Gene
KWMTBOMO00949
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 1.534 Nuclear Reliability : 1.537
Sequence
CDS
ATGTGTATTCTTCCATGCTTACTGTACGGGTGTCAAACTTGGGCGTTGACAGAAGAACTATCAAACAAATTAAAAATTTGTCAAAACGGAATAGAGGGAAGTACAATAGGCGTAAAAAGAAAAGACCGAATAAAACTAAAAGACATAAAGAGCAAGACAAACTTTAAGAACGCATACACAACTTACAAACAGCTAAAATGGCGATGGTCTGGACATATGATCAGAGAGAAAAAGGAGAAATGGACCAGTTTAATAACTGAATGGTATCCACGAGATGGTAAAAGGAATAGAGGTAGACAGACTAAAAGATGGGAGGACGACATTATAAAAATAGCTGGTACTACATGGACGCGTGCAGCAAAGGACAGGACACAGTGGAAATCTTTAGAGGAGGCCTTTGTCGAAGGACAAGCTGCGATACCACAACCACCGTTTGCCGATATTTAA
Protein
MCILPCLLYGCQTWALTEELSNKLKICQNGIEGSTIGVKRKDRIKLKDIKSKTNFKNAYTTYKQLKWRWSGHMIREKKEKWTSLITEWYPRDGKRNRGRQTKRWEDDIIKIAGTTWTRAAKDRTQWKSLEEAFVEGQAAIPQPPFADI
Summary
Uniprot
A0A2H1V663
A0A2W1BCS2
A0A2H1W4U6
A0A2W1BSQ8
A0A2W1BDX2
D7F179
+ More
A0A2H1VY30 D7F176 A4KWF4 A4KWF2 D7F178 A4KWG0 A0A131YK20 A0A2A4J025 A0A023EXV2 D7F158 D7F157 A0A069DQ34 A0A2H1WM45 D7F165 A0A224XSN7 D7F164 H9JNQ9 D7F159 D7F177 D7F160 A0A224Y0J1 A0A2H1WG43 A0A131Z9K5 W4YXJ1 E2AL50 D7F166 A0A2W1BLC9 A0A2G8JIM1 A0A023EWU8 A0A2W1BGB6 A0A2W1B5F8 A0A2A4J310 A0A069DX96 A0A0P4VWD0 A0A2W1BMB6 A0A0M3HSL8 A0A2G8JFM9 A0A2W1BZV1 A0A2P8YXE9 A0A069DZM5 A0A2G8K8G7 A0A0V0G6F7 A0A0L7LQA3 W4YTY1 A0A016VZE9 A0A016VZ67 A0A0B2VA05 A0A2H1WF27 A0A2G8K4F2 A0A2G8KFR2 A0A016W5P9 A0A2W1BI26 A0A183UMK8 A0A016SG46 A0A016TK11 A0A016W4R1 A0A1E1X2U1 A0A2W5U8K4 A0A016WNU6 A0A1E1XN17 A0A2W1B5M8 A0A2G8L598 A0A016UGZ5 A0A016UGH8 A0A2W1BSC0 W4XMW4 A0A0B2W3Y4 W4YEN5 A0A0B2UUK1 A0A183UG87 A0A3P7G895 A0A183UCP0 A0A0R3Q573 W4YU42 A0A016V419 W4YFQ5 W4YJQ5 W4XDK0 W4YRL2 A0A0K0FAW8 W4XCB7 A0A016S1D2 A0A2H2IR56 A0A183TZB3 A0A2G8L2G5 W4ZHY7 A0A016V557
A0A2H1VY30 D7F176 A4KWF4 A4KWF2 D7F178 A4KWG0 A0A131YK20 A0A2A4J025 A0A023EXV2 D7F158 D7F157 A0A069DQ34 A0A2H1WM45 D7F165 A0A224XSN7 D7F164 H9JNQ9 D7F159 D7F177 D7F160 A0A224Y0J1 A0A2H1WG43 A0A131Z9K5 W4YXJ1 E2AL50 D7F166 A0A2W1BLC9 A0A2G8JIM1 A0A023EWU8 A0A2W1BGB6 A0A2W1B5F8 A0A2A4J310 A0A069DX96 A0A0P4VWD0 A0A2W1BMB6 A0A0M3HSL8 A0A2G8JFM9 A0A2W1BZV1 A0A2P8YXE9 A0A069DZM5 A0A2G8K8G7 A0A0V0G6F7 A0A0L7LQA3 W4YTY1 A0A016VZE9 A0A016VZ67 A0A0B2VA05 A0A2H1WF27 A0A2G8K4F2 A0A2G8KFR2 A0A016W5P9 A0A2W1BI26 A0A183UMK8 A0A016SG46 A0A016TK11 A0A016W4R1 A0A1E1X2U1 A0A2W5U8K4 A0A016WNU6 A0A1E1XN17 A0A2W1B5M8 A0A2G8L598 A0A016UGZ5 A0A016UGH8 A0A2W1BSC0 W4XMW4 A0A0B2W3Y4 W4YEN5 A0A0B2UUK1 A0A183UG87 A0A3P7G895 A0A183UCP0 A0A0R3Q573 W4YU42 A0A016V419 W4YFQ5 W4YJQ5 W4XDK0 W4YRL2 A0A0K0FAW8 W4XCB7 A0A016S1D2 A0A2H2IR56 A0A183TZB3 A0A2G8L2G5 W4ZHY7 A0A016V557
Pubmed
EMBL
ODYU01000693
SOQ35892.1
KZ150485
PZC70710.1
ODYU01006342
SOQ48110.1
+ More
KZ149912 PZC78089.1 KZ150108 PZC73382.1 FJ265564 ADI61832.1 ODYU01004982 SOQ45402.1 FJ265561 ADI61829.1 EF113399 EF113400 ABO45233.1 EF113398 EF113401 ABO45231.1 FJ265563 ADI61831.1 EF113402 ABO45239.1 GEDV01010296 JAP78261.1 NWSH01004509 PCG65028.1 GBBI01004876 JAC13836.1 FJ265543 ADI61811.1 FJ265542 ADI61810.1 GBGD01002934 JAC85955.1 ODYU01009605 SOQ54153.1 FJ265550 ADI61818.1 GFTR01005437 JAW10989.1 FJ265549 ADI61817.1 BABH01035234 BABH01035235 FJ265544 ADI61812.1 FJ265562 ADI61830.1 FJ265545 ADI61813.1 GFTR01002495 JAW13931.1 ODYU01008431 SOQ52031.1 GEDV01000518 JAP88039.1 AAGJ04144474 GL440455 EFN65840.1 FJ265551 ADI61819.1 KZ149985 PZC75698.1 MRZV01001865 PIK35593.1 GBBI01004877 JAC13835.1 KZ150065 PZC74169.1 KZ150317 PZC71439.1 NWSH01003335 PCG66527.1 GBGD01000453 JAC88436.1 GDKW01001596 JAI54999.1 KZ150072 PZC74016.1 MRZV01002136 PIK34564.1 KZ149907 PZC78256.1 PYGN01000302 PSN48926.1 GBGD01000380 JAC88509.1 MRZV01000786 PIK44294.1 GECL01002465 JAP03659.1 JTDY01000331 KOB77663.1 AAGJ04012759 JARK01001338 EYC32701.1 EYC32700.1 JPKZ01002136 KHN78309.1 ODYU01008249 SOQ51673.1 MRZV01000899 PIK42833.1 MRZV01000618 PIK46838.1 JARK01000732 EYC35149.1 KZ150147 PZC72867.1 UYWY01020264 VDM41049.1 JARK01001565 EYB89683.1 JARK01001676 JARK01001446 JARK01001433 EYB83199.1 EYC01126.1 EYC02928.1 JARK01001337 EYC34625.1 GFAC01005640 JAT93548.1 QFQQ01000179 PZR20596.1 JARK01000189 EYC40932.1 GFAA01002978 JAU00457.1 KZ150308 PZC71479.1 MRZV01000215 PIK55436.1 JARK01001376 EYC14440.1 EYC14439.1 KZ149953 PZC76544.1 AAGJ04013248 JPKZ01000255 KHN88272.1 AAGJ04035986 JPKZ01003153 KHN73088.1 UYWY01019694 VDM38828.1 UYWY01019463 VDM37564.1 UZAG01000503 VDO08661.1 AAGJ04112537 JARK01001354 EYC21986.1 AAGJ04006812 AAGJ04005502 AAGJ04011852 AAGJ04112759 AAGJ04065374 JARK01001655 EYB84321.1 UYWY01001253 VDM26453.1 MRZV01000248 PIK54443.1 AAGJ04015353 JARK01001352 EYC22819.1
KZ149912 PZC78089.1 KZ150108 PZC73382.1 FJ265564 ADI61832.1 ODYU01004982 SOQ45402.1 FJ265561 ADI61829.1 EF113399 EF113400 ABO45233.1 EF113398 EF113401 ABO45231.1 FJ265563 ADI61831.1 EF113402 ABO45239.1 GEDV01010296 JAP78261.1 NWSH01004509 PCG65028.1 GBBI01004876 JAC13836.1 FJ265543 ADI61811.1 FJ265542 ADI61810.1 GBGD01002934 JAC85955.1 ODYU01009605 SOQ54153.1 FJ265550 ADI61818.1 GFTR01005437 JAW10989.1 FJ265549 ADI61817.1 BABH01035234 BABH01035235 FJ265544 ADI61812.1 FJ265562 ADI61830.1 FJ265545 ADI61813.1 GFTR01002495 JAW13931.1 ODYU01008431 SOQ52031.1 GEDV01000518 JAP88039.1 AAGJ04144474 GL440455 EFN65840.1 FJ265551 ADI61819.1 KZ149985 PZC75698.1 MRZV01001865 PIK35593.1 GBBI01004877 JAC13835.1 KZ150065 PZC74169.1 KZ150317 PZC71439.1 NWSH01003335 PCG66527.1 GBGD01000453 JAC88436.1 GDKW01001596 JAI54999.1 KZ150072 PZC74016.1 MRZV01002136 PIK34564.1 KZ149907 PZC78256.1 PYGN01000302 PSN48926.1 GBGD01000380 JAC88509.1 MRZV01000786 PIK44294.1 GECL01002465 JAP03659.1 JTDY01000331 KOB77663.1 AAGJ04012759 JARK01001338 EYC32701.1 EYC32700.1 JPKZ01002136 KHN78309.1 ODYU01008249 SOQ51673.1 MRZV01000899 PIK42833.1 MRZV01000618 PIK46838.1 JARK01000732 EYC35149.1 KZ150147 PZC72867.1 UYWY01020264 VDM41049.1 JARK01001565 EYB89683.1 JARK01001676 JARK01001446 JARK01001433 EYB83199.1 EYC01126.1 EYC02928.1 JARK01001337 EYC34625.1 GFAC01005640 JAT93548.1 QFQQ01000179 PZR20596.1 JARK01000189 EYC40932.1 GFAA01002978 JAU00457.1 KZ150308 PZC71479.1 MRZV01000215 PIK55436.1 JARK01001376 EYC14440.1 EYC14439.1 KZ149953 PZC76544.1 AAGJ04013248 JPKZ01000255 KHN88272.1 AAGJ04035986 JPKZ01003153 KHN73088.1 UYWY01019694 VDM38828.1 UYWY01019463 VDM37564.1 UZAG01000503 VDO08661.1 AAGJ04112537 JARK01001354 EYC21986.1 AAGJ04006812 AAGJ04005502 AAGJ04011852 AAGJ04112759 AAGJ04065374 JARK01001655 EYB84321.1 UYWY01001253 VDM26453.1 MRZV01000248 PIK54443.1 AAGJ04015353 JARK01001352 EYC22819.1
Proteomes
Interpro
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
A0A2H1V663
A0A2W1BCS2
A0A2H1W4U6
A0A2W1BSQ8
A0A2W1BDX2
D7F179
+ More
A0A2H1VY30 D7F176 A4KWF4 A4KWF2 D7F178 A4KWG0 A0A131YK20 A0A2A4J025 A0A023EXV2 D7F158 D7F157 A0A069DQ34 A0A2H1WM45 D7F165 A0A224XSN7 D7F164 H9JNQ9 D7F159 D7F177 D7F160 A0A224Y0J1 A0A2H1WG43 A0A131Z9K5 W4YXJ1 E2AL50 D7F166 A0A2W1BLC9 A0A2G8JIM1 A0A023EWU8 A0A2W1BGB6 A0A2W1B5F8 A0A2A4J310 A0A069DX96 A0A0P4VWD0 A0A2W1BMB6 A0A0M3HSL8 A0A2G8JFM9 A0A2W1BZV1 A0A2P8YXE9 A0A069DZM5 A0A2G8K8G7 A0A0V0G6F7 A0A0L7LQA3 W4YTY1 A0A016VZE9 A0A016VZ67 A0A0B2VA05 A0A2H1WF27 A0A2G8K4F2 A0A2G8KFR2 A0A016W5P9 A0A2W1BI26 A0A183UMK8 A0A016SG46 A0A016TK11 A0A016W4R1 A0A1E1X2U1 A0A2W5U8K4 A0A016WNU6 A0A1E1XN17 A0A2W1B5M8 A0A2G8L598 A0A016UGZ5 A0A016UGH8 A0A2W1BSC0 W4XMW4 A0A0B2W3Y4 W4YEN5 A0A0B2UUK1 A0A183UG87 A0A3P7G895 A0A183UCP0 A0A0R3Q573 W4YU42 A0A016V419 W4YFQ5 W4YJQ5 W4XDK0 W4YRL2 A0A0K0FAW8 W4XCB7 A0A016S1D2 A0A2H2IR56 A0A183TZB3 A0A2G8L2G5 W4ZHY7 A0A016V557
A0A2H1VY30 D7F176 A4KWF4 A4KWF2 D7F178 A4KWG0 A0A131YK20 A0A2A4J025 A0A023EXV2 D7F158 D7F157 A0A069DQ34 A0A2H1WM45 D7F165 A0A224XSN7 D7F164 H9JNQ9 D7F159 D7F177 D7F160 A0A224Y0J1 A0A2H1WG43 A0A131Z9K5 W4YXJ1 E2AL50 D7F166 A0A2W1BLC9 A0A2G8JIM1 A0A023EWU8 A0A2W1BGB6 A0A2W1B5F8 A0A2A4J310 A0A069DX96 A0A0P4VWD0 A0A2W1BMB6 A0A0M3HSL8 A0A2G8JFM9 A0A2W1BZV1 A0A2P8YXE9 A0A069DZM5 A0A2G8K8G7 A0A0V0G6F7 A0A0L7LQA3 W4YTY1 A0A016VZE9 A0A016VZ67 A0A0B2VA05 A0A2H1WF27 A0A2G8K4F2 A0A2G8KFR2 A0A016W5P9 A0A2W1BI26 A0A183UMK8 A0A016SG46 A0A016TK11 A0A016W4R1 A0A1E1X2U1 A0A2W5U8K4 A0A016WNU6 A0A1E1XN17 A0A2W1B5M8 A0A2G8L598 A0A016UGZ5 A0A016UGH8 A0A2W1BSC0 W4XMW4 A0A0B2W3Y4 W4YEN5 A0A0B2UUK1 A0A183UG87 A0A3P7G895 A0A183UCP0 A0A0R3Q573 W4YU42 A0A016V419 W4YFQ5 W4YJQ5 W4XDK0 W4YRL2 A0A0K0FAW8 W4XCB7 A0A016S1D2 A0A2H2IR56 A0A183TZB3 A0A2G8L2G5 W4ZHY7 A0A016V557
Ontologies
Topology
SignalP
Position: 1 - 15,
Likelihood: 0.597849
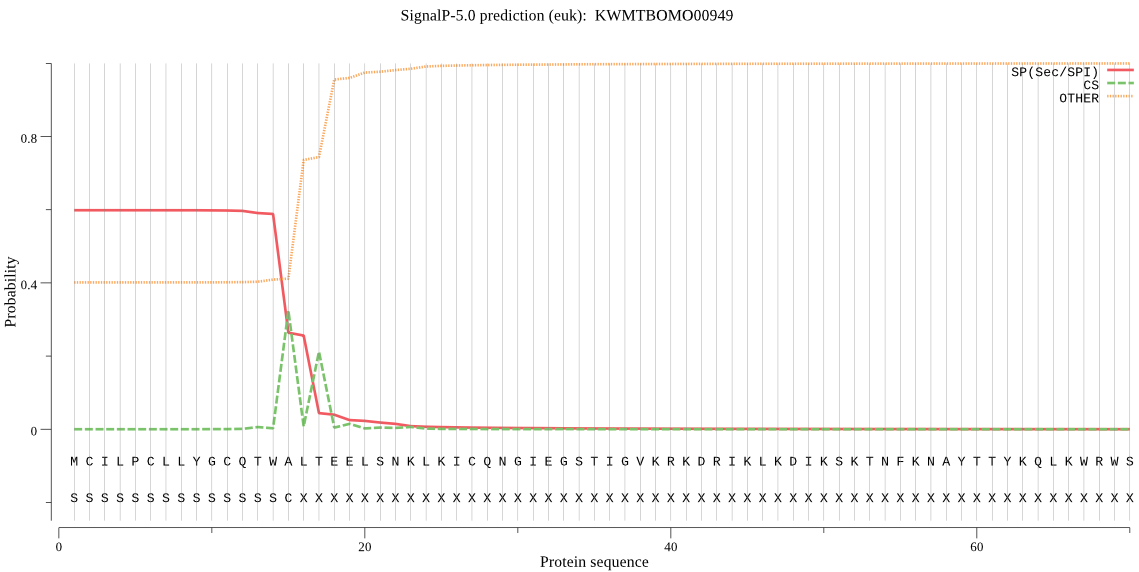
Length:
148
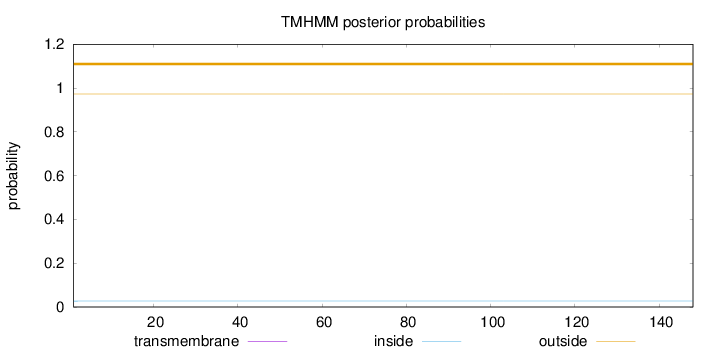
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00016
Exp number, first 60 AAs:
0.00016
Total prob of N-in:
0.02707
outside
1 - 148
Population Genetic Test Statistics
Pi
187.235554
Theta
78.634146
Tajima's D
4.321055
CLR
0
CSRT
0.999750012499375
Interpretation
Uncertain
