Gene
KWMTBOMO00810
Annotation
reverse_transcriptase_[Lasius_niger]
Location in the cell
Mitochondrial Reliability : 1.711 Nuclear Reliability : 1.722
Sequence
CDS
ATGAGGAAATCGCCAACCGACCCTTGGGCCAAGAGTACAACTGAAAAAGCTGAGCTATTTGCAAATTACTTGTATGAAATATTTACCCCTAACCTATCACCAATGGAGGACTATGAAAAAGAAGTTGATGTCATGATTAATAGTGACCAGCAGCTATCCCTGCCACTCAAATCCGTCACCCCCAGGGAACTTACGCAATGTATACAGTCTCTTATAAATAAAAAAGCTCCTGGTTTTGATTTAATAACTGGTGAGGTACTTAAAAAATTGCCCAAAAAAGTAATAATATTTTTAACAATGTTATTCAATGCTATCCTCAGAGTACATTATTACCCAAAACTCTGGCAAATAGCACAAGTCTGCATGGTTATAAAGCCTGGAAAACCAGCGTCAGATCCAAGTTCATATAGGCCCATAAGTCTTCTACCTGTGTTGTCTAAGGTGTTTGAAACGGTATTTCTAAAAAGACTGAAGCCGCTTCTTGACCAGGAAAAGATTATACCGGATCATCAGTTTGGATTTCGGGCGAAGCATTCTACTGTCGAACAAGTACACAGAGTAGTTCATAAAATTCGACAATCCTTAGAAAAGAAGGAATACTGTTCTGCGGCATTTATTGACATCAAGCAAGCTTTCGATAAAGTCTGGCATAAGGGATTACTATATAAATTGAAAACCCTCCTACCGAATTCTTTTTTCATGCTCCTGAGATCCTACTTAAAACACAGAAAATTCCAAGTCAAATTTGGAGAGGACTGTTCGAAACTGTATAATATTAACGCCTCCGTACCCCAGGGATCAGTGCTTGGTCCGGTCCTGTTTTCTATCTATACTGCAGACCTGCCTCAATCCCAAGATGTTGTAACTGCGACTTACGCTGACGATACAGCTTGTCTGGCAAGTGACATAGATCCACATAATGCATCATTACAACTACAAGTACAGCTTGACAAAGTAGATGCTTGGTTGCAGAGATGGCGATTAAAACCTAGTGTAACTAAGTCAACGCAGGTAACATTCACCCTCCGGAAAGGTATCTGTGGCCCTGTTTACATGGAAGGTAAAGCTTTACCAAATGCCGACTCTGCGAAATACTTGGGATTACACCTAGACAGAAGATTAACCTGGGCGAACCATATAAAAGCAAAAAAACGAGAAGCGAACTTGCAATTTCTCTCTTTGAATTGGTTACTGGGACGCCGGTCTCCACTGTCACTAAGTAACAAACTCCTGGTCTACAAAGCGATTATAAAACCTGTCTGGACTTATGGAATAGAACTGTGGGGTTCCGCTAGCAACTCGAATATCGAGATCTTACAGAGATTCCAGAACATTGCTCTTAGAACTATCTCAAACGCGCATTGGTTTGTAAGAAATAGTGAGATTCACGAATATTTAGAAATGCCGACTGTTCGAGAGGAAGTCAACAAGAACAGCAAGAGATATAAGGAAAGACTAAATGAGCACTCAAATGAATTGGCAAGGTGCCTGCTAGACACAACTGGAGATGTGAAACGTTTAAAAAGATGGCAAATTTTAGACTTAGACCAAAGATACTAA
Protein
MRKSPTDPWAKSTTEKAELFANYLYEIFTPNLSPMEDYEKEVDVMINSDQQLSLPLKSVTPRELTQCIQSLINKKAPGFDLITGEVLKKLPKKVIIFLTMLFNAILRVHYYPKLWQIAQVCMVIKPGKPASDPSSYRPISLLPVLSKVFETVFLKRLKPLLDQEKIIPDHQFGFRAKHSTVEQVHRVVHKIRQSLEKKEYCSAAFIDIKQAFDKVWHKGLLYKLKTLLPNSFFMLLRSYLKHRKFQVKFGEDCSKLYNINASVPQGSVLGPVLFSIYTADLPQSQDVVTATYADDTACLASDIDPHNASLQLQVQLDKVDAWLQRWRLKPSVTKSTQVTFTLRKGICGPVYMEGKALPNADSAKYLGLHLDRRLTWANHIKAKKREANLQFLSLNWLLGRRSPLSLSNKLLVYKAIIKPVWTYGIELWGSASNSNIEILQRFQNIALRTISNAHWFVRNSEIHEYLEMPTVREEVNKNSKRYKERLNEHSNELARCLLDTTGDVKRLKRWQILDLDQRY
Summary
Uniprot
A0A2A4K8E7
A0A2A4K918
A0A0J7K4E0
A0A2J7PVE7
A0A2J7R7D0
A0A1Y1M7E9
+ More
A0A2J7Q9Z1 A0A2J7PSM7 A0A2J7R4U3 A0A2J7QES2 A0A2A4IUM2 A0A2P8Z520 A0A2J7PVX3 A0A1Y1K0N4 A0A2J7QB68 A0A2J7R579 A0A2J7R4B1 A0A2J7QPI2 A0A1Y1LKN6 A0A1L8DIZ6 A0A2J7R8R8 A0A2J7R098 A0A2J7RQ52 A0A2J7PMN4 A0A0J7K811 A0A2J7PDM0 J9LYS0 A0A224X6T4 A0A2J7Q0S7 A0A0J7MYU2 A0A1Y1JXT3 A0A2S2NI07 A0A1B6KCY8 A0A2S2QLY2 A0A2J7PEB1 A0A142LX26 A0A069DVA6 A0A0A9YU04 A0A023F6S4 A0A069DZC5 A0A2S2Q7E7 A0A2J7RD70 A0A069DW01 A0A0A9YJ00 A0A224X6U5 X1WVQ4 A0A2S2NPJ7 A0A1B6EGD8 A0A224X746 A0A023F7D1 A0A1Y1JWM7 A0A224XK92 A0A2S2P0Y0 A0A023F069 A0A224X724 A0A023F700 A0A2H8U0H5 A0A2H8TL70 A0A034VFD2 A0A2S2P6S3 A0A0M4EWX7 A0A224XJI0 A0A2S2N6Q7 A0A2S2N9P4 A0A2S2N814 A0A2S2PNT5 A0A2S2NH02 X1WWD6 A0A2H8TKI9 A0A224XI99 A0A0J7K7J8 A0A0J7KBP6 Q6R811 A0A224X6R6 Q04135 A0A023F754 A0A1Y1KFR3 A0A2S2PH29 A0A069DX78 A0A224XHW8 J9LBC4 A0A0V0G5B6 A0A2S2P3X8 X1XSU7 A0A2S2NPG3 Q961V7 A0A2S2PX53 A0A2S2N629 A0A2S2P698 A0A142LX49 A0A2H8TF37 X1WUG2 A0A023F0I3 A0A2S2QJM7 A0A2H8TV33 A0A2S2NNQ6 A0A0A1WW72 A0A2S2QTZ8 J9M4H5 A0A2S2P7B2
A0A2J7Q9Z1 A0A2J7PSM7 A0A2J7R4U3 A0A2J7QES2 A0A2A4IUM2 A0A2P8Z520 A0A2J7PVX3 A0A1Y1K0N4 A0A2J7QB68 A0A2J7R579 A0A2J7R4B1 A0A2J7QPI2 A0A1Y1LKN6 A0A1L8DIZ6 A0A2J7R8R8 A0A2J7R098 A0A2J7RQ52 A0A2J7PMN4 A0A0J7K811 A0A2J7PDM0 J9LYS0 A0A224X6T4 A0A2J7Q0S7 A0A0J7MYU2 A0A1Y1JXT3 A0A2S2NI07 A0A1B6KCY8 A0A2S2QLY2 A0A2J7PEB1 A0A142LX26 A0A069DVA6 A0A0A9YU04 A0A023F6S4 A0A069DZC5 A0A2S2Q7E7 A0A2J7RD70 A0A069DW01 A0A0A9YJ00 A0A224X6U5 X1WVQ4 A0A2S2NPJ7 A0A1B6EGD8 A0A224X746 A0A023F7D1 A0A1Y1JWM7 A0A224XK92 A0A2S2P0Y0 A0A023F069 A0A224X724 A0A023F700 A0A2H8U0H5 A0A2H8TL70 A0A034VFD2 A0A2S2P6S3 A0A0M4EWX7 A0A224XJI0 A0A2S2N6Q7 A0A2S2N9P4 A0A2S2N814 A0A2S2PNT5 A0A2S2NH02 X1WWD6 A0A2H8TKI9 A0A224XI99 A0A0J7K7J8 A0A0J7KBP6 Q6R811 A0A224X6R6 Q04135 A0A023F754 A0A1Y1KFR3 A0A2S2PH29 A0A069DX78 A0A224XHW8 J9LBC4 A0A0V0G5B6 A0A2S2P3X8 X1XSU7 A0A2S2NPG3 Q961V7 A0A2S2PX53 A0A2S2N629 A0A2S2P698 A0A142LX49 A0A2H8TF37 X1WUG2 A0A023F0I3 A0A2S2QJM7 A0A2H8TV33 A0A2S2NNQ6 A0A0A1WW72 A0A2S2QTZ8 J9M4H5 A0A2S2P7B2
Pubmed
EMBL
NWSH01000032
PCG80497.1
PCG80496.1
LBMM01014395
KMQ85192.1
NEVH01020963
+ More
PNF20308.1 NEVH01006736 PNF36716.1 GEZM01038405 JAV81653.1 NEVH01016340 PNF25410.1 NEVH01021925 PNF19341.1 NEVH01007402 PNF35856.1 NEVH01015305 PNF27092.1 NWSH01007753 PCG62823.1 PYGN01000192 PSN51582.1 NEVH01020940 PNF20481.1 GEZM01097906 JAV54051.1 NEVH01016302 PNF25823.1 NEVH01007393 PNF35987.1 NEVH01007578 PNF35646.1 NEVH01012089 PNF30491.1 GEZM01053022 JAV74219.1 GFDF01007661 JAV06423.1 NEVH01006721 PNF37225.1 NEVH01008277 PNF34262.1 NEVH01001347 PNF42959.1 NEVH01023979 PNF17579.1 LBMM01012078 KMQ86419.1 NEVH01026386 PNF14434.1 ABLF02041861 GFTR01008261 JAW08165.1 NEVH01019963 PNF22185.1 LBMM01013518 KMQ85605.1 GEZM01101509 JAV52620.1 GGMR01003787 MBY16406.1 GEBQ01030670 JAT09307.1 GGMS01009480 MBY78683.1 NEVH01026106 PNF14658.1 KU543673 AMS38352.1 GBGD01000881 JAC88008.1 GBHO01021814 GBHO01010549 JAG21790.1 JAG33055.1 GBBI01001646 JAC17066.1 GBGD01000880 JAC88009.1 GGMS01004444 MBY73647.1 NEVH01005295 PNF38784.1 GBGD01000799 JAC88090.1 GBHO01010547 GBHO01010546 JAG33057.1 JAG33058.1 GFTR01008251 JAW08175.1 ABLF02023279 ABLF02041474 GGMR01006491 MBY19110.1 GEDC01000337 JAS36961.1 GFTR01008151 JAW08275.1 GBBI01001570 JAC17142.1 GEZM01101506 JAV52621.1 GFTR01007997 JAW08429.1 GGMR01010373 MBY22992.1 GBBI01003810 JAC14902.1 GFTR01008169 JAW08257.1 GBBI01001699 JAC17013.1 GFXV01007093 MBW18898.1 GFXV01003078 MBW14883.1 GAKP01017773 JAC41179.1 GGMR01012443 MBY25062.1 CP012527 ALC48130.1 GFTR01008257 JAW08169.1 GGMR01000226 MBY12845.1 GGMR01000857 MBY13476.1 GGMR01000676 MBY13295.1 GGMR01018428 MBY31047.1 GGMR01003864 MBY16483.1 ABLF02042106 GFXV01002840 MBW14645.1 GFTR01008256 JAW08170.1 LBMM01012041 KMQ86458.1 LBMM01010106 KMQ87671.1 AY508487 AAS13459.1 GFTR01008269 JAW08157.1 X17551 CAA35587.1 GBBI01001642 JAC17070.1 GEZM01084969 JAV60363.1 GGMR01016114 MBY28733.1 GBGD01000389 JAC88500.1 GFTR01008254 JAW08172.1 ABLF02036794 GECL01003538 JAP02586.1 GGMR01011511 MBY24130.1 ABLF02028354 ABLF02028357 ABLF02062343 GGMR01006408 MBY19027.1 AY047531 AAK77263.1 GGMS01000737 MBY69940.1 GGMR01000005 MBY12624.1 GGMR01012305 MBY24924.1 KU543685 AMS38375.1 GFXV01000908 MBW12713.1 ABLF02016613 GBBI01003747 JAC14965.1 GGMS01008766 MBY77969.1 GFXV01005866 MBW17671.1 GGMR01006148 MBY18767.1 GBXI01011604 JAD02688.1 GGMS01012005 MBY81208.1 ABLF02017668 ABLF02021394 ABLF02055728 GGMR01012676 MBY25295.1
PNF20308.1 NEVH01006736 PNF36716.1 GEZM01038405 JAV81653.1 NEVH01016340 PNF25410.1 NEVH01021925 PNF19341.1 NEVH01007402 PNF35856.1 NEVH01015305 PNF27092.1 NWSH01007753 PCG62823.1 PYGN01000192 PSN51582.1 NEVH01020940 PNF20481.1 GEZM01097906 JAV54051.1 NEVH01016302 PNF25823.1 NEVH01007393 PNF35987.1 NEVH01007578 PNF35646.1 NEVH01012089 PNF30491.1 GEZM01053022 JAV74219.1 GFDF01007661 JAV06423.1 NEVH01006721 PNF37225.1 NEVH01008277 PNF34262.1 NEVH01001347 PNF42959.1 NEVH01023979 PNF17579.1 LBMM01012078 KMQ86419.1 NEVH01026386 PNF14434.1 ABLF02041861 GFTR01008261 JAW08165.1 NEVH01019963 PNF22185.1 LBMM01013518 KMQ85605.1 GEZM01101509 JAV52620.1 GGMR01003787 MBY16406.1 GEBQ01030670 JAT09307.1 GGMS01009480 MBY78683.1 NEVH01026106 PNF14658.1 KU543673 AMS38352.1 GBGD01000881 JAC88008.1 GBHO01021814 GBHO01010549 JAG21790.1 JAG33055.1 GBBI01001646 JAC17066.1 GBGD01000880 JAC88009.1 GGMS01004444 MBY73647.1 NEVH01005295 PNF38784.1 GBGD01000799 JAC88090.1 GBHO01010547 GBHO01010546 JAG33057.1 JAG33058.1 GFTR01008251 JAW08175.1 ABLF02023279 ABLF02041474 GGMR01006491 MBY19110.1 GEDC01000337 JAS36961.1 GFTR01008151 JAW08275.1 GBBI01001570 JAC17142.1 GEZM01101506 JAV52621.1 GFTR01007997 JAW08429.1 GGMR01010373 MBY22992.1 GBBI01003810 JAC14902.1 GFTR01008169 JAW08257.1 GBBI01001699 JAC17013.1 GFXV01007093 MBW18898.1 GFXV01003078 MBW14883.1 GAKP01017773 JAC41179.1 GGMR01012443 MBY25062.1 CP012527 ALC48130.1 GFTR01008257 JAW08169.1 GGMR01000226 MBY12845.1 GGMR01000857 MBY13476.1 GGMR01000676 MBY13295.1 GGMR01018428 MBY31047.1 GGMR01003864 MBY16483.1 ABLF02042106 GFXV01002840 MBW14645.1 GFTR01008256 JAW08170.1 LBMM01012041 KMQ86458.1 LBMM01010106 KMQ87671.1 AY508487 AAS13459.1 GFTR01008269 JAW08157.1 X17551 CAA35587.1 GBBI01001642 JAC17070.1 GEZM01084969 JAV60363.1 GGMR01016114 MBY28733.1 GBGD01000389 JAC88500.1 GFTR01008254 JAW08172.1 ABLF02036794 GECL01003538 JAP02586.1 GGMR01011511 MBY24130.1 ABLF02028354 ABLF02028357 ABLF02062343 GGMR01006408 MBY19027.1 AY047531 AAK77263.1 GGMS01000737 MBY69940.1 GGMR01000005 MBY12624.1 GGMR01012305 MBY24924.1 KU543685 AMS38375.1 GFXV01000908 MBW12713.1 ABLF02016613 GBBI01003747 JAC14965.1 GGMS01008766 MBY77969.1 GFXV01005866 MBW17671.1 GGMR01006148 MBY18767.1 GBXI01011604 JAD02688.1 GGMS01012005 MBY81208.1 ABLF02017668 ABLF02021394 ABLF02055728 GGMR01012676 MBY25295.1
Proteomes
Interpro
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
A0A2A4K8E7
A0A2A4K918
A0A0J7K4E0
A0A2J7PVE7
A0A2J7R7D0
A0A1Y1M7E9
+ More
A0A2J7Q9Z1 A0A2J7PSM7 A0A2J7R4U3 A0A2J7QES2 A0A2A4IUM2 A0A2P8Z520 A0A2J7PVX3 A0A1Y1K0N4 A0A2J7QB68 A0A2J7R579 A0A2J7R4B1 A0A2J7QPI2 A0A1Y1LKN6 A0A1L8DIZ6 A0A2J7R8R8 A0A2J7R098 A0A2J7RQ52 A0A2J7PMN4 A0A0J7K811 A0A2J7PDM0 J9LYS0 A0A224X6T4 A0A2J7Q0S7 A0A0J7MYU2 A0A1Y1JXT3 A0A2S2NI07 A0A1B6KCY8 A0A2S2QLY2 A0A2J7PEB1 A0A142LX26 A0A069DVA6 A0A0A9YU04 A0A023F6S4 A0A069DZC5 A0A2S2Q7E7 A0A2J7RD70 A0A069DW01 A0A0A9YJ00 A0A224X6U5 X1WVQ4 A0A2S2NPJ7 A0A1B6EGD8 A0A224X746 A0A023F7D1 A0A1Y1JWM7 A0A224XK92 A0A2S2P0Y0 A0A023F069 A0A224X724 A0A023F700 A0A2H8U0H5 A0A2H8TL70 A0A034VFD2 A0A2S2P6S3 A0A0M4EWX7 A0A224XJI0 A0A2S2N6Q7 A0A2S2N9P4 A0A2S2N814 A0A2S2PNT5 A0A2S2NH02 X1WWD6 A0A2H8TKI9 A0A224XI99 A0A0J7K7J8 A0A0J7KBP6 Q6R811 A0A224X6R6 Q04135 A0A023F754 A0A1Y1KFR3 A0A2S2PH29 A0A069DX78 A0A224XHW8 J9LBC4 A0A0V0G5B6 A0A2S2P3X8 X1XSU7 A0A2S2NPG3 Q961V7 A0A2S2PX53 A0A2S2N629 A0A2S2P698 A0A142LX49 A0A2H8TF37 X1WUG2 A0A023F0I3 A0A2S2QJM7 A0A2H8TV33 A0A2S2NNQ6 A0A0A1WW72 A0A2S2QTZ8 J9M4H5 A0A2S2P7B2
A0A2J7Q9Z1 A0A2J7PSM7 A0A2J7R4U3 A0A2J7QES2 A0A2A4IUM2 A0A2P8Z520 A0A2J7PVX3 A0A1Y1K0N4 A0A2J7QB68 A0A2J7R579 A0A2J7R4B1 A0A2J7QPI2 A0A1Y1LKN6 A0A1L8DIZ6 A0A2J7R8R8 A0A2J7R098 A0A2J7RQ52 A0A2J7PMN4 A0A0J7K811 A0A2J7PDM0 J9LYS0 A0A224X6T4 A0A2J7Q0S7 A0A0J7MYU2 A0A1Y1JXT3 A0A2S2NI07 A0A1B6KCY8 A0A2S2QLY2 A0A2J7PEB1 A0A142LX26 A0A069DVA6 A0A0A9YU04 A0A023F6S4 A0A069DZC5 A0A2S2Q7E7 A0A2J7RD70 A0A069DW01 A0A0A9YJ00 A0A224X6U5 X1WVQ4 A0A2S2NPJ7 A0A1B6EGD8 A0A224X746 A0A023F7D1 A0A1Y1JWM7 A0A224XK92 A0A2S2P0Y0 A0A023F069 A0A224X724 A0A023F700 A0A2H8U0H5 A0A2H8TL70 A0A034VFD2 A0A2S2P6S3 A0A0M4EWX7 A0A224XJI0 A0A2S2N6Q7 A0A2S2N9P4 A0A2S2N814 A0A2S2PNT5 A0A2S2NH02 X1WWD6 A0A2H8TKI9 A0A224XI99 A0A0J7K7J8 A0A0J7KBP6 Q6R811 A0A224X6R6 Q04135 A0A023F754 A0A1Y1KFR3 A0A2S2PH29 A0A069DX78 A0A224XHW8 J9LBC4 A0A0V0G5B6 A0A2S2P3X8 X1XSU7 A0A2S2NPG3 Q961V7 A0A2S2PX53 A0A2S2N629 A0A2S2P698 A0A142LX49 A0A2H8TF37 X1WUG2 A0A023F0I3 A0A2S2QJM7 A0A2H8TV33 A0A2S2NNQ6 A0A0A1WW72 A0A2S2QTZ8 J9M4H5 A0A2S2P7B2
Ontologies
KEGG
GO
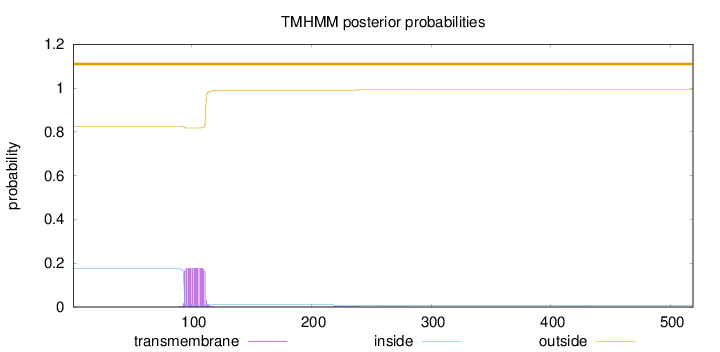
Topology
Length:
519
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
3.35904000000001
Exp number, first 60 AAs:
0.00105
Total prob of N-in:
0.17634
outside
1 - 519
Population Genetic Test Statistics
Pi
168.486964
Theta
123.732333
Tajima's D
0.682273
CLR
137.44788
CSRT
0.56057197140143
Interpretation
Uncertain
