Gene
KWMTBOMO02398
Annotation
PREDICTED:_LOW_QUALITY_PROTEIN:_uncharacterized_protein_LOC106708942_[Papilio_machaon]
Location in the cell
Mitochondrial Reliability : 1.088 Nuclear Reliability : 1.783
Sequence
CDS
ATGCAGATCGTTGAGTCTCACATACCCCGCGCGCCCCTTTTGCGCGGGGACCTCGTAGGAGGTTCGGCCCTCTACCCGAAAAAAAAAAAAAAAAAAAAAAAAAAGGAGGGTACCTCCCGCTCAGCGGGAGGAGAATCCCTCCGTGTGGCCGACTCGTCGGACGCACGCTGCCCCGCGTACTCTGGGGGGGGCAAAAACTGCCTAGACGTCAGGTCCGGGAAGGACGGAGTTCAAAAGAAGAAAATTATGTCCGAGAAGGACCCTGAAGTAAGCTCCTTAAGAGCCGATGTCGTGTCGGACAACGATATGGCCACAGTTGGTTCGGCCCAGTCGTCGATCCATTCGTCGCCGACACGACGACGGCGGAAAAGGGTGCTCAATATAAGTTCGGACGGGCTGAGTAGCAGCTCCGGCTCGGAGGCTCAGGATCCGACACGTGTGAAGCCCAGAGCAGTCACGCCGTCAAAGCGTGGACGGGGACGCCCTGCGACCACAGGGCAGTACGTTGGCATGGCCGCTGCCAGGCAAGCGTACCTCAAGGCACAAAAAGAGGAGAAGGAGCTCACTGAGAGCGCGCGCACCACACGCCACAAGAGGAGAGCGGCTGAAGGGTCTGTCCCTAAGCCCGATGCCGAGCGTAGGCTCCATCAAAGTGCAGAGGTGGCCCTGATGGTCGCCGCCAAATCCAGCAGCCCGAAGGGCACGTACGTGCGTGCACTGAAGGAGTTGGCAGTCACTGTCAAGGAGGCCACGGAGGACCTTGTGGGCTACACCGCAGCCGAAGAGGCTAAGAGACTGCGAGCGGTCAACGTCAAGCAGGAGCAGGAGATCCTGCAACTGCGTAAGGAGCTCGATGAAGTTAGAAGGGAGTTGGCGAGGGTTTCGCGGGTGCTAGCGTTGCCCACATCGACTGCGACCCCGGCCCCGAACACCAAGGAGGAAGAGGAGGAGTTGCGCCTACAGCGTATTATGCGAGCTGTCGGCACTATGCTCGACGCTCGCTTTGAAGGGCTGGAGGCACGGCTTCTACCGGAGCCCCGAATGCGACCACCGCTAGCGGCGGACAAGCAGAGGAGCCGGGAAAAATCGTCTAATCCCCCTGTCGTCGTTGACGCTGCACCGGATGCGGCACCGTCATCAGTGCCACCGATGACATTGGCGAAAAAGAGGATGCGTGGCCCCAAGAGAAGTGTCACCGCGCAAGCGGCTTCATCCGAAACGCACACCTTCCTCCCGGCTCCAGCTTCCATGATCGAGAGCTGGTCAACGGTCACCCGTAGGGATGCTCGACCGAGGGAGGAAGCAGTGAGGACGCTTCCAGCGGCAGCTATCGCTGAAGCGGAAAGGAAAGACCGGTCTTCTGTAAAGAGGAAAGAGAAGAAGCTGCGCCCTCCACGCACTCAGGCCGTTGTCCTCAAGCTGCAACCAGAGGCCGTGGAAAGAGGACTCACGTACCGCACTGTACTCGCCGAAGCGAGGGCCAAGGTAGATCCTGGGGCACTTGGAATACCCATCCAGCGCATCCGGTCTGCAGTGACTGGGTCTAAGGTGCTGGTAGTGGAAGGTGCAGACCAGAGCGCCAAGGCCGATCTCCTGGCCCAAAAGCTTCGTGAGGTTCTGCCCGCAGAAGGGGTCATAGTGACCAGGCCAGTAACGACTGCAGCCATCCGCATCAGCGGACTGGACGACTCCCTGATGCCGGAGGAACTTACCGCGGAGCTTGCCCGGATTGGCGAATGCGCTCCTGGTGCGGTGAAGATCGGAGACATCAAGTATGGCCCAGGCGGGATGGGTCAAGTATTGGTCCGGTTGCCCGTCGTTGCCACGATGAAGGTTCTCGCCGTTCCAAAACTGAGGGTTGGTTGGAGCGTGCTCCGCGCTTGTTTGTTAGAGGCTAAGAAGCTACAGTGCTTCCGATGCCATGAGCTAGGGCACGTGAGCGCCCGATGTCCTTCGTCGGTCGACCGCAGCTGGGATTGTTATCGGTGCGGCCAGACCGGCCACGTAGCGGCCGGCTGCTCTCTTGCACCACACTGCGCTGGGAATCTCAACCATTGCGCCAGAGCTCAGGACCTTTTGTTCCAGAGCATGGCGGAGGGGTTGACCCATCTTGCGGTGGTCGCCGAGCCGTATCGGGTCCCTTCGAGCCCCGATTGGGCGGCCGATTTGGAGGGCCGAGTGGCAATCATCCGGCGTTGTTGTGTGGGTGCTCCGCCTACGTTTGCCGTTGTTGAAAGAGGTCGCGGCTTCGTTGCCGTCCTCTGGGCCGAGATATTCGTGTTGGGAGTGTACTTTTCCCCAAACAGGACGCTCGCCGAGTTTGAGGTTTTCCTCAGCGAGCTCAGCCGCGTCGTTGGAAGGTCGCACTTCCGACGGATTCTCGTTCTAGGTGACCTAAATGCCAAGTCATTGTCTTGGGGTTCCTCTAGGACGTGCCCCAGAGGTAGAGCGATGGAGGAGTGGCTGGTCGGAAGCGCTCTCCTCATCCTCAATCGCGGTACGGAACTCACATGCGTGCGACGTTTGGACGGGTCCGTGGTGGACGTTACATTCGCCACGCCTGACGTCGCATATCGCGTACGCGGTTGGGCGGTGATGGTTGGCGAGGAGACCCTCTCCGACCACCGTTACATCCGATTCAGTGTCGCCGCGACCCCGGTGGGTTCTGTTCGGGGCTTTTCCTTGCCCATCAGTGGCAGCGCTAGGGGTCCTCGTTGGGCCCAGAAGCGCCTCAACATCGAGCGGTTGCGTGAGGCGGCCATAGTGCAGGCGTGGCGTCTTGACTCGCTTGGCGAGCCAGCAGACGTGTGCGAGGGGGTGGAGCGTCTGCGCGAGGCAATGTCACGGGTGTGTGACGCTGCTATGCCTCGCGTGAGAGCTCTCGCTCCTAAGCGCCAAGTTCATTGGTGGACCGAGGAGATCGCCAGCCAGCGCCGGCTATGCGATATTAGTCGTCGCACATACCAGCGATATCGACGACGAAGAACGCGCCGGGACCCCGACGAGGAGGACCGCCTGTACGAAGTGTACAGGACGGCGATAAGAACCCTGCGCTTGGCTATCGGGGAGGCGAAGGAGGCCGCCTGGAACGACCTACTGGCCTCGCTGGACCGTGACCCGTGGGGGCGGCCCTACAGGCTGGCGCGTAATGCGCTTCGCCGTTGGGCTCCTCCCGCAACCAGCACCCTGCCGCCGGAAACATTGCAGCGGGTAGTCGGGGGTCTGTTTCCCGATTTATCCGGGACGGCCTTCGTCCCCCCTGTGATGACAATGGCGCGGATTGCTGATAGCGAAGAGCTTGAGGACGTCTCCCAGGTGGAGTTCGATTTGGCTGTGCAGAGGATGCGGGCTAAGCGCACGGCACCCGGTCCCGATGGGATCTCGTCCCAAGCGTGGGCGCTCGCCTTGACAGGTGACGGCTTGGGGCCTGCCCTCCAAGGGCTATTCAGCAGGTGCCTCCGTGAGGGCAGGTTCCCAGAGCCATGGAAGACTGGTCGGCTTGTCCTTATTCCTAAGGAAGGCCGGCCACGTGACGAGCCAAGCGGGTACCGCCCAATCGTCGTGCTGGACGAGGCTGGTAAGCTCCTCGAGCGCATCGTCGCCAGTCGCCTCGTCCAGCACCTCGAAAGCGTTGGGCCTGACCTGGCTCCTAACCAATATGGTTTCCGGAGAGGTCGCTCCACCATGGACGCGGTCTTGCGCGTCCGCCACCTCTCCGATCGTGCGTGCTCCGAGGGGGGCGTGTTGTTGGCTGTGTCGATTGATATCGCCAACGCCTTCAACACGATCCCTTGGAGCACGATCGTGGAATCGCTCCGGTTTCACCGCGTCCCCCCTAGTCTCCGCACCCTGATAGAGGATTACCTCTCAGGGCGAAATGTAGTCTTCCCCGAGAGGAGGGGGTGGGGACGGAAAGCGGTGTCGTGCGGGGTCCCGCAGGGGTCGGTACTGGGACCACTCCTGTGGGACATCGGTTTCGACTGGGTCCTGCGCGGTGCTAGCCTGCGTGGCGTCGACGTAGTGTGCTATGCCGACGACACGCTGGTGACGGCCCGCGGAGCCGACTACAGAGCCGCAGCGATCCTTGCGACGGCGGCGGTCTCCACCGTCGTTAGTCGCATTCGGAGATTAGGTCTTGAGGTGGCCCTCCGCAAGTCCGAAGCGGTGTGCTTTCACCCAGTCCGGAGGAGACCTCCTCCGGGAGCGAATCTCATAGTCGGCGGAGTATCGATCGCTGTCCAGCCGAAGCTCAAATATTTGGGCCTTGTGCTGGACAGTCGATGGCGCTTCGACCACCACTTTGGTGAGTTAGTCCCAAAGCTGCTGGGGATGGCGGGCGCACTAGCCCGTCTTCTCCCCAACGTCGGTGGTTGCAGCGCCGGCGTCCGGCGTCTGTACCTGGGGGTCGTGCGCAGTATGGCTTTGTACGGCGCTCCCGTGTGGTCGCCCGCACTCTCTGCGCGCAATGCAGCTTTGCTGCTACGAGTGCAGCGGGTGCTCGCGGTGAGAGTCATCAGGGGGTACCGTACGATCTCCCGGGAGGTCGCCTGCGCCCTCGCCGGTTCCCTTCCTTGGGATCTCGAGGCTGAGGTAGCAGCTGCGGTATACCGGCGTAGAACACAGTCCCTGAGTCGGGGACGGACGCCCGGCCCGTCGGCTGTCGGTCGGTGGAGGCGTGCTGCGCGTCATCTGGCGTACGCCAAGTGGAGGGAGCGGTTGCTGGAGGAGCTTGGCCAAGCTTCAGTCACTCGCCGACGCACCCTCGAGGCCCTGGTGCCCGTGCTGGAGGCATGGTCAGATAGACGACACGGCGTGCTCTCCTTCCACTTAACGCAGGTCCTCTCGGGGCATGGCTGCTTCGGGAGGTACCTGTGGAAGGTCTGCAGGAGAGAGCCACATCCGGGTTGCCACCAGTGCGGGCATCCGGACGATGACGCTCAGCACGCACTCGAAGCGTGCCCGCGTTGGGAGCGCTCGCGGCGAGACCTCATCGCGGTGCTGGGGAGAGACCTCTCCTTGCGGGCTGTCATCGCCCGCATGCTAGAGGACCAGAATTCTTGGGAGGCCATGACCAGGTTTTGCGACCTGGTCATGGCGTTCCGTGAGGCTGAGGAGCGCGAAAGGGAGTGGGATCCCTCTTCGGCTTCACAGCGAAGAAGGCGGAGTGGGCGGCACGGGCGCGAAGCTCCCGTTCATCTCCCGTAG
Protein
MQIVESHIPRAPLLRGDLVGGSALYPKKKKKKKKKEGTSRSAGGESLRVADSSDARCPAYSGGGKNCLDVRSGKDGVQKKKIMSEKDPEVSSLRADVVSDNDMATVGSAQSSIHSSPTRRRRKRVLNISSDGLSSSSGSEAQDPTRVKPRAVTPSKRGRGRPATTGQYVGMAAARQAYLKAQKEEKELTESARTTRHKRRAAEGSVPKPDAERRLHQSAEVALMVAAKSSSPKGTYVRALKELAVTVKEATEDLVGYTAAEEAKRLRAVNVKQEQEILQLRKELDEVRRELARVSRVLALPTSTATPAPNTKEEEEELRLQRIMRAVGTMLDARFEGLEARLLPEPRMRPPLAADKQRSREKSSNPPVVVDAAPDAAPSSVPPMTLAKKRMRGPKRSVTAQAASSETHTFLPAPASMIESWSTVTRRDARPREEAVRTLPAAAIAEAERKDRSSVKRKEKKLRPPRTQAVVLKLQPEAVERGLTYRTVLAEARAKVDPGALGIPIQRIRSAVTGSKVLVVEGADQSAKADLLAQKLREVLPAEGVIVTRPVTTAAIRISGLDDSLMPEELTAELARIGECAPGAVKIGDIKYGPGGMGQVLVRLPVVATMKVLAVPKLRVGWSVLRACLLEAKKLQCFRCHELGHVSARCPSSVDRSWDCYRCGQTGHVAAGCSLAPHCAGNLNHCARAQDLLFQSMAEGLTHLAVVAEPYRVPSSPDWAADLEGRVAIIRRCCVGAPPTFAVVERGRGFVAVLWAEIFVLGVYFSPNRTLAEFEVFLSELSRVVGRSHFRRILVLGDLNAKSLSWGSSRTCPRGRAMEEWLVGSALLILNRGTELTCVRRLDGSVVDVTFATPDVAYRVRGWAVMVGEETLSDHRYIRFSVAATPVGSVRGFSLPISGSARGPRWAQKRLNIERLREAAIVQAWRLDSLGEPADVCEGVERLREAMSRVCDAAMPRVRALAPKRQVHWWTEEIASQRRLCDISRRTYQRYRRRRTRRDPDEEDRLYEVYRTAIRTLRLAIGEAKEAAWNDLLASLDRDPWGRPYRLARNALRRWAPPATSTLPPETLQRVVGGLFPDLSGTAFVPPVMTMARIADSEELEDVSQVEFDLAVQRMRAKRTAPGPDGISSQAWALALTGDGLGPALQGLFSRCLREGRFPEPWKTGRLVLIPKEGRPRDEPSGYRPIVVLDEAGKLLERIVASRLVQHLESVGPDLAPNQYGFRRGRSTMDAVLRVRHLSDRACSEGGVLLAVSIDIANAFNTIPWSTIVESLRFHRVPPSLRTLIEDYLSGRNVVFPERRGWGRKAVSCGVPQGSVLGPLLWDIGFDWVLRGASLRGVDVVCYADDTLVTARGADYRAAAILATAAVSTVVSRIRRLGLEVALRKSEAVCFHPVRRRPPPGANLIVGGVSIAVQPKLKYLGLVLDSRWRFDHHFGELVPKLLGMAGALARLLPNVGGCSAGVRRLYLGVVRSMALYGAPVWSPALSARNAALLLRVQRVLAVRVIRGYRTISREVACALAGSLPWDLEAEVAAAVYRRRTQSLSRGRTPGPSAVGRWRRAARHLAYAKWRERLLEELGQASVTRRRTLEALVPVLEAWSDRRHGVLSFHLTQVLSGHGCFGRYLWKVCRREPHPGCHQCGHPDDDAQHALEACPRWERSRRDLIAVLGRDLSLRAVIARMLEDQNSWEAMTRFCDLVMAFREAEEREREWDPSSASQRRRRSGRHGREAPVHLP
Summary
Uniprot
EMBL
LBMM01004272
KMQ92618.1
LBMM01005474
KMQ91485.1
LBMM01012412
KMQ86235.1
+ More
LBMM01003988 KMQ92922.1 LBMM01008278 KMQ89009.1 ABLF02011238 LBMM01006977 KMQ90149.1 ABLF02018942 ABLF02041885 ABLF02011183 ABLF02041884 KQ971388 KYB24913.1 KQ971310 KYB29499.1 KYB29500.1 ABLF02007989 ABLF02018098 ABLF02004854 ABLF02041317 ABLF02018808 ABLF02012448 ABLF02012450 ABLF02012455 ABLF02012460
LBMM01003988 KMQ92922.1 LBMM01008278 KMQ89009.1 ABLF02011238 LBMM01006977 KMQ90149.1 ABLF02018942 ABLF02041885 ABLF02011183 ABLF02041884 KQ971388 KYB24913.1 KQ971310 KYB29499.1 KYB29500.1 ABLF02007989 ABLF02018098 ABLF02004854 ABLF02041317 ABLF02018808 ABLF02012448 ABLF02012450 ABLF02012455 ABLF02012460
Proteomes
Interpro
Gene 3D
ProteinModelPortal
PDB
1WDU
E-value=1.59601e-06,
Score=130
Ontologies
GO
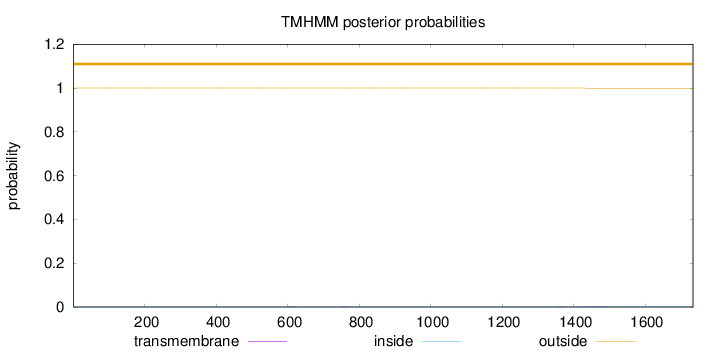
Topology
Length:
1734
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.05311
Exp number, first 60 AAs:
0.00039
Total prob of N-in:
0.00008
outside
1 - 1734
Population Genetic Test Statistics
Pi
0
Theta
0
Tajima's D
0
CLR
102.652478
CSRT
0
Interpretation
Uncertain
