Gene
KWMTBOMO01894
Pre Gene Modal
BGIBMGA005944
Annotation
PREDICTED:_serine/threonine-protein_phosphatase_1_regulatory_subunit_10-like_isoform_X2_[Papilio_machaon]
Location in the cell
Nuclear Reliability : 3.963
Sequence
CDS
ATGTCCAGCATAGAACCAAAACAACTTCTTGGGTGCCTTAGTGGACTTCTTTCTAATAATGGTGGCATTAAACGCAGAAACGAAGTCCAAAGGATTGCACGTCTTATGTCTAAATATTCTAAGAAATTAGTATCTAAATGTATCTACATACAAATTCTGAAGTGTACAGAAACTGAACTATTGGATTTGTTCATGAGGTATAATGGATGGCTTTTAATTCACTTATGGTTAACAGAAAGCATAGTAGCAAAAAACTGGCCTCTTGTGAAAGAATTACTGGAGTTATTGTTGTTGTGTCCTGTCGACTTAAATCGACTTAAAACTAATAATTGTCCAAAACTCGTTAAAGAATTATCTAAGGATGGAGATCACTTTGCTATTAGAGCACTGGCTTCAAAACTGGTTGAGCAATGGTTAAAAACTGTACAAGGAGAAAAAGTAGTTCCTGTTTTACTGTCTGATATTCCTCAGATAATTAGTGATGGCAACGGTGAAATAAAACCGGAGATAGTTAAACTAAAATCTGAGGTAGAACAAGCAAAAGATACTGCAGAAAATGAAGAGAGTACTGTTAAAACTGTTAAAACTGAGGAATGCTGCAAGAGTGAAAGTATAGAAATAACGCCCACAGTTAAGGAAGAAAACCAAGAACCTTCCAAAACTGAAATCAAAATAGAAACAGACCAACAACTTGAGACTTTGCCTGTCCTTAAAATTAGTTTAAAAGATGGTGTTCAAACAGTGTTGAAAGTTGAAGATACTGAAAACAAAGATTTGAAAGAATCCCCAACCAGTGAAAAGTTGAAAGAAAACTCCAAAAATAAAGATAAAGATAAATCGAGTGAGAAAACCAAAAGTAGTCACACGAAATCAAATAGTTCGAAACATTCGAGTAAAGCACGTTCTTCGAGTGATAAACATAAGAGTAGCAGCAGCTCTAGTCGACACAGTAGTAGCAAGGACAAGAAAAAAGATAAGGATAAGGATAGAAGTTCTCATAGTTCTAAATCTAAAAGTGATAGTAGTAAGAAATCATCCAGTAGTAGTAGTAGTAGTAATAGTAGTAGTAGTAGTAAAACAAAAGATGATCGTAGCACTAAAAATTCTGAAAAAGAGAAAAGAGAGAAAGATGAGAAACTTGGTAAGAAAAAAGATAAAGACACAAAAGAAAAATCGGATGAAAGTAATGAACATAAGACACCTTCAATATCGAAGCTAGGAAAAATTCCAAAGCTTAGTGATTTAAAAAAGGAAAAGCCTTCTATATCAATAGAAGTGAGAAAGCCTGACGATCCAAAACCGAAAACTGTAAAAGCTTACAATTCAAAATTTAGGAAACATGGATTGGAGGAGGAGATAAAACCGCCGCCATCGCGCTCTGTACTATTAAATAAAAAAGCACCACCACCACCCATACCTCCCATAGTCTCTATACCGAAAAGACATTCTCCAGTACATAATGAAACACCACCTGAAAAGAAACTTAAAACTATGGAAATAGTAGAAAAACCTGGTTCTATTAAATTAATACCTCCCAAACCAAAACCCATGACACTTTTGGAAAGCGATATGTTTATGGATGCACTCAATGCTTCAGCGACAAATAAGAAGGAACCAAAGAAAAGAAAAAGGCGTACGAGCGGATCGAAGGATGGCAATCCTAATGACGGATCACCTCCACCTACACCGACATCACTCAGTAGTCCCACAAGCGAAAGTAAATCTGTACCGCCAAGGTTTTACCAAGATACTTTGGATGCAGAAGAACGTAAAGACAAAAGTCATGAAAGTAAAGAACCAGTTAGTCCCGTGACCACAGAAAATGAAGTCGAAGACGACAAGATGGACGTTTCTGGGCCGCATACATTAACAGTTAATGGATTAAAAGGGGTTTTATGTTATCATAGAAAAAAAGGTCCCAAAAAGAACATTAAATGGAAACCTGACAGTGAACTAGAAACAATTCATGTATTTGACCTGGATGAGACTGAAAGAATAAATGTCACCAAAACATTCACGGATATGAAATATTTAGAAAGAATACACGAAAGGGAAGCTTTCGAAAAAGCAAGAAACCTTAGTAATGACGACGTTATGGAAGAACGTACAAATTGGAAGCCTTTGATTTTAGTAGATTTAGAAGGACAATTACAAGTAGATTATGGCAAAAACAGTAAAGAGAAAGATATACAGGCACTACGACAAAAAGTAACCTTACAACCTTTATATTTCAACAAGGCAATGATTCCTGATTCCCCAATTGAACCAGAATTAGAAACTCACACCTATTCTGAACCCTCAATAATTCCCTTGGAAGATGTCACTGGAAATCAAGATAATATAAGTGATTTTAGAAATATGCCTTGGCCTGAACCGAGAGGTAATGCTCCACCCCAAACACCGAGTAATATGACTGTTGGTGCTAGTCCATTATTCCCTAACAATCTCACACAATTTTCAAACACTTTTCCTAATACGCAATTTCCTGGAGTACCACCTGCCTTCCAAGGCCCTAACATCGTTCCTGGTGAGTGGCAAAATGGAATACCCCAACAGATGATACCAAATGGAATGCCTGGTCCAATAGGAACCCACAATTTAGCACCAGGTGTGATGCCTCCCGGAAGCCATCCACAAGGTATGATGATTTCACCTGAGACTATGATGATGGCTCCGGAGATGTTCGGAAACCCAACTCCAATGATTCCTGTACCACCGGAAAGTTTCAATATGCAACAGAATATGTATCCGATGGATTTCAACATGGTAGGTCCTCCGGGACCGGGCCCCGAAGGTTTTAATGGCCCTGGCCCTGGTTCTGGTCCCGGACCTGGTAATTTTAGAGGAGCCATGCGTGGTCGCGGTGTCAATGGTCATTGGCGCGGAAAGGGTGCGGGGAACTGGGAAAATCCGTCCAGGGGTGGTAGAGGAGGAGGCCCTCGCGGGAACCGAAAGACAATTTGTATGTATTTTCAAAGGAAGGGATCTTGTCGGCAGGGCGATAATTGTACATTCTTGCATCCAGGTGTGAACTGTCCTTTTTAA
Protein
MSSIEPKQLLGCLSGLLSNNGGIKRRNEVQRIARLMSKYSKKLVSKCIYIQILKCTETELLDLFMRYNGWLLIHLWLTESIVAKNWPLVKELLELLLLCPVDLNRLKTNNCPKLVKELSKDGDHFAIRALASKLVEQWLKTVQGEKVVPVLLSDIPQIISDGNGEIKPEIVKLKSEVEQAKDTAENEESTVKTVKTEECCKSESIEITPTVKEENQEPSKTEIKIETDQQLETLPVLKISLKDGVQTVLKVEDTENKDLKESPTSEKLKENSKNKDKDKSSEKTKSSHTKSNSSKHSSKARSSSDKHKSSSSSSRHSSSKDKKKDKDKDRSSHSSKSKSDSSKKSSSSSSSSNSSSSSKTKDDRSTKNSEKEKREKDEKLGKKKDKDTKEKSDESNEHKTPSISKLGKIPKLSDLKKEKPSISIEVRKPDDPKPKTVKAYNSKFRKHGLEEEIKPPPSRSVLLNKKAPPPPIPPIVSIPKRHSPVHNETPPEKKLKTMEIVEKPGSIKLIPPKPKPMTLLESDMFMDALNASATNKKEPKKRKRRTSGSKDGNPNDGSPPPTPTSLSSPTSESKSVPPRFYQDTLDAEERKDKSHESKEPVSPVTTENEVEDDKMDVSGPHTLTVNGLKGVLCYHRKKGPKKNIKWKPDSELETIHVFDLDETERINVTKTFTDMKYLERIHEREAFEKARNLSNDDVMEERTNWKPLILVDLEGQLQVDYGKNSKEKDIQALRQKVTLQPLYFNKAMIPDSPIEPELETHTYSEPSIIPLEDVTGNQDNISDFRNMPWPEPRGNAPPQTPSNMTVGASPLFPNNLTQFSNTFPNTQFPGVPPAFQGPNIVPGEWQNGIPQQMIPNGMPGPIGTHNLAPGVMPPGSHPQGMMISPETMMMAPEMFGNPTPMIPVPPESFNMQQNMYPMDFNMVGPPGPGPEGFNGPGPGSGPGPGNFRGAMRGRGVNGHWRGKGAGNWENPSRGGRGGGPRGNRKTICMYFQRKGSCRQGDNCTFLHPGVNCPF
Summary
Uniprot
EMBL
Proteomes
Pfam
PF08711 Med26
Interpro
Gene 3D
ProteinModelPortal
Ontologies
GO
Topology
Subcellular location
Nucleus
Length:
1014
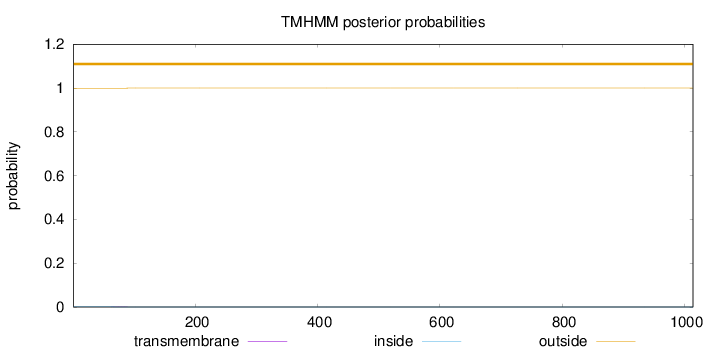
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.02123
Exp number, first 60 AAs:
0.00139
Total prob of N-in:
0.00101
outside
1 - 1014
Population Genetic Test Statistics
Pi
256.727023
Theta
204.124551
Tajima's D
0.765656
CLR
0.108598
CSRT
0.590070496475176
Interpretation
Uncertain
