Gene
KWMTBOMO00933
Annotation
reverse_transcriptase_[Danaus_plexippus]
Location in the cell
Nuclear Reliability : 1.328
Sequence
CDS
ATGATGGCAGACTTAACGAAAAAATTACCCCAGGAGGGTACCACTCTAGGTGGAGAATCCCTCCGTGCTTCCACCTCGGTGGGAGCATGCTGCCCCACGTATTCTGGGGGGGGCAAAGCATGCCACGATGAAGGGGGCTGTGCGACGGGCAGCGGAAAACCCGTCGAACTGAGGGAGACCCGCTCGGGGAAGGGCGCACCGTCACCGGCGCCGTCATCGGCGTCGATGTCTGTCGCCTCGCTGGACGGGCGGACGACAGAAGAGGAGGAGGGCGGTCTTCGGGACCCTGCCTTGGTCATAGGCAGGGCCATGAAGACCGTCAAGTCGTATGCGGCGACCAGTGCCAAAACCAAAGCTGCTGGCCGCAGGCTGCGGGAAGTCGTCTCGCGAGTCTCGAAGGTTCTGCCTTCGGTTGCTTCCGCCAATGACCAGGTGGTGCAGCTGATGGCGGAGAATTTGAGGCTCCGAGCACAGCTACAGGCGCAGCGGCGGCCCGATGACAAGGTTGCTGCTACCAAGCGGGTAGCCGCGACGAGGTCGTCGCCCGCGTTGCCACGTGCCGCTGGGTCGCGGACTCCTCCGGCATCTGTGCCGGCGGTGTCGCCGCTGGTCTCCGGGGTCCCTCGGGGTCCCTGTCCCTCAGCGGTGGCTGGCGACCCGTGGCCCGTGCTGCCACCATCGCGCGCGGTACAGCGGCGTTCGCGTCTCCCGTCGCCGCGGTCTAGGGAGGCGACGGCCGTCGAGCGGCAGCGGGGATCCGCGGGGCCACGGATCGTGGGCCTGCCGGAACCGAATTGCTACAGCCTGGACGAGATTATCGATCGCACCGTCCAGGCTGTAATGGAAAGGCTGGGTGGCGGGCTGGCGGCACTCGAAGCGAAACGTATTGTCGCTAGTGACGCTGAACGGCCTGCGTCTACACATGCCGCGGTGTCGGGTGCTGCAGTGACCCGCTCCACCCCCGTACAGGCTCCAGCCACTAGGGCGGGCCCGGGGCGAGACAGAGCCAAGCGGGGGGGCGGCGCTAACGCGAAGTGCACATCGCAAGCGCCTCAACCCCGTCCGCTGCCCCCGCCCCCGTCTAACATGGACGAGGGGTGGAATGTGGTGGCCCGGCGGGGTGCCAAGAGGGCCGTACCGACAGCTCCCCAAGTGGGTGGAGCACCGGCCACTGCGGCGTCTCAGGCTGCTCCCCAACAGGGTCGAGCGCCTGTGGCCGCCAAGAAAAAGAAGATGAAGAAGAAGTCGTCGCGCGCGCCGCGGTCGGCGGCAGTGGTCTTAACGCTGCTGCCGGCCGCCGAGGCCAAGGGCGTGTCATACGGAGATGTAGTCTCCCGGGCGCGCGCGGCTATTGATCTGGACGAGATCGGGGCGGGGGAGGGCCTCCGCTGTAGGCTGACGGCCAATGGGGCCAAGCTGCTAGAGTGTCCCGGTGCCGATAGTTCCGGCATCGCGGAGCGTCTGGCGGCGAAGCTCCGTACGGTTCTGCCTGAGGAAGAGGTTCGAGTGCACAGGCCGGTCAAAATGGCTGAATTGAGAGTGACCGGCCTGGACGACTGCACCACTAAGGATGAGGTGGCGGCGGCTATTGCCGCCCGGGGCAACTGCGCCCTGGAGAATATAACTGTAGGCGAGCTGCGCCTCGGTTATACCGGGGCTGGCTCTGTTTGGGTGCGTTGCCCGGTAGAGGCGGCCACCTCCTTGGCTGCTCCTCCGCCCGGTAGACCGTCGGGGGAACCGGGCAGGTTGCGCGTGGGCTGGATAGCGGCCCGCGTGCGCCTTCTCGAACCACGCGCCTGGCGGTGCCTCAGATGCTTCAGCACTGGGCACTGCCTCGCTAAATGCGCGAGTGCGGTTGACCGCAGCGGATTGTGCTTTCGCTGTGGTCAGCCTGGACACAAGGCGGCCGTGTGCTCGGCTGCGCCGCATTGCTTATTATGTGCGGCGGCCGGGCGCAAGGCTGACCACCGGGCCGGGGGCAAGGCCTGCCCTCCGGCCAGCAAGAATGCAGAACGAAATCGGCGCCGCCAAGCTAAGCGGCGTCAAAAGAAGCGGTTGGCCGGGGGAGCACCGGCCGGCACTTTGTCCCCCGCTGAGGCGTTCGCGCCCGATAGCGGGGGCAATGGAGTAGGGGAGGGAGCGATGGATGTGAAGAACCGCGATGGTCCCCCCCTGGCGATGGTGACCAGGGGCCCCGGGATCGTCGCGGTACAGTTAGAAAATATCGTCGTGATCGGGGTGTACTTTTCTCCGAATCGGCCTACCGTCGAGTTCGAGCGGTTCCTGGACGGGCTAGAGGTGATCGCTCGTCACCTCGCTCCCCGTTCAGTGATACTGGCGGGGGACTTCAATGCGAAGTCTGTCGCTTGGGGTTCCCCATTCACGGACGCTCGTGGTAGGCTGCTGGAGGAGTGGGCGGTCGCGGTCGGTCTCTGTATCGTTAATAGGGGCTCGATCGCGACTTGCGTGCGGTGGACGGGCGAGTCTATCGTGGACTTGACGTTCGCGAGCTCGCCCGTCGCACGGCGCATCCTCGGCTGGAGAGTGGTGGTGGGGGCGGAATCGTTGTCCGATCACCGGTATATCCGGTACGATCTTTCTGCCCCTGCGCCGACGCCGGCTGCAGGTGCCCGCGGGGTTGACCCCCCGCAGAGTGCATCCCGGTCATTCCCTAGGTGGGCACTGAAGCTCCTGAACAGGGAGCTCCTGGTGGAGGCCTCTATGGTGGCGGCGTGGGCGCCCAAGCGCCCACAACCTGTCGATGTGGAAACCGAGGCCGAATGGTTCCGGGGCGCGATGCACCGCATATGTGATGCCTCGATGCCCCGGGTCGGCTCTCGGCACTCTGGACGGCGACAGTCGCCCTGGTGGTCGCCCGAAATTGCAAGACTGCGTGCGGTCTCCATTCGGCGAGCCGCCGGTGCACTAGACACCGCCGACGCCCTGCGGCTGGCCATCAGGCGGGCCAAGGAGCAGAACATGGAGAGACTCTTGGAGGCGCTCGAAGCGGATCCGTGGGGGAGCCCATACAAATTGGTACGCGGCAAGTTGCGCCCGTGGGCCCCCTCTATGACTGAGCGTCTCCAGCCCCAGCAGCTGCGGGACATCGTTTCGGCGTTGTTCCCGCAGGAGCGGGAGGGTTTTGTCCCTCCCGCTATGGGCGGGCCGTCTGACATCGTCGACGGCCAAGCTCCTGCTGAGGTGCCCCGTATTACGGAGGCAGAGCTCCGGGTGGCCGTTGTCAAGATGACTGCGAAAGACACGGCCCCCGGCCCGGACGGAGTCCACGGCCGGGTATTGGCCTTGGCCCTCGGTGCCCTGGGGGACCGGCTCCTTGAGCTATATAATGGCTGCTTGGAGTCGGGACGGTTTCCGTCGCTCTGGCGGACGGGTAGACTTGTGTTGTTGAGGAAGGAGGGGCGCCCGGTGGATACAGCCGCCGGATATCGTCCTATCGTGCTGCTGGACGAGGCGGGAAAATTGCTGGAACGTATTCTGGCTGCCCGCATCGTTCGGCACCTGGTCGGGGTGGGGCCTGACCTGTCGGCGGAGCAGTACGGCTTCCGGGAGGGCCGTTCGACCGTGGATGCAATTCTTCGCGTGCGGTCCCTCTCGGACGAGGCCGTCTCTCGGGGGTCGGTTCTCGGCCCCCTCTTGTGGAATATCGGGTACGACTGGGTGCTGAGGGGCGCCCTCCTCCCGGGCCTCCGCGTTATTTGTTACGCGGACGACACGTTGGTCGTGGCCCGGGGGGATGATTTTAGGGAGTCTGCCCGTCTCGCCACAGCGGGAGTGGCCCTCGTCGTCGGAAGGATAAGGAGGCTGGGTCTCGACGTGGCGCTCAATAAATCCGAGGCTCTGTGGTTTCACGGGCCGCGGAGGGCGCCACCCGTTGACACCCACATCGTGGTTGGAGGCGTCCGGATAGGGGTCGGGGTGCAGTTGAAGTACCTCGGCCTCGTGTTGGACAGCCGATGGGCCTTTCGTGCTCACTTTGCGGAGCTGGTCCCCCGATTGATGGGGACGGCCGGTTCTTTGAGCCGGCTGCTCCCGAATATTGGGGGACCGGATCAGGTTGTGCGCCGTCTCTACGCGGGGGTGGTGCGATCGATGGCCCTGTACGGTGCACCTGTGTGGGCTGAGCCGCGCAACCGGGCCACTATGGCTCGGTTCCTGCGCCGGCCGCAGCGCACCGTTGCCATCAGGGTCATCCGCGGATATCGCACCGTCTCCTTCGAGGCGGCGTGTGTTTTGGCGGGGACGCCGCCATGGGAGCTGGAGGCGGAGTCGCTCGCTGCCGACTATCGGTGGCGCAGCGAGCTTCGTGCTCGGGGCGTGGCGCGTGTCCCCGAGAGTGAGCTGCGGGCGCGGAAGGCCCATTCTCGGCGGTCCGTGCTCGAGTCGTGGTCGAGGCGATTGGCCAACCCCACGTGGGGGCTACGGACCGTCGAGGCGGTTTACCCGGTCTTTGATGACTGGGTGAATCGTGGCGAGGGACGTCTCACCTTTCGTCTGGTGCAGGTGCTGACCGGGCACGGATGCTTCGGGAAGTACCTGCGCCGGATAGGGGCTGAGCCGACGACGAGGTGTCACCATTGTGGACACGACCTGGACACGGCGGAGCATACGCTCGCTGTCTGCCCCGCTTGGGAGGTGCAGCGCCGTGTCCTGGTCGCAAAGATAGGACCTGACTTGTCGCTGCCTGGCGTCGTGGCGTCGATGCTTGGCAGCGATGAGTCATGGAAGGCTATGCTCGACTTCTGCGAGTGCACCATCTCGCAGAAGGAGGCGGCGGGGCGAGTGAGGGAAAGCTCTCCTCACTACGCAGAAACCCGCCGCCGCCGAGCAGGGGGTCGGGACCGGGGTCGTATCCGTGACCTGGCCCCCTAA
Protein
MMADLTKKLPQEGTTLGGESLRASTSVGACCPTYSGGGKACHDEGGCATGSGKPVELRETRSGKGAPSPAPSSASMSVASLDGRTTEEEEGGLRDPALVIGRAMKTVKSYAATSAKTKAAGRRLREVVSRVSKVLPSVASANDQVVQLMAENLRLRAQLQAQRRPDDKVAATKRVAATRSSPALPRAAGSRTPPASVPAVSPLVSGVPRGPCPSAVAGDPWPVLPPSRAVQRRSRLPSPRSREATAVERQRGSAGPRIVGLPEPNCYSLDEIIDRTVQAVMERLGGGLAALEAKRIVASDAERPASTHAAVSGAAVTRSTPVQAPATRAGPGRDRAKRGGGANAKCTSQAPQPRPLPPPPSNMDEGWNVVARRGAKRAVPTAPQVGGAPATAASQAAPQQGRAPVAAKKKKMKKKSSRAPRSAAVVLTLLPAAEAKGVSYGDVVSRARAAIDLDEIGAGEGLRCRLTANGAKLLECPGADSSGIAERLAAKLRTVLPEEEVRVHRPVKMAELRVTGLDDCTTKDEVAAAIAARGNCALENITVGELRLGYTGAGSVWVRCPVEAATSLAAPPPGRPSGEPGRLRVGWIAARVRLLEPRAWRCLRCFSTGHCLAKCASAVDRSGLCFRCGQPGHKAAVCSAAPHCLLCAAAGRKADHRAGGKACPPASKNAERNRRRQAKRRQKKRLAGGAPAGTLSPAEAFAPDSGGNGVGEGAMDVKNRDGPPLAMVTRGPGIVAVQLENIVVIGVYFSPNRPTVEFERFLDGLEVIARHLAPRSVILAGDFNAKSVAWGSPFTDARGRLLEEWAVAVGLCIVNRGSIATCVRWTGESIVDLTFASSPVARRILGWRVVVGAESLSDHRYIRYDLSAPAPTPAAGARGVDPPQSASRSFPRWALKLLNRELLVEASMVAAWAPKRPQPVDVETEAEWFRGAMHRICDASMPRVGSRHSGRRQSPWWSPEIARLRAVSIRRAAGALDTADALRLAIRRAKEQNMERLLEALEADPWGSPYKLVRGKLRPWAPSMTERLQPQQLRDIVSALFPQEREGFVPPAMGGPSDIVDGQAPAEVPRITEAELRVAVVKMTAKDTAPGPDGVHGRVLALALGALGDRLLELYNGCLESGRFPSLWRTGRLVLLRKEGRPVDTAAGYRPIVLLDEAGKLLERILAARIVRHLVGVGPDLSAEQYGFREGRSTVDAILRVRSLSDEAVSRGSVLGPLLWNIGYDWVLRGALLPGLRVICYADDTLVVARGDDFRESARLATAGVALVVGRIRRLGLDVALNKSEALWFHGPRRAPPVDTHIVVGGVRIGVGVQLKYLGLVLDSRWAFRAHFAELVPRLMGTAGSLSRLLPNIGGPDQVVRRLYAGVVRSMALYGAPVWAEPRNRATMARFLRRPQRTVAIRVIRGYRTVSFEAACVLAGTPPWELEAESLAADYRWRSELRARGVARVPESELRARKAHSRRSVLESWSRRLANPTWGLRTVEAVYPVFDDWVNRGEGRLTFRLVQVLTGHGCFGKYLRRIGAEPTTRCHHCGHDLDTAEHTLAVCPAWEVQRRVLVAKIGPDLSLPGVVASMLGSDESWKAMLDFCECTISQKEAAGRVRESSPHYAETRRRRAGGRDRGRIRDLAP
Summary
Uniprot
EMBL
Proteomes
Interpro
Gene 3D
ProteinModelPortal
PDB
1WDU
E-value=7.27186e-11,
Score=168
Ontologies
GO
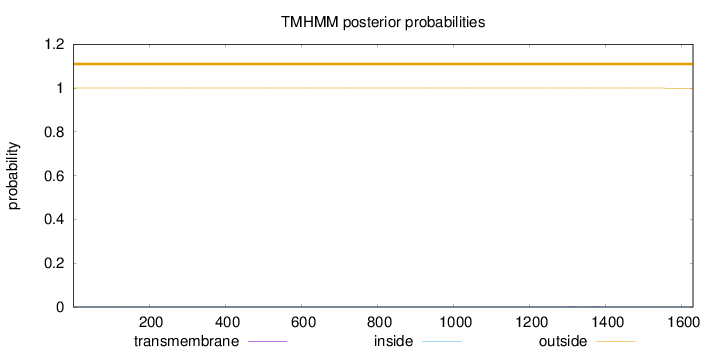
Topology
Length:
1629
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.01144
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00006
outside
1 - 1629
Population Genetic Test Statistics
Pi
151.461093
Theta
109.154343
Tajima's D
0
CLR
186.925241
CSRT
0.367131643417829
Interpretation
Uncertain
