Gene
KWMTBOMO02337
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Full name
Integrin beta
+ More
Sodium/hydrogen exchanger
Sodium/hydrogen exchanger
Location in the cell
Mitochondrial Reliability : 2.092 Nuclear Reliability : 1.572
Sequence
CDS
ATGGCAACCCCCCCCAACATGGCTGACACAAATAAAAACTGGCCCCCAGTCCCCGGCAGCTCCGCTATTCCTGGCTACGGTAGCGGTCAAGGTAACGGCGGGGCAGGGGGTGCTAAGAATCTCCGGCAGAGATCCGGCTACCATAAAAAACGACGACTGGACCTGGCAACATACAACGTGCGAACACTGCGGACCGATGAGAAGATTGTGGAACTCGAGGAAGAATTGAGCAGGTTACGGTGGGACATTGTGGGCTTATCTGAAGTCCGAAGAGAGGGAGAGGACACGATAACTCTGAGGTCCGGTAACTTGCTCTATTTTCGAGAAGGTGATCATCTGTCTCAAGGCGGTGTCGGATTTTTCGTCCACAAGTCCCTTGTGAATAATGTGCTCAAAATCGAGAGTGTGTCAACCAGGGTGGCGTACCTTGTACTCAGAATCACCAAAAAGTATTCGTTGAAGATCATACAGGTTTACGCTCCGACCTCGGCACACTCCGACGATGAGGTAGAGACTATGTATGAGGACATTTCCAGAGCCATGCATAGCTTTAAAACTCACATCACCATTGTCATGGGGGATTTCAACGCCAGACTGGGTAAACGGTCCGAGGGGGAGCTGCGAGTGGGACCATTTGGATTTGGGAATCGAAATCATCGGGGAAACCTACTTGCCGGTTTCATGGAGAAAGAAGAATTATTTATGATGAACTCTTTCTTCCAAAAACGGAATCACAGGAAGTGGACCTGGATGAGCCCCAATGGTGCCATCAAAAATGAGATAGATTTCATTATGTCGACAAAAAGGCAAATATTCAATGATGTCTCTGTGATCAATATGGTTAAAACAGGAAGTGATCATCGGATGGTAAGAGGCACATTGAACCTCAATATTAAGCTTGAGAGATTCCGACTGATAAGGTCTACGCTCCGTCCAGCACCAGCTCAGATCCAAAGTTCTGGGCACTTTCAGCTCGAACTTCAAAACCGCTTCGATTGCCTAGGAGGCTCCGTGGACGATTACAATGACGGGTTTGTGAATGCTGTCCATACGATTGGTTCCAAGTTTTTTGGAGCCCGTCGTACAAACAGAACCAAAAAACTCTCAGACCCTACTCTCAAACTCATGGAAGAACGTCGCGGAATGATTTTGCAGTCTTCAGAAGATTCGTTTAAATATCGGTCTCTCAACAGACAGATCTCAAAGTCTCTTAAATTTGATCTACGCCGTTTTAATACTGAGCGTATTAAAGAGGCTATAGAACGAAACAAAGGCTCCAAGGTGTTCGCGAGAGATTTGTCTCTTGGGAAAAGCCAGTTGACAAGACTGAAGATCGATGATGGCAGGATCATTTCGTCCAAGCCTGAGCTTTTGAGAGAGGTCGAGAAGTTCTATGGACAGCTATACACATCTACTCGACAACCCGTGGAATATCTAGCAAAGGATCCAAGAGCTAGATTAGTCCGACACTATACCGAAGATATCCCCGACATCAGCCTGTGCGAGATTAGAATGGCTCTCAAGCAACTTAAGGACAACAAGGCACCGGGCGAGGATGGAATTACCGTCGAACTTCTGAAGGCGGGTGGAAAACCAGTCCTTCGGGCCCTACAAAATTTATTTAATTCCGTCATTCTCGATGGGAATGTGCCCCTGGCATGGACCAGAAGTGTGGTGGTACTGTTCTTCAAAAAGGGTGACAATACTCGTTTGAAGAATTATAGACCCATCTCACTTCTGAGTCATGTCTATAAGTTGTTCTCGAGAGTTATTACGAACCGTCTCGCGCGCAGGTTTGATGACTTCCAGCCACCCGAACAAGCCGGTTTCCGAAAAGGCTATAGTACCGTAGACCACATTCATACGTTGCGGCAGATTGTACAAAAGACCGAGGAATATAACCTGCCGCTTTGCTTAGCATTTGTGGACTACGAGAAAGCCTTTGATTCGATTGAAACCTGGGCCGTGTTACAGTCTCTCCAACGGTGCCAAATTGATTATCGGTATATCGAAGTGTTGAGGTGTTTGTATGAAAACGCTACTATGTCAGTCCGAGTACAGGACCGGGCTTCAGAACCTATCCTTCTGCAGCGGGGCGTGAGGCAAGGAGATGTTATTTCTCCGAAACTATTTACCGCTGCGCTGGAAGATGTTTTCAAAGTTCTGGACTGGAAAGGACTAGGCATCAACATAAACGGCGAGTACATAACTCACCTTCGGTTCGCCGATGATATTGTGATTATGGCCGAGACCATGGAGGACCTCAGTACAATGCTCAAAGACCTCAGTAGGGCTTCTATCCGAGTGGGTCTAAATATGAACAAAGAAAAAACAAAAATTATGTTGAATGCTCATGTAGCGCCCACTCCAGTAAAGATCGGAGGCTCTACGCTCGAAGTTGTTGACGAGTATATTTACCTGGGACACACTGTCCAGCTAGGTAAGTCCAACTTCGAAAAAGAGGTCAACCGTCGAATCCAGCTCGGATGGGCAGCGTTCGGAAAGCTACGTAAAATCTTCTCATCACAAATACCACAGTGTCTCAAGACGAAAGTCTTTGACCAGTGTGTGTTGCCAGTGATGACATATGGCACTGAGACGTGGTCGCTCACAATGGGCCTTATGAGAAGGCTCAAGGTCACCCAAAGAGCAATGGAGAGGGCTATGCTCGGAGTTTCCCTGCGAGACCAAATCAGAAATGAGGAGATTCGCAGGAAAACCATGGTAACCGACATAGCCCAAAGGATTGCGACACTGAAGTGGCAGTGGGCAGGACACATTGCTCGCGGAACAGATGGCCGTTGGGGCCAGAAGGTTCTTGAGTGGCGACCGCGTACTGGGAGACGTAGCGTGGGTAGGCCTCCCACAAGGTGGGCCGACGACCTATTAAGGTTCGCGGGGATCCGCTGGATGCAGGCAGCGCAGGACCGGTCTTTGTGGCAAGGCTTGGGGGAGGCCTATGTCCAGCAGTGGACGTCCATGGGCTGA
Protein
MATPPNMADTNKNWPPVPGSSAIPGYGSGQGNGGAGGAKNLRQRSGYHKKRRLDLATYNVRTLRTDEKIVELEEELSRLRWDIVGLSEVRREGEDTITLRSGNLLYFREGDHLSQGGVGFFVHKSLVNNVLKIESVSTRVAYLVLRITKKYSLKIIQVYAPTSAHSDDEVETMYEDISRAMHSFKTHITIVMGDFNARLGKRSEGELRVGPFGFGNRNHRGNLLAGFMEKEELFMMNSFFQKRNHRKWTWMSPNGAIKNEIDFIMSTKRQIFNDVSVINMVKTGSDHRMVRGTLNLNIKLERFRLIRSTLRPAPAQIQSSGHFQLELQNRFDCLGGSVDDYNDGFVNAVHTIGSKFFGARRTNRTKKLSDPTLKLMEERRGMILQSSEDSFKYRSLNRQISKSLKFDLRRFNTERIKEAIERNKGSKVFARDLSLGKSQLTRLKIDDGRIISSKPELLREVEKFYGQLYTSTRQPVEYLAKDPRARLVRHYTEDIPDISLCEIRMALKQLKDNKAPGEDGITVELLKAGGKPVLRALQNLFNSVILDGNVPLAWTRSVVVLFFKKGDNTRLKNYRPISLLSHVYKLFSRVITNRLARRFDDFQPPEQAGFRKGYSTVDHIHTLRQIVQKTEEYNLPLCLAFVDYEKAFDSIETWAVLQSLQRCQIDYRYIEVLRCLYENATMSVRVQDRASEPILLQRGVRQGDVISPKLFTAALEDVFKVLDWKGLGININGEYITHLRFADDIVIMAETMEDLSTMLKDLSRASIRVGLNMNKEKTKIMLNAHVAPTPVKIGGSTLEVVDEYIYLGHTVQLGKSNFEKEVNRRIQLGWAAFGKLRKIFSSQIPQCLKTKVFDQCVLPVMTYGTETWSLTMGLMRRLKVTQRAMERAMLGVSLRDQIRNEEIRRKTMVTDIAQRIATLKWQWAGHIARGTDGRWGQKVLEWRPRTGRRSVGRPPTRWADDLLRFAGIRWMQAAQDRSLWQGLGEAYVQQWTSMG
Summary
Similarity
Belongs to the integrin beta chain family.
Belongs to the monovalent cation:proton antiporter 1 (CPA1) transporter (TC 2.A.36) family.
Belongs to the monovalent cation:proton antiporter 1 (CPA1) transporter (TC 2.A.36) family.
Feature
chain Integrin beta
Uniprot
D7F164
D7F158
A0A437BHD1
D7F159
D7F157
D7F165
+ More
D7F160 D7F166 A0A437B0R4 A0A2W1BZV1 A0A2H1VS24 A0A023EXV2 A0A0K0FAW8 D7F179 D7F176 A0A016SG46 A0A016TK11 A0A016W4R1 D7F177 A0A023EWU8 A0A2G8JFM9 D7F178 A0A016S1D2 A0A0P4VWD0 A0A0N5C0B1 A0A016V419 E3MH09 E3NC15 A0A016ULU9 A0A016WPY4 A0A016T7J7 A0A016WCB8 K7HDM3 K7HDM2 K7ICT2 A0A069DZM5 K7I2A3 A0A2H1WF27 K7ICT1 A0A2H2JFN0 K7IG35 K7IBZ5 K7IBZ6 K7H0Q9 K7H0R0 A0A2H2J8I4 A0A016RUR3 A0A016S795 A0A016WA79 A0A016S5B2 A0A016RV73 A0A016VVN3 A0A016RRX8 K7I2A2 K7IG36 A0A016SLV8 K7I2A1 E3N0F4 A0A016V6A2 Q9TZX4 K7H0Q8 E3NGJ2 A0A016U3L3 A0A016U352 E3NJQ7 A0A016SVA2 A0A016SW37 A0A016T9D3 A0A016WE57 A0A016T9A5 A0A069DX96 A0A016THC6 A0A016V4Q8 A0A2W5U8K4 A0A016SX67 A0A016SPR9 A0A016UN79 A0A2H2ITH9 A0A2H2JCS8 A0A2H2JKG1 A0A016T4C4 A0A2H2JKX8 A0A016TQJ1 A0A016VT25 A0A016VGA0 A0A016WYN9 A0A016W1X2 A0A016VLP0 A0A016SY13 A0A016U6B7 A0A016U123
D7F160 D7F166 A0A437B0R4 A0A2W1BZV1 A0A2H1VS24 A0A023EXV2 A0A0K0FAW8 D7F179 D7F176 A0A016SG46 A0A016TK11 A0A016W4R1 D7F177 A0A023EWU8 A0A2G8JFM9 D7F178 A0A016S1D2 A0A0P4VWD0 A0A0N5C0B1 A0A016V419 E3MH09 E3NC15 A0A016ULU9 A0A016WPY4 A0A016T7J7 A0A016WCB8 K7HDM3 K7HDM2 K7ICT2 A0A069DZM5 K7I2A3 A0A2H1WF27 K7ICT1 A0A2H2JFN0 K7IG35 K7IBZ5 K7IBZ6 K7H0Q9 K7H0R0 A0A2H2J8I4 A0A016RUR3 A0A016S795 A0A016WA79 A0A016S5B2 A0A016RV73 A0A016VVN3 A0A016RRX8 K7I2A2 K7IG36 A0A016SLV8 K7I2A1 E3N0F4 A0A016V6A2 Q9TZX4 K7H0Q8 E3NGJ2 A0A016U3L3 A0A016U352 E3NJQ7 A0A016SVA2 A0A016SW37 A0A016T9D3 A0A016WE57 A0A016T9A5 A0A069DX96 A0A016THC6 A0A016V4Q8 A0A2W5U8K4 A0A016SX67 A0A016SPR9 A0A016UN79 A0A2H2ITH9 A0A2H2JCS8 A0A2H2JKG1 A0A016T4C4 A0A2H2JKX8 A0A016TQJ1 A0A016VT25 A0A016VGA0 A0A016WYN9 A0A016W1X2 A0A016VLP0 A0A016SY13 A0A016U6B7 A0A016U123
EMBL
FJ265549
ADI61817.1
FJ265543
ADI61811.1
RSAL01000057
RVE49824.1
+ More
FJ265544 ADI61812.1 FJ265542 ADI61810.1 FJ265550 ADI61818.1 FJ265545 ADI61813.1 FJ265551 ADI61819.1 RSAL01000220 RVE44071.1 KZ149907 PZC78256.1 ODYU01004088 SOQ43598.1 GBBI01004876 JAC13836.1 FJ265564 ADI61832.1 FJ265561 ADI61829.1 JARK01001565 EYB89683.1 JARK01001676 JARK01001446 JARK01001433 EYB83199.1 EYC01126.1 EYC02928.1 JARK01001337 EYC34625.1 FJ265562 ADI61830.1 GBBI01004877 JAC13835.1 MRZV01002136 PIK34564.1 FJ265563 ADI61831.1 JARK01001655 EYB84321.1 GDKW01001596 JAI54999.1 JARK01001354 EYC21986.1 DS268444 EFP01729.1 DS268591 EFO92565.1 JARK01001370 EYC16150.1 JARK01000173 EYC41342.1 JARK01001736 JARK01001462 JARK01001379 JARK01001338 EYB80775.1 EYB98943.1 EYC13646.1 EYC33347.1 JARK01000388 EYC37469.1 GBGD01000380 JAC88509.1 ODYU01008249 SOQ51673.1 JARK01001711 EYB81734.1 JARK01001621 EYB86109.1 JARK01000460 EYC36749.1 JARK01001626 EYB85833.1 JARK01001700 JARK01001393 EYB82228.1 EYC10181.1 JARK01001340 EYC30828.1 JARK01001730 EYB81048.1 JARK01001543 EYB91356.1 DS268505 EFP13372.1 JARK01001353 EYC22263.1 AF054983 AAC72298.1 DS268655 EFO97114.1 JARK01001395 EYC09730.1 EYC09729.1 DS268752 EFP01106.1 JARK01001505 EYB94618.1 EYB94617.1 JARK01001459 EYB99276.1 JARK01000343 EYC38099.1 EYB99275.1 GBGD01000453 JAC88436.1 JARK01001438 EYC02111.1 EYC21713.1 QFQQ01000179 PZR20596.1 JARK01001500 JARK01001343 EYB95040.1 EYC28825.1 JARK01001531 EYB92324.1 EYC16376.1 JARK01001475 EYB97550.1 JARK01001421 EYC04912.1 EYC30744.1 JARK01001346 EYC26445.1 JARK01000063 EYC44387.1 EYC33297.1 JARK01001344 EYC28216.1 JARK01001497 EYB95322.1 JARK01001389 EYC10874.1 JARK01001401 EYC08686.1
FJ265544 ADI61812.1 FJ265542 ADI61810.1 FJ265550 ADI61818.1 FJ265545 ADI61813.1 FJ265551 ADI61819.1 RSAL01000220 RVE44071.1 KZ149907 PZC78256.1 ODYU01004088 SOQ43598.1 GBBI01004876 JAC13836.1 FJ265564 ADI61832.1 FJ265561 ADI61829.1 JARK01001565 EYB89683.1 JARK01001676 JARK01001446 JARK01001433 EYB83199.1 EYC01126.1 EYC02928.1 JARK01001337 EYC34625.1 FJ265562 ADI61830.1 GBBI01004877 JAC13835.1 MRZV01002136 PIK34564.1 FJ265563 ADI61831.1 JARK01001655 EYB84321.1 GDKW01001596 JAI54999.1 JARK01001354 EYC21986.1 DS268444 EFP01729.1 DS268591 EFO92565.1 JARK01001370 EYC16150.1 JARK01000173 EYC41342.1 JARK01001736 JARK01001462 JARK01001379 JARK01001338 EYB80775.1 EYB98943.1 EYC13646.1 EYC33347.1 JARK01000388 EYC37469.1 GBGD01000380 JAC88509.1 ODYU01008249 SOQ51673.1 JARK01001711 EYB81734.1 JARK01001621 EYB86109.1 JARK01000460 EYC36749.1 JARK01001626 EYB85833.1 JARK01001700 JARK01001393 EYB82228.1 EYC10181.1 JARK01001340 EYC30828.1 JARK01001730 EYB81048.1 JARK01001543 EYB91356.1 DS268505 EFP13372.1 JARK01001353 EYC22263.1 AF054983 AAC72298.1 DS268655 EFO97114.1 JARK01001395 EYC09730.1 EYC09729.1 DS268752 EFP01106.1 JARK01001505 EYB94618.1 EYB94617.1 JARK01001459 EYB99276.1 JARK01000343 EYC38099.1 EYB99275.1 GBGD01000453 JAC88436.1 JARK01001438 EYC02111.1 EYC21713.1 QFQQ01000179 PZR20596.1 JARK01001500 JARK01001343 EYB95040.1 EYC28825.1 JARK01001531 EYB92324.1 EYC16376.1 JARK01001475 EYB97550.1 JARK01001421 EYC04912.1 EYC30744.1 JARK01001346 EYC26445.1 JARK01000063 EYC44387.1 EYC33297.1 JARK01001344 EYC28216.1 JARK01001497 EYB95322.1 JARK01001389 EYC10874.1 JARK01001401 EYC08686.1
Proteomes
Pfam
Interpro
IPR000477
RT_dom
+ More
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR032695 Integrin_dom_sf
IPR036465 vWFA_dom_sf
IPR015812 Integrin_bsu
IPR002035 VWF_A
IPR002369 Integrin_bsu_VWA
IPR020846 MFS_dom
IPR002293 AA/rel_permease1
IPR002213 UDP_glucos_trans
IPR017441 Protein_kinase_ATP_BS
IPR008271 Ser/Thr_kinase_AS
IPR011009 Kinase-like_dom_sf
IPR000719 Prot_kinase_dom
IPR001507 ZP_dom
IPR019430 7TM_GPCR_serpentine_rcpt_Srx
IPR017452 GPCR_Rhodpsn_7TM
IPR018422 Cation/H_exchanger_CPA1
IPR006153 Cation/H_exchanger
IPR004709 NaH_exchanger
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR032695 Integrin_dom_sf
IPR036465 vWFA_dom_sf
IPR015812 Integrin_bsu
IPR002035 VWF_A
IPR002369 Integrin_bsu_VWA
IPR020846 MFS_dom
IPR002293 AA/rel_permease1
IPR002213 UDP_glucos_trans
IPR017441 Protein_kinase_ATP_BS
IPR008271 Ser/Thr_kinase_AS
IPR011009 Kinase-like_dom_sf
IPR000719 Prot_kinase_dom
IPR001507 ZP_dom
IPR019430 7TM_GPCR_serpentine_rcpt_Srx
IPR017452 GPCR_Rhodpsn_7TM
IPR018422 Cation/H_exchanger_CPA1
IPR006153 Cation/H_exchanger
IPR004709 NaH_exchanger
Gene 3D
CDD
ProteinModelPortal
D7F164
D7F158
A0A437BHD1
D7F159
D7F157
D7F165
+ More
D7F160 D7F166 A0A437B0R4 A0A2W1BZV1 A0A2H1VS24 A0A023EXV2 A0A0K0FAW8 D7F179 D7F176 A0A016SG46 A0A016TK11 A0A016W4R1 D7F177 A0A023EWU8 A0A2G8JFM9 D7F178 A0A016S1D2 A0A0P4VWD0 A0A0N5C0B1 A0A016V419 E3MH09 E3NC15 A0A016ULU9 A0A016WPY4 A0A016T7J7 A0A016WCB8 K7HDM3 K7HDM2 K7ICT2 A0A069DZM5 K7I2A3 A0A2H1WF27 K7ICT1 A0A2H2JFN0 K7IG35 K7IBZ5 K7IBZ6 K7H0Q9 K7H0R0 A0A2H2J8I4 A0A016RUR3 A0A016S795 A0A016WA79 A0A016S5B2 A0A016RV73 A0A016VVN3 A0A016RRX8 K7I2A2 K7IG36 A0A016SLV8 K7I2A1 E3N0F4 A0A016V6A2 Q9TZX4 K7H0Q8 E3NGJ2 A0A016U3L3 A0A016U352 E3NJQ7 A0A016SVA2 A0A016SW37 A0A016T9D3 A0A016WE57 A0A016T9A5 A0A069DX96 A0A016THC6 A0A016V4Q8 A0A2W5U8K4 A0A016SX67 A0A016SPR9 A0A016UN79 A0A2H2ITH9 A0A2H2JCS8 A0A2H2JKG1 A0A016T4C4 A0A2H2JKX8 A0A016TQJ1 A0A016VT25 A0A016VGA0 A0A016WYN9 A0A016W1X2 A0A016VLP0 A0A016SY13 A0A016U6B7 A0A016U123
D7F160 D7F166 A0A437B0R4 A0A2W1BZV1 A0A2H1VS24 A0A023EXV2 A0A0K0FAW8 D7F179 D7F176 A0A016SG46 A0A016TK11 A0A016W4R1 D7F177 A0A023EWU8 A0A2G8JFM9 D7F178 A0A016S1D2 A0A0P4VWD0 A0A0N5C0B1 A0A016V419 E3MH09 E3NC15 A0A016ULU9 A0A016WPY4 A0A016T7J7 A0A016WCB8 K7HDM3 K7HDM2 K7ICT2 A0A069DZM5 K7I2A3 A0A2H1WF27 K7ICT1 A0A2H2JFN0 K7IG35 K7IBZ5 K7IBZ6 K7H0Q9 K7H0R0 A0A2H2J8I4 A0A016RUR3 A0A016S795 A0A016WA79 A0A016S5B2 A0A016RV73 A0A016VVN3 A0A016RRX8 K7I2A2 K7IG36 A0A016SLV8 K7I2A1 E3N0F4 A0A016V6A2 Q9TZX4 K7H0Q8 E3NGJ2 A0A016U3L3 A0A016U352 E3NJQ7 A0A016SVA2 A0A016SW37 A0A016T9D3 A0A016WE57 A0A016T9A5 A0A069DX96 A0A016THC6 A0A016V4Q8 A0A2W5U8K4 A0A016SX67 A0A016SPR9 A0A016UN79 A0A2H2ITH9 A0A2H2JCS8 A0A2H2JKG1 A0A016T4C4 A0A2H2JKX8 A0A016TQJ1 A0A016VT25 A0A016VGA0 A0A016WYN9 A0A016W1X2 A0A016VLP0 A0A016SY13 A0A016U6B7 A0A016U123
PDB
6AR3
E-value=2.78436e-06,
Score=126
Ontologies
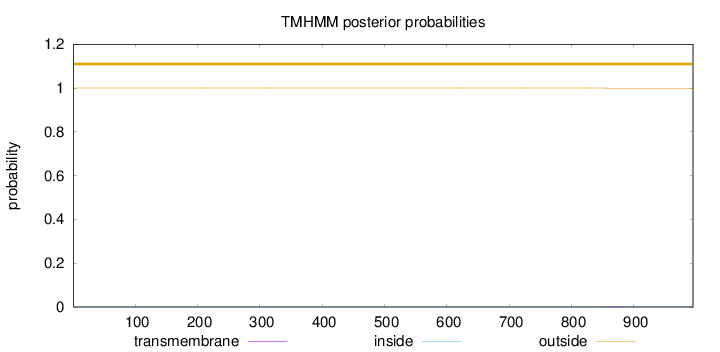
Topology
Subcellular location
Membrane
Length:
995
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.02371
Exp number, first 60 AAs:
0.00082
Total prob of N-in:
0.00008
outside
1 - 995
Population Genetic Test Statistics
Pi
14.344572
Theta
18.816744
Tajima's D
-0.967766
CLR
40.119689
CSRT
0.138243087845608
Interpretation
Uncertain
