Gene
KWMTBOMO02349
Pre Gene Modal
BGIBMGA014553
Annotation
hypothetical_protein_TcasGA2_TC034914?_partial_[Tribolium_castaneum]
Location in the cell
Cytoplasmic Reliability : 1.898 Mitochondrial Reliability : 1.424
Sequence
CDS
ATGGTTTTTAAAGTGTTACAGATAAATATAGACCGAGGGAAAGCTGCGCACGACCTACTTCTGGCAACTGCGAGCAGTATGAACGCGGATCTCGTGATGCTCTCTGAACCCAATAGAAAACTGGCAAAAGCTGCTGGATGGGAGTGCGACACCAGGGGAGATGCAGCCCTAGTGATTCTTAACAAAACACTGACGGTTTCTGAAATTTATAGGGGGAAGGGATTTGTTGGGTTCAAGATGAAGCACTTTGCCGCGTTCAGCTGTTACATATCTCCCAACTGCAGTTTTGAGGAGTTCAAAGCTGACGTCGATGAGCTTTGGACACTACTTAAAAAACAAAAGACACCGATCCTAGTGGCAGGAGATTTTAACTCTGCATCGAGAGCGTGGGGCAGTAAAGTAACCAACAAAAGGGGCCACTACCTGCTGGAAATGGTGGCGGACCTCGACACTGTCATCATGAATGAAGGCGTGACCCCAACTTTTGAAAGAGGAGACAGAACATCCAGGCCCGACATCACCATGGCCTCTGAAACGCTGGCAGGCAAGATCAGTAATTGGCAAGTGCTCGACGAGGAGACCATGAGCGGACATAAATATATAGCCTACGAGATAAAGCATGTATTGCCTCCATCCCAGTGCTATGCACAGATGAAAAGGTGGAACCTCCAAAGGCTGGACGTAGATAAGTTCAGTAAAAGCCTGAAAGGAGCGCAGATCTCGACTCCCATGGAACTGGTGCAGGCTACGACGGCAGCGTGTGACGCTTCTATGCCAAAGCAAAACAACCGCAAGAAGAAAGGAGTCTACTGGTGGAACAAGGAAATTGCGGAAGCAAGGAAGGAGTGTTTGAAATGCCGAAGGAGGTACACAAGGAATAACAAAAAGCTAGGAAAAGGGAAAGACAGGGAAAGTAACGAAGAAAATGAACACAAAAAGCAGTATAAGGAAGCGAAACTGGAGCTACAAAGACGAATATGGAAAGCGAAAGACGATAAATGGAGGGAAGTGTGCGAGGAAGTCGACAATGATATATGGGGACTGGGATACAAAATAGTGTCAAAAAAGCTACGCGGAAACTCACCAATAAATATGGGCATGGCCCAAATTCAAAGCATAGCTGATGGCCTCTTCCCGGAACACATAGATTTCGAACGGAAAACGAAACCGTCACAGGAGTTCCCAGAGGTGACCGAGGAGGAGATCATCCAACTAAAAGCTCGCATAAAGCTTAGGAAAGCCCCAGGGCCCGACCGGATTCCGCCGGAGATAGTGACTCTAGCAGTAGCAGTAATTCCAGACAGGATAAGGCTTGTGTTGCAGAAGGTGCTGGAGTCTGGCGTCTTCCCCAACAGCTGGAAAACAGCCAGACTAGTCCTGCTAAGAAAACCAGGCAAACCCGCTGAAGATCCTGCGGGGCACAGGCCGCTATGTCTACTGGACTGCTACGGAAAGCTGCTGGAGTCGCTGATCCTAGGCAGAATAGAAAAACAACTAGTGCATTCGGACTGTTTACAAGATTCACAGTACGGGTTCAGAAAGGGGAGGTCCACAGTGGATGCAGCCAACAAGCTACTTGAGATAGCGGAAGACGCGAGACGAGGAACCCACCGCACCAGAAGAATCTGCGCAGTAGTGGCCCTCGACGTCCGTAATGCTTTCAATTCTGCATCGTGGGAGCTCATCCTGGAGGAAATGGCTGCAATAGGTATGCCGCAATACATCTATAATATCATAAACAGCTACCTCCAGGACAGATGCATTATACTGACAGACGCCGAGGGAGTGGTTGTAGAGAAAAAGGTTACCGGTGGTGTTCCGCAGGGCTCGATACTGGGCCCCACGCTGTGGAACATATTGTATGATGGGGTACTGCGCCTCGAACTGGGGGACGGGGCACAATTAATAGGCTTCGCTGACGATTTAGCTCTGGTGGTGTCAGCAAAGAAGGAAGCTGAACTGATGTCAATAGCGAATGTAGCCCTAGAGAAGATCAGCGCTTGGATGAAGCAAAAGCGGCTACACTTGGCCCCAGAAAAGACCGAGGCAATTGTGCTAAGTGGGCGAAGAAAGCTTAGCGAGATTTGCTTTACCCTCCAGGGCGTCAGGGTGGTTCCAACCGACAAACTGAAATACCTCGGGTTTTGGTTTGACAAAAGTCTGAAGTTCGGCAAACACATTGAAGAAACAACTGGTAAAGCCGAAAGACTGACGACCGCCCTTGGCCAAATCATGCCCAGGAAAGGAGGGCCGAGGTCGAGCAAGCGCAGACTATTGGCGTCCGCAATACAATCTGTGATGTTATATGGAGCACCGGTGTGGGCAAAGGTAATGAATGTAGCGAAGTACAGGAACATGTATAGCAAAGTGCAACGACAGCTGGCAATACGCATCTGTGGAGCTTACCGTACCATATCGCTAGACGCCGTCTTGGTGATAGCGGGCCTGATACCGATAGAACGGCTAGCAAAGGAACGGCAACGCTACTACCACGAGGGGACAAGAACAAGAGAAGAGGAAAGAGCACATACAATAAAAGAATGGGAACGAGACTGGCAGAAGGAGGGAAAAGGTAAGTGGACTCGTACAATCATACCCAACCTTAGGCTCTGGATGGGACGGAAACATGGCGAAACCGACAGACTGCTGACACAGGCCCTAAGTGGGCATGGAGTTTTCAACACTTACCTGCATGCAATAGGGAAAGCAGAAAGCGCAAGTTGCCCATACTGCGAACATGTTGACACACCGGAACATGCCCTATTTCAATGCATAAAGTTCTCAGAACTAAGAGAGAAAACGGAGGGAAGTGACGGAAAGGTGACTGTGGAAAACCTGATGGTGAGAATGATCCATAGTAAAGAAGAATGGGATAAATATGCGGGAATGCTCAAAGAGATAATGAGAGTGCGGGAAAAGGACGAAAGACTAAGGCAAAAATCTATCTATAATAATATATAA
Protein
MVFKVLQINIDRGKAAHDLLLATASSMNADLVMLSEPNRKLAKAAGWECDTRGDAALVILNKTLTVSEIYRGKGFVGFKMKHFAAFSCYISPNCSFEEFKADVDELWTLLKKQKTPILVAGDFNSASRAWGSKVTNKRGHYLLEMVADLDTVIMNEGVTPTFERGDRTSRPDITMASETLAGKISNWQVLDEETMSGHKYIAYEIKHVLPPSQCYAQMKRWNLQRLDVDKFSKSLKGAQISTPMELVQATTAACDASMPKQNNRKKKGVYWWNKEIAEARKECLKCRRRYTRNNKKLGKGKDRESNEENEHKKQYKEAKLELQRRIWKAKDDKWREVCEEVDNDIWGLGYKIVSKKLRGNSPINMGMAQIQSIADGLFPEHIDFERKTKPSQEFPEVTEEEIIQLKARIKLRKAPGPDRIPPEIVTLAVAVIPDRIRLVLQKVLESGVFPNSWKTARLVLLRKPGKPAEDPAGHRPLCLLDCYGKLLESLILGRIEKQLVHSDCLQDSQYGFRKGRSTVDAANKLLEIAEDARRGTHRTRRICAVVALDVRNAFNSASWELILEEMAAIGMPQYIYNIINSYLQDRCIILTDAEGVVVEKKVTGGVPQGSILGPTLWNILYDGVLRLELGDGAQLIGFADDLALVVSAKKEAELMSIANVALEKISAWMKQKRLHLAPEKTEAIVLSGRRKLSEICFTLQGVRVVPTDKLKYLGFWFDKSLKFGKHIEETTGKAERLTTALGQIMPRKGGPRSSKRRLLASAIQSVMLYGAPVWAKVMNVAKYRNMYSKVQRQLAIRICGAYRTISLDAVLVIAGLIPIERLAKERQRYYHEGTRTREEERAHTIKEWERDWQKEGKGKWTRTIIPNLRLWMGRKHGETDRLLTQALSGHGVFNTYLHAIGKAESASCPYCEHVDTPEHALFQCIKFSELREKTEGSDGKVTVENLMVRMIHSKEEWDKYAGMLKEIMRVREKDERLRQKSIYNNI
Summary
Uniprot
A0A139WAH4
F7IYV5
A0A139W940
F7IYU8
J9KJG8
A0A139W9B3
+ More
D6WC19 A0A139W939 F7IYV2 V5I881 A0A2H8TR36 A0A139WNC9 A0A139WN93 A0A139W9J0 A0A224XIZ2 T1DG59 T1DCM0 T1DG89 F7IYV6 A0A142LX45 J9KUM6 A0A2M4CS98 A0A2M4ADW9 A0A2M4BBV9 J9JWJ3 A0A2S2NQQ6 J9LVU7 A0A2M4BC01 A0A2M4BC28 A0A034WR48 A0A2M4BC24 A0A139W9A6 A0A2M4BC18 A0A2H8TIE5 A0A142LX43 A0A142LX37 A0A0K8VPH1 W8ANK7 X1WZ64 W8ADE4 J9KM84 F7IYV3 A0A2M4BDI3 A0A2M4CSA9 Q868S4 F7IYV7 A0A2M4CSI6 A0A1W7R6I9 A0A023EW96 A0A139W8G2 A0A2S2PRL8 A0A0K8S5I5 D7EIB0 A0A139W942 Q868S6 A0A2M4BB78 A0A2M3Z1C1 A0A2M4BB89 A0A139WA31 A0A2M4BBA0 Q9N9Z1 A0A1S4EMG7 J9L6Z9 A0A2S2PEZ7 A0A2M4BCE1 Q868S0 A0A139W8J5 X1WXT5 Q868T0 A0A2M4AK35 J9LM37 J9KT61 K7JP42 J9K6F0 J9JY94 X1WJY4 A0A2S2NJ82 J9KNS1 A0A2M4CJ33 J9KNN8 Q868S8 V9GZD3 Q868R6 A0A2S2P7D1 Q868R2 A0A2S2P488 Q868R0 A0A0J7KIY5 W8AMZ9 A0A2M4AGH2 J9LBG0 A0A2M4CL98 Q868R4 J9LQQ1 V5G2Q2 A0A2M4CJ59 A0A2M4CKD9 A0A2M4CKQ7
D6WC19 A0A139W939 F7IYV2 V5I881 A0A2H8TR36 A0A139WNC9 A0A139WN93 A0A139W9J0 A0A224XIZ2 T1DG59 T1DCM0 T1DG89 F7IYV6 A0A142LX45 J9KUM6 A0A2M4CS98 A0A2M4ADW9 A0A2M4BBV9 J9JWJ3 A0A2S2NQQ6 J9LVU7 A0A2M4BC01 A0A2M4BC28 A0A034WR48 A0A2M4BC24 A0A139W9A6 A0A2M4BC18 A0A2H8TIE5 A0A142LX43 A0A142LX37 A0A0K8VPH1 W8ANK7 X1WZ64 W8ADE4 J9KM84 F7IYV3 A0A2M4BDI3 A0A2M4CSA9 Q868S4 F7IYV7 A0A2M4CSI6 A0A1W7R6I9 A0A023EW96 A0A139W8G2 A0A2S2PRL8 A0A0K8S5I5 D7EIB0 A0A139W942 Q868S6 A0A2M4BB78 A0A2M3Z1C1 A0A2M4BB89 A0A139WA31 A0A2M4BBA0 Q9N9Z1 A0A1S4EMG7 J9L6Z9 A0A2S2PEZ7 A0A2M4BCE1 Q868S0 A0A139W8J5 X1WXT5 Q868T0 A0A2M4AK35 J9LM37 J9KT61 K7JP42 J9K6F0 J9JY94 X1WJY4 A0A2S2NJ82 J9KNS1 A0A2M4CJ33 J9KNN8 Q868S8 V9GZD3 Q868R6 A0A2S2P7D1 Q868R2 A0A2S2P488 Q868R0 A0A0J7KIY5 W8AMZ9 A0A2M4AGH2 J9LBG0 A0A2M4CL98 Q868R4 J9LQQ1 V5G2Q2 A0A2M4CJ59 A0A2M4CKD9 A0A2M4CKQ7
Pubmed
EMBL
KQ971388
KYB24913.1
AB593325
BAK38646.1
KQ972691
KXZ75806.1
+ More
AB593321 BAK38639.1 ABLF02041886 KQ972544 KXZ75869.1 KQ971309 EEZ99146.1 KXZ75807.1 AB593323 BAK38643.1 GALX01005145 JAB63321.1 GFXV01003903 MBW15708.1 KQ971310 KYB29500.1 KYB29499.1 KQ972226 KYB24577.1 GFTR01008427 JAW07999.1 GALA01000302 JAA94550.1 GALA01001777 JAA93075.1 GALA01001776 JAA93076.1 AB593326 BAK38647.1 KU543683 AMS38371.1 ABLF02041557 GGFL01004044 MBW68222.1 GGFK01005660 MBW38981.1 GGFJ01001369 MBW50510.1 ABLF02013358 ABLF02013361 ABLF02054869 GGMR01006838 MBY19457.1 ABLF02034925 GGFJ01001370 MBW50511.1 GGFJ01001439 MBW50580.1 GAKP01002292 JAC56660.1 GGFJ01001454 MBW50595.1 KXZ75870.1 GGFJ01001453 MBW50594.1 GFXV01002090 MBW13895.1 KU543682 AMS38369.1 KU543679 AMS38363.1 GDHF01029467 GDHF01011533 JAI22847.1 JAI40781.1 GAMC01020517 JAB86038.1 ABLF02011183 ABLF02041884 GAMC01020519 GAMC01020515 JAB86040.1 ABLF02018942 ABLF02041885 AB593324 BAK38644.1 GGFJ01001727 MBW50868.1 GGFL01004046 MBW68224.1 AB090815 BAC57906.1 AB593327 BAK38648.1 GGFL01004047 MBW68225.1 GEHC01000915 JAV46730.1 GAPW01000192 JAC13406.1 KQ973343 KXZ75566.1 GGMR01019493 MBY32112.1 GBRD01017378 JAG48449.1 AB593322 BAK38641.1 KXZ75805.1 AB090814 BAC57904.1 GGFJ01001164 MBW50305.1 GGFM01001554 MBW22305.1 GGFJ01001165 MBW50306.1 KQ971675 KYB24771.1 GGFJ01001166 MBW50307.1 AJ278684 CAB99192.1 ABLF02021100 GGMR01015159 MBY27778.1 GGFJ01001541 MBW50682.1 AB090817 BAC57910.1 KQ973256 KXZ75595.1 ABLF02004854 ABLF02041317 AB090812 BAC57900.1 GGFK01007771 MBW41092.1 ABLF02014650 ABLF02009073 ABLF02030702 ABLF02042963 AAZX01012143 AAZX01012194 ABLF02012448 ABLF02012450 ABLF02012455 ABLF02012460 ABLF02007989 ABLF02011238 GGMR01004600 MBY17219.1 ABLF02018808 GGFL01001192 MBW65370.1 ABLF02023257 AB090813 BAC57902.1 M93690 AAA29363.1 AB090819 BAC57914.1 GGMR01012748 MBY25367.1 AB090821 BAC57918.1 GGMR01011682 MBY24301.1 AB090822 BAC57920.1 LBMM01006954 KMQ90166.1 GAMC01019173 JAB87382.1 GGFK01006563 MBW39884.1 ABLF02021645 GGFL01001916 MBW66094.1 AB090820 BAC57916.1 ABLF02030663 ABLF02041312 GALX01004171 JAB64295.1 GGFL01001189 MBW65367.1 GGFL01001190 MBW65368.1 GGFL01001583 MBW65761.1
AB593321 BAK38639.1 ABLF02041886 KQ972544 KXZ75869.1 KQ971309 EEZ99146.1 KXZ75807.1 AB593323 BAK38643.1 GALX01005145 JAB63321.1 GFXV01003903 MBW15708.1 KQ971310 KYB29500.1 KYB29499.1 KQ972226 KYB24577.1 GFTR01008427 JAW07999.1 GALA01000302 JAA94550.1 GALA01001777 JAA93075.1 GALA01001776 JAA93076.1 AB593326 BAK38647.1 KU543683 AMS38371.1 ABLF02041557 GGFL01004044 MBW68222.1 GGFK01005660 MBW38981.1 GGFJ01001369 MBW50510.1 ABLF02013358 ABLF02013361 ABLF02054869 GGMR01006838 MBY19457.1 ABLF02034925 GGFJ01001370 MBW50511.1 GGFJ01001439 MBW50580.1 GAKP01002292 JAC56660.1 GGFJ01001454 MBW50595.1 KXZ75870.1 GGFJ01001453 MBW50594.1 GFXV01002090 MBW13895.1 KU543682 AMS38369.1 KU543679 AMS38363.1 GDHF01029467 GDHF01011533 JAI22847.1 JAI40781.1 GAMC01020517 JAB86038.1 ABLF02011183 ABLF02041884 GAMC01020519 GAMC01020515 JAB86040.1 ABLF02018942 ABLF02041885 AB593324 BAK38644.1 GGFJ01001727 MBW50868.1 GGFL01004046 MBW68224.1 AB090815 BAC57906.1 AB593327 BAK38648.1 GGFL01004047 MBW68225.1 GEHC01000915 JAV46730.1 GAPW01000192 JAC13406.1 KQ973343 KXZ75566.1 GGMR01019493 MBY32112.1 GBRD01017378 JAG48449.1 AB593322 BAK38641.1 KXZ75805.1 AB090814 BAC57904.1 GGFJ01001164 MBW50305.1 GGFM01001554 MBW22305.1 GGFJ01001165 MBW50306.1 KQ971675 KYB24771.1 GGFJ01001166 MBW50307.1 AJ278684 CAB99192.1 ABLF02021100 GGMR01015159 MBY27778.1 GGFJ01001541 MBW50682.1 AB090817 BAC57910.1 KQ973256 KXZ75595.1 ABLF02004854 ABLF02041317 AB090812 BAC57900.1 GGFK01007771 MBW41092.1 ABLF02014650 ABLF02009073 ABLF02030702 ABLF02042963 AAZX01012143 AAZX01012194 ABLF02012448 ABLF02012450 ABLF02012455 ABLF02012460 ABLF02007989 ABLF02011238 GGMR01004600 MBY17219.1 ABLF02018808 GGFL01001192 MBW65370.1 ABLF02023257 AB090813 BAC57902.1 M93690 AAA29363.1 AB090819 BAC57914.1 GGMR01012748 MBY25367.1 AB090821 BAC57918.1 GGMR01011682 MBY24301.1 AB090822 BAC57920.1 LBMM01006954 KMQ90166.1 GAMC01019173 JAB87382.1 GGFK01006563 MBW39884.1 ABLF02021645 GGFL01001916 MBW66094.1 AB090820 BAC57916.1 ABLF02030663 ABLF02041312 GALX01004171 JAB64295.1 GGFL01001189 MBW65367.1 GGFL01001190 MBW65368.1 GGFL01001583 MBW65761.1
Proteomes
PRIDE
Pfam
Interpro
IPR000477
RT_dom
+ More
IPR001878 Znf_CCHC
IPR036875 Znf_CCHC_sf
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR000663 Natr_peptide
IPR021896 Transposase_37
IPR026960 RVT-Znf
IPR016067 S-AdoMet_deCO2ase_core
IPR012337 RNaseH-like_sf
IPR002156 RNaseH_domain
IPR036397 RNaseH_sf
IPR001878 Znf_CCHC
IPR036875 Znf_CCHC_sf
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR000663 Natr_peptide
IPR021896 Transposase_37
IPR026960 RVT-Znf
IPR016067 S-AdoMet_deCO2ase_core
IPR012337 RNaseH-like_sf
IPR002156 RNaseH_domain
IPR036397 RNaseH_sf
Gene 3D
ProteinModelPortal
A0A139WAH4
F7IYV5
A0A139W940
F7IYU8
J9KJG8
A0A139W9B3
+ More
D6WC19 A0A139W939 F7IYV2 V5I881 A0A2H8TR36 A0A139WNC9 A0A139WN93 A0A139W9J0 A0A224XIZ2 T1DG59 T1DCM0 T1DG89 F7IYV6 A0A142LX45 J9KUM6 A0A2M4CS98 A0A2M4ADW9 A0A2M4BBV9 J9JWJ3 A0A2S2NQQ6 J9LVU7 A0A2M4BC01 A0A2M4BC28 A0A034WR48 A0A2M4BC24 A0A139W9A6 A0A2M4BC18 A0A2H8TIE5 A0A142LX43 A0A142LX37 A0A0K8VPH1 W8ANK7 X1WZ64 W8ADE4 J9KM84 F7IYV3 A0A2M4BDI3 A0A2M4CSA9 Q868S4 F7IYV7 A0A2M4CSI6 A0A1W7R6I9 A0A023EW96 A0A139W8G2 A0A2S2PRL8 A0A0K8S5I5 D7EIB0 A0A139W942 Q868S6 A0A2M4BB78 A0A2M3Z1C1 A0A2M4BB89 A0A139WA31 A0A2M4BBA0 Q9N9Z1 A0A1S4EMG7 J9L6Z9 A0A2S2PEZ7 A0A2M4BCE1 Q868S0 A0A139W8J5 X1WXT5 Q868T0 A0A2M4AK35 J9LM37 J9KT61 K7JP42 J9K6F0 J9JY94 X1WJY4 A0A2S2NJ82 J9KNS1 A0A2M4CJ33 J9KNN8 Q868S8 V9GZD3 Q868R6 A0A2S2P7D1 Q868R2 A0A2S2P488 Q868R0 A0A0J7KIY5 W8AMZ9 A0A2M4AGH2 J9LBG0 A0A2M4CL98 Q868R4 J9LQQ1 V5G2Q2 A0A2M4CJ59 A0A2M4CKD9 A0A2M4CKQ7
D6WC19 A0A139W939 F7IYV2 V5I881 A0A2H8TR36 A0A139WNC9 A0A139WN93 A0A139W9J0 A0A224XIZ2 T1DG59 T1DCM0 T1DG89 F7IYV6 A0A142LX45 J9KUM6 A0A2M4CS98 A0A2M4ADW9 A0A2M4BBV9 J9JWJ3 A0A2S2NQQ6 J9LVU7 A0A2M4BC01 A0A2M4BC28 A0A034WR48 A0A2M4BC24 A0A139W9A6 A0A2M4BC18 A0A2H8TIE5 A0A142LX43 A0A142LX37 A0A0K8VPH1 W8ANK7 X1WZ64 W8ADE4 J9KM84 F7IYV3 A0A2M4BDI3 A0A2M4CSA9 Q868S4 F7IYV7 A0A2M4CSI6 A0A1W7R6I9 A0A023EW96 A0A139W8G2 A0A2S2PRL8 A0A0K8S5I5 D7EIB0 A0A139W942 Q868S6 A0A2M4BB78 A0A2M3Z1C1 A0A2M4BB89 A0A139WA31 A0A2M4BBA0 Q9N9Z1 A0A1S4EMG7 J9L6Z9 A0A2S2PEZ7 A0A2M4BCE1 Q868S0 A0A139W8J5 X1WXT5 Q868T0 A0A2M4AK35 J9LM37 J9KT61 K7JP42 J9K6F0 J9JY94 X1WJY4 A0A2S2NJ82 J9KNS1 A0A2M4CJ33 J9KNN8 Q868S8 V9GZD3 Q868R6 A0A2S2P7D1 Q868R2 A0A2S2P488 Q868R0 A0A0J7KIY5 W8AMZ9 A0A2M4AGH2 J9LBG0 A0A2M4CL98 Q868R4 J9LQQ1 V5G2Q2 A0A2M4CJ59 A0A2M4CKD9 A0A2M4CKQ7
PDB
1WDU
E-value=1.41205e-10,
Score=163
Ontologies
KEGG
GO
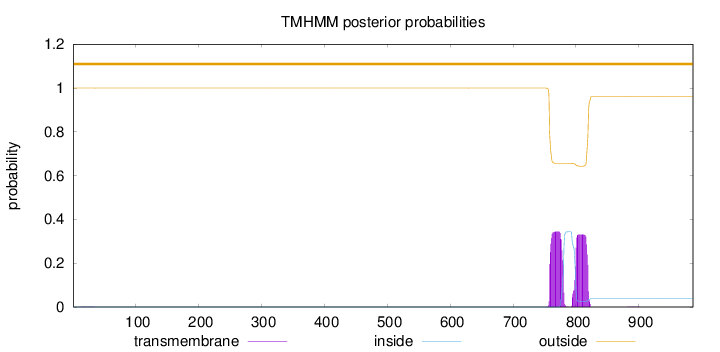
Topology
Length:
986
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
14.21218
Exp number, first 60 AAs:
0.00482
Total prob of N-in:
0.00034
outside
1 - 986
Population Genetic Test Statistics
Pi
3.474093
Theta
11.32497
Tajima's D
-1.954946
CLR
160.449003
CSRT
0.0177491125443728
Interpretation
Uncertain
