8ANK7 A0A2M4BC24 A0A0J7NKG1 J9KJG8 X1WXT5 A0A139WAH4 A0A142LX37 Q868S0 A0A2M4AHG6 A0A139WA31 F7IYU8 A0A2H8TR36 Q868R0 J9LB02 D6WC19 A0A142LX45 A0A2M4CJ33
PDB
1WDU
E-value=5.56599e-12,
Score=175
Ontologies
GO
Topology
Length:
926
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.16959
Exp number, first 60 AAs:
0.00086
Total prob of N-in:
0.00497
outside
1 - 926
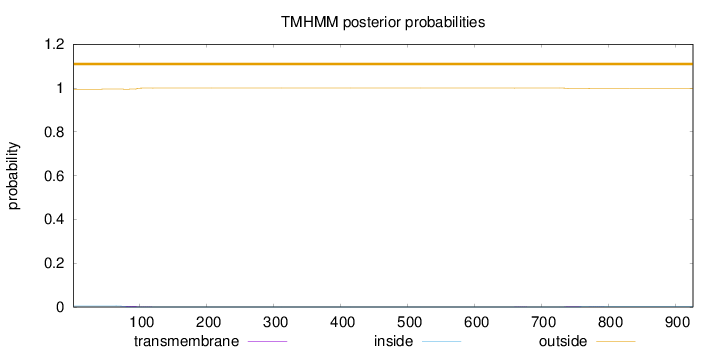
Population Genetic Test Statistics
Pi
11.479583
Theta
13.027453
Tajima's D
-2.292113
CLR
308.15499
CSRT
0.00259987000649968
Interpretation
Uncertain
