Gene
KWMTBOMO00701
Annotation
PREDICTED:_uncharacterized_protein_LOC103309518_[Acyrthosiphon_pisum]
Location in the cell
Nuclear Reliability : 1.142 PlasmaMembrane Reliability : 1.16
Sequence
CDS
ATGGTAGACACGGGGAGCCAGGCATCGTTTATTACCGAAGCTTGTGTGCAGCGCTTAGGTCTAGCGCGCCGGCATAGCAACGTGCCGGTGTTCGGTATTGGTGACACCGGACCCGTGCATCCAAAAGGTTTAGTTTCGTGTTCCATTGCGCCTAAAGGTCAACAACAACCTGTCATTCCGGTAAATGCGTTAATTTTGCCCGAATTAATTCCAAAAATGCCAAGTGTACCTTTACCGTATACCGGGTGGTCACATTTGAAAAATCTGAAGTTAGCTGATCCGGACTTTCATACGCCCCAGCCTGTTGAAATGTTACTAGGAGCGGATATACTATCCCACGTGTTACTAGGCAACACTATCGTAGGCCCACCGGGCACGCCGATCGCTATGAATAGTGTGTTCGGTTATTTACTTTTAGGCAAATTAGATTTAGATTCTTCTGTTCCAACTTCATTTCAAGTGTGTTTTAGCTCGTTTGACAACGACAACCTTCAGAGATTTTGGGAGCTGGAATCAATACCAGAGAAGCGGAGTTACACTCCAGATGAGGAGTTGTGCGAATCCTTTTTTCAAAAGACGCACACTCGTAATGCGGACGGGCGATACGTGGTCGCCCTCCCGTTTAAGCCCGACGCACCGTCTCTCGGCGAGTCGCGGTCCATTGCGCTAGCTCGGTTTCATAAACTAGAATATCGGTTAGAACGCAACCTCAAGTTGAAGGCTGATTACCACGCTTGTCTCCAGGAATACGTAGACCTCAACCACATGGAGCTCGTCGATGATCAACCTTCAGTTAGTGAAAGTTATTACATTCCGCACCACTGCGTGGTCAAAGAATCGAGTGAGTCGACGCCTACACGTGTGGTGTACGACGCCGGTTGTCGCTCAACTAGTGGGTATTCGCTCAATGACGTTTTACTAACGGGTCCTAAGCTTCACATGGACATAGTGGACGTCCTTCTTAAATTTCGTGTCCATTCAATCGCGCTTACTGCCGACATAAAACAAATGTATCGTAATATACTTGTTCGTGAAAGTGACAAAGATTTTCAGCGTATTGTGTGGCGCACATCACCAGAGCAAACTATTCGTGACTACCGTTTGCGTACAGTCACATTCGGTGTGAAATCGTCACCTTACCTCGCTCTGCGTACAATAAAACAACTCGCTCAAGATGAGGCCGAGCGGTTCAAACTCGCGTCGCCCGTTTTACTTAACGATGTTTTCGTTGACGACGTCGTATCTGGCGAGGACTCTGAAGTCAGCGCACTCGCGCTACAACAAGAGCTGATAGGTATTTGTGGGGCGGCTGGGTTTGAATTGCGCAAATGGCACAGCAACTCACCAGCGTTACTCGCTGCGGTGCTGCCATCAGAGTCTCATGGTGAGCGGCCGGAGAACGTTCTCTTCGCTGAAATGGAGATCGACAAAAAGGTTAAGGTTCTGGGCTTACAGTGGAACCCTAAATCTGATTCATTCAATTTTAAAGTCCAAGTTTCATCGATTAAGTGTACAAAGCGTGTCATTCTTTCAGAAATAGCAAAAATATACGACCCCCTCGGGCTATTATCACCGGTCACATTGTTTGCCAAACATTTGATTCAACTCTTATGGATAGCCAAAGTCGATTGGGATGAGACGCCGCCTGTTGACATAGTAAATTTGTGGTCTTCTTTCGTTGACGAGCTCCCACTCATCTCACAAATCTCATTTCCGCGTTATATATTTAACAATACCAATCCCGAACCGATACAAATTCACGGTTTTGCTGACGCTTCCGAGAAGGGTTTCGGAGCATGTGTATATATTCGTTACAAAGATCAAGAAGATTTCATTCAAACTCACCTGATTATAGCCAAAACTCGCGTCGCACCGTTGAGTGTGCGTCTCACTATTCCTCGCCTCGAACTAATGGCGGCAGTCCTCTTATCTAAATTAATTGAAAAGGTAATGCTCACTTACAGTGGTCGTGTTCGCTTCGACCAGGTCTACGCTTGGTCCGACGCATCCATCGTGCTAGCATGGCTACACTCATCTCCGCACGAATGGAAAACCTTCGTTTCGAACCGTGTCAGTGAGATTTTGCGCCGCATACCCGCGGACCGCTGGCGCCACGTCCCGTCGGCCGACAATCCCGCCGACGCTGCGTCTCGTGGCCTTTTGCCCGCCGCGCTCGTACAGCATGACTTATGGTACCACGGACCCTCGTGGCTACATCTGGGTCAGAACTCCTGGCCGATTCAAAATCCCGTAAGAGACACGTGCGAAGAAAAACGCAACGTATCGTCCGTTCGCTACGCGCTTTTACAACGCTATTGGATTTTGTCAAGCCGTAACATCGTGCGGCAGCGCATACATCGCTGCGTGCAGTGCTTCCGCGCGCGCCCGCCGCACAATCAACCGCGCATGGGACCACTCCCAGCAGTTCGTGTTCGCCCCGCGCGACCGTTTCTAAAGACCGCGGTAGATTTTGCTGGTCCATTTTACGTGCGCGCCAATAAGGACAAGGGCGTCGCCCTTCGTGAAGGCACTGTTGTGGTGGTATGCGACGACCGCTTGCCGCCGTTGCAATGGAGGTTGGCGCGCATCCACGAACTTCATCCCGGCTCGGACAATATCACACGAGTTGTAACCATTAAAATGGGAAATTCCCTTTATAAACGACCCGTGGTTAAAATATGTCCTCTTCCCTTGGAGTAG
Protein
MVDTGSQASFITEACVQRLGLARRHSNVPVFGIGDTGPVHPKGLVSCSIAPKGQQQPVIPVNALILPELIPKMPSVPLPYTGWSHLKNLKLADPDFHTPQPVEMLLGADILSHVLLGNTIVGPPGTPIAMNSVFGYLLLGKLDLDSSVPTSFQVCFSSFDNDNLQRFWELESIPEKRSYTPDEELCESFFQKTHTRNADGRYVVALPFKPDAPSLGESRSIALARFHKLEYRLERNLKLKADYHACLQEYVDLNHMELVDDQPSVSESYYIPHHCVVKESSESTPTRVVYDAGCRSTSGYSLNDVLLTGPKLHMDIVDVLLKFRVHSIALTADIKQMYRNILVRESDKDFQRIVWRTSPEQTIRDYRLRTVTFGVKSSPYLALRTIKQLAQDEAERFKLASPVLLNDVFVDDVVSGEDSEVSALALQQELIGICGAAGFELRKWHSNSPALLAAVLPSESHGERPENVLFAEMEIDKKVKVLGLQWNPKSDSFNFKVQVSSIKCTKRVILSEIAKIYDPLGLLSPVTLFAKHLIQLLWIAKVDWDETPPVDIVNLWSSFVDELPLISQISFPRYIFNNTNPEPIQIHGFADASEKGFGACVYIRYKDQEDFIQTHLIIAKTRVAPLSVRLTIPRLELMAAVLLSKLIEKVMLTYSGRVRFDQVYAWSDASIVLAWLHSSPHEWKTFVSNRVSEILRRIPADRWRHVPSADNPADAASRGLLPAALVQHDLWYHGPSWLHLGQNSWPIQNPVRDTCEEKRNVSSVRYALLQRYWILSSRNIVRQRIHRCVQCFRARPPHNQPRMGPLPAVRVRPARPFLKTAVDFAGPFYVRANKDKGVALREGTVVVVCDDRLPPLQWRLARIHELHPGSDNITRVVTIKMGNSLYKRPVVKICPLPLE
Summary
Uniprot
V5GHM7
A0A1Y1KMX0
X1XU26
A0A2S2QPS5
X1WY95
X1XTX7
+ More
X1WRA3 J9JNP4 J9LPA9 J9JLG6 A0A0J7K7U0 A0A2S2PM34 A0A1S3DIZ2 X1WY64 A0A1S3DJC0 V5GC07 A0A2S2NCY5 A0A1S3DJJ0 A0A1S3DJ34 J9JVD6 A0A182HA33 A0A1B6KIX6 A0A226D417 A0A0J7KDN6 A0A1B6LLT3 A0A0J7ND98 A0A1B6MT91 J9KD60 A0A1U8N896 X1WZY4 X1WYQ8 A0A1B6GGX5 A0A2S2R0Y5 A0A1I8M1N7 A0A2S2P2U4 A0A0A9Z001 A0A1S3DMK8 A0A1S4EIX5 A0A0K8SX96 A0A1Y1LM33 A0A0A9VYL4 A0A0J7N465 A0A034WA88 A0A0A9VSF7 A0A1Y1LIG5 A0A0A9X2P1 A0A0A9ZJJ6 A0A3L8DF49 A0A2S2P1X6 A0A0J7KBZ0 A0A0A9WBL7 A0A3L8D6N4 A0A0J7N314 J9JUM9 A0A2W1BJC9 A0A0A9WAJ3 A0A0A9ZG52 A0A355B931 A0A1Y1LIC8 A0A1B6MUZ9 X1WTX8 A0A1Y1LBZ3 A0A1B6KZV2 A0A0A9W8L8 A0A0N0PAG9 A0A0J7N158 A0A0A9Z275 A0A0A9ZHU0 A0A087T9T6 A0A0A9YYJ5 A0A146KRI6 A0A0A9WP39 A0A3Q0J7N1 A0A146L5E8 A0A0A9WHD6 A0A3Q0J7K9 W8AH72 A0A0J7N189 A0A0J7KBS8 A0A0J7K8M4 A0A3Q0JFQ3 A0A0J7N2B8 J9K0A1 A0A0J7KCI2 A0A023F0J1 A0A0A9WE49 A0A146M9H0 X1WVF9 A0A226E0X4 X1WNM7 A0A146LBU6 A0A226EPI5 A0A2S2NNC8 A0A146L7F7 A0A2S2PLU2 A0A0A9WW84 A0A0J7KMJ0 X1WZF3 A0A023F096
X1WRA3 J9JNP4 J9LPA9 J9JLG6 A0A0J7K7U0 A0A2S2PM34 A0A1S3DIZ2 X1WY64 A0A1S3DJC0 V5GC07 A0A2S2NCY5 A0A1S3DJJ0 A0A1S3DJ34 J9JVD6 A0A182HA33 A0A1B6KIX6 A0A226D417 A0A0J7KDN6 A0A1B6LLT3 A0A0J7ND98 A0A1B6MT91 J9KD60 A0A1U8N896 X1WZY4 X1WYQ8 A0A1B6GGX5 A0A2S2R0Y5 A0A1I8M1N7 A0A2S2P2U4 A0A0A9Z001 A0A1S3DMK8 A0A1S4EIX5 A0A0K8SX96 A0A1Y1LM33 A0A0A9VYL4 A0A0J7N465 A0A034WA88 A0A0A9VSF7 A0A1Y1LIG5 A0A0A9X2P1 A0A0A9ZJJ6 A0A3L8DF49 A0A2S2P1X6 A0A0J7KBZ0 A0A0A9WBL7 A0A3L8D6N4 A0A0J7N314 J9JUM9 A0A2W1BJC9 A0A0A9WAJ3 A0A0A9ZG52 A0A355B931 A0A1Y1LIC8 A0A1B6MUZ9 X1WTX8 A0A1Y1LBZ3 A0A1B6KZV2 A0A0A9W8L8 A0A0N0PAG9 A0A0J7N158 A0A0A9Z275 A0A0A9ZHU0 A0A087T9T6 A0A0A9YYJ5 A0A146KRI6 A0A0A9WP39 A0A3Q0J7N1 A0A146L5E8 A0A0A9WHD6 A0A3Q0J7K9 W8AH72 A0A0J7N189 A0A0J7KBS8 A0A0J7K8M4 A0A3Q0JFQ3 A0A0J7N2B8 J9K0A1 A0A0J7KCI2 A0A023F0J1 A0A0A9WE49 A0A146M9H0 X1WVF9 A0A226E0X4 X1WNM7 A0A146LBU6 A0A226EPI5 A0A2S2NNC8 A0A146L7F7 A0A2S2PLU2 A0A0A9WW84 A0A0J7KMJ0 X1WZF3 A0A023F096
Pubmed
EMBL
GALX01004947
JAB63519.1
GEZM01085145
GEZM01085143
JAV60187.1
ABLF02001506
+ More
ABLF02008029 ABLF02035899 ABLF02037719 ABLF02046327 ABLF02064627 GGMS01010524 MBY79727.1 ABLF02030854 ABLF02067347 ABLF02006304 ABLF02036268 ABLF02066034 ABLF02005684 ABLF02035780 ABLF02030698 ABLF02030705 ABLF02030709 LBMM01012238 KMQ86334.1 GGMR01017890 MBY30509.1 ABLF02012239 ABLF02012241 ABLF02017372 ABLF02042178 ABLF02066436 GALX01000776 JAB67690.1 GGMR01002421 MBY15040.1 ABLF02008594 JXUM01030959 KQ560903 KXJ80414.1 GEBQ01028569 JAT11408.1 LNIX01000036 OXA39969.1 LBMM01009202 KMQ88311.1 GEBQ01015458 JAT24519.1 LBMM01006539 KMQ90520.1 GEBQ01000825 JAT39152.1 ABLF02041863 ABLF02036131 ABLF02041527 ABLF02064049 ABLF02028666 ABLF02066713 GECZ01008071 JAS61698.1 GGMS01014443 MBY83646.1 GGMR01011131 MBY23750.1 GBHO01006408 JAG37196.1 GBRD01007977 JAG57844.1 GEZM01054766 GEZM01054765 JAV73410.1 GBHO01044316 JAF99287.1 LBMM01010417 KMQ87475.1 GAKP01007887 JAC51065.1 GBHO01044317 GBHO01044315 JAF99286.1 JAF99288.1 JAV73413.1 GBHO01028617 JAG14987.1 GBHO01000059 JAG43545.1 QOIP01000009 RLU19024.1 GGMR01010723 MBY23342.1 LBMM01009964 KMQ87756.1 GBHO01039691 GBHO01039690 JAG03913.1 JAG03914.1 QOIP01000012 RLU15919.1 LBMM01011051 KMQ87060.1 ABLF02004035 ABLF02004036 ABLF02004037 ABLF02004038 ABLF02008018 ABLF02008020 ABLF02012635 ABLF02061171 KZ150008 PZC75149.1 GBHO01038800 JAG04804.1 GBHO01000060 JAG43544.1 DNYX01000039 HBK82233.1 GEZM01054789 JAV73399.1 GEBQ01007199 GEBQ01000222 JAT32778.1 JAT39755.1 ABLF02054908 GEZM01060254 JAV71154.1 GEBQ01022991 GEBQ01006185 JAT16986.1 JAT33792.1 GBHO01039838 GBHO01006034 JAG03766.1 JAG37570.1 KQ458473 KPJ05885.1 LBMM01012146 KMQ86385.1 GBHO01007689 JAG35915.1 GBHO01000061 JAG43543.1 KK114194 KFM61875.1 GBHO01012499 GBHO01012498 GBHO01012497 GBHO01007479 GBHO01007451 GBHO01007450 JAG31105.1 JAG31106.1 JAG31107.1 JAG36125.1 JAG36153.1 JAG36154.1 GDHC01019528 JAP99100.1 GBHO01034065 GBHO01034061 JAG09539.1 JAG09543.1 GDHC01016272 JAQ02357.1 GBHO01037676 JAG05928.1 GAMC01021577 GAMC01021573 JAB84982.1 LBMM01012077 KMQ86420.1 LBMM01009888 KMQ87813.1 LBMM01011781 KMQ86634.1 LBMM01011492 KMQ86800.1 ABLF02067195 LBMM01009688 KMQ87939.1 GBBI01004211 JAC14501.1 GBHO01038805 GBHO01038804 GBHO01038801 JAG04799.1 JAG04800.1 JAG04803.1 GDHC01002235 JAQ16394.1 ABLF02026240 ABLF02055027 LNIX01000007 OXA51382.1 ABLF02042806 GDHC01014057 JAQ04572.1 LNIX01000002 OXA59522.1 GGMR01005823 MBY18442.1 GDHC01014268 JAQ04361.1 GGMR01017803 MBY30422.1 GBHO01031928 GBHO01031926 JAG11676.1 JAG11678.1 LBMM01005506 KMQ91466.1 ABLF02041614 GBBI01004276 JAC14436.1
ABLF02008029 ABLF02035899 ABLF02037719 ABLF02046327 ABLF02064627 GGMS01010524 MBY79727.1 ABLF02030854 ABLF02067347 ABLF02006304 ABLF02036268 ABLF02066034 ABLF02005684 ABLF02035780 ABLF02030698 ABLF02030705 ABLF02030709 LBMM01012238 KMQ86334.1 GGMR01017890 MBY30509.1 ABLF02012239 ABLF02012241 ABLF02017372 ABLF02042178 ABLF02066436 GALX01000776 JAB67690.1 GGMR01002421 MBY15040.1 ABLF02008594 JXUM01030959 KQ560903 KXJ80414.1 GEBQ01028569 JAT11408.1 LNIX01000036 OXA39969.1 LBMM01009202 KMQ88311.1 GEBQ01015458 JAT24519.1 LBMM01006539 KMQ90520.1 GEBQ01000825 JAT39152.1 ABLF02041863 ABLF02036131 ABLF02041527 ABLF02064049 ABLF02028666 ABLF02066713 GECZ01008071 JAS61698.1 GGMS01014443 MBY83646.1 GGMR01011131 MBY23750.1 GBHO01006408 JAG37196.1 GBRD01007977 JAG57844.1 GEZM01054766 GEZM01054765 JAV73410.1 GBHO01044316 JAF99287.1 LBMM01010417 KMQ87475.1 GAKP01007887 JAC51065.1 GBHO01044317 GBHO01044315 JAF99286.1 JAF99288.1 JAV73413.1 GBHO01028617 JAG14987.1 GBHO01000059 JAG43545.1 QOIP01000009 RLU19024.1 GGMR01010723 MBY23342.1 LBMM01009964 KMQ87756.1 GBHO01039691 GBHO01039690 JAG03913.1 JAG03914.1 QOIP01000012 RLU15919.1 LBMM01011051 KMQ87060.1 ABLF02004035 ABLF02004036 ABLF02004037 ABLF02004038 ABLF02008018 ABLF02008020 ABLF02012635 ABLF02061171 KZ150008 PZC75149.1 GBHO01038800 JAG04804.1 GBHO01000060 JAG43544.1 DNYX01000039 HBK82233.1 GEZM01054789 JAV73399.1 GEBQ01007199 GEBQ01000222 JAT32778.1 JAT39755.1 ABLF02054908 GEZM01060254 JAV71154.1 GEBQ01022991 GEBQ01006185 JAT16986.1 JAT33792.1 GBHO01039838 GBHO01006034 JAG03766.1 JAG37570.1 KQ458473 KPJ05885.1 LBMM01012146 KMQ86385.1 GBHO01007689 JAG35915.1 GBHO01000061 JAG43543.1 KK114194 KFM61875.1 GBHO01012499 GBHO01012498 GBHO01012497 GBHO01007479 GBHO01007451 GBHO01007450 JAG31105.1 JAG31106.1 JAG31107.1 JAG36125.1 JAG36153.1 JAG36154.1 GDHC01019528 JAP99100.1 GBHO01034065 GBHO01034061 JAG09539.1 JAG09543.1 GDHC01016272 JAQ02357.1 GBHO01037676 JAG05928.1 GAMC01021577 GAMC01021573 JAB84982.1 LBMM01012077 KMQ86420.1 LBMM01009888 KMQ87813.1 LBMM01011781 KMQ86634.1 LBMM01011492 KMQ86800.1 ABLF02067195 LBMM01009688 KMQ87939.1 GBBI01004211 JAC14501.1 GBHO01038805 GBHO01038804 GBHO01038801 JAG04799.1 JAG04800.1 JAG04803.1 GDHC01002235 JAQ16394.1 ABLF02026240 ABLF02055027 LNIX01000007 OXA51382.1 ABLF02042806 GDHC01014057 JAQ04572.1 LNIX01000002 OXA59522.1 GGMR01005823 MBY18442.1 GDHC01014268 JAQ04361.1 GGMR01017803 MBY30422.1 GBHO01031928 GBHO01031926 JAG11676.1 JAG11678.1 LBMM01005506 KMQ91466.1 ABLF02041614 GBBI01004276 JAC14436.1
Proteomes
Pfam
Interpro
IPR041588
Integrase_H2C2
+ More
IPR036397 RNaseH_sf
IPR021109 Peptidase_aspartic_dom_sf
IPR008737 Peptidase_asp_put
IPR040676 DUF5641
IPR012337 RNaseH-like_sf
IPR008042 Retrotrans_Pao
IPR001584 Integrase_cat-core
IPR001878 Znf_CCHC
IPR005312 DUF1759
IPR036875 Znf_CCHC_sf
IPR000477 RT_dom
IPR001969 Aspartic_peptidase_AS
IPR001995 Peptidase_A2_cat
IPR041373 RT_RNaseH
IPR041577 RT_RNaseH_2
IPR000727 T_SNARE_dom
IPR036397 RNaseH_sf
IPR021109 Peptidase_aspartic_dom_sf
IPR008737 Peptidase_asp_put
IPR040676 DUF5641
IPR012337 RNaseH-like_sf
IPR008042 Retrotrans_Pao
IPR001584 Integrase_cat-core
IPR001878 Znf_CCHC
IPR005312 DUF1759
IPR036875 Znf_CCHC_sf
IPR000477 RT_dom
IPR001969 Aspartic_peptidase_AS
IPR001995 Peptidase_A2_cat
IPR041373 RT_RNaseH
IPR041577 RT_RNaseH_2
IPR000727 T_SNARE_dom
Gene 3D
ProteinModelPortal
V5GHM7
A0A1Y1KMX0
X1XU26
A0A2S2QPS5
X1WY95
X1XTX7
+ More
X1WRA3 J9JNP4 J9LPA9 J9JLG6 A0A0J7K7U0 A0A2S2PM34 A0A1S3DIZ2 X1WY64 A0A1S3DJC0 V5GC07 A0A2S2NCY5 A0A1S3DJJ0 A0A1S3DJ34 J9JVD6 A0A182HA33 A0A1B6KIX6 A0A226D417 A0A0J7KDN6 A0A1B6LLT3 A0A0J7ND98 A0A1B6MT91 J9KD60 A0A1U8N896 X1WZY4 X1WYQ8 A0A1B6GGX5 A0A2S2R0Y5 A0A1I8M1N7 A0A2S2P2U4 A0A0A9Z001 A0A1S3DMK8 A0A1S4EIX5 A0A0K8SX96 A0A1Y1LM33 A0A0A9VYL4 A0A0J7N465 A0A034WA88 A0A0A9VSF7 A0A1Y1LIG5 A0A0A9X2P1 A0A0A9ZJJ6 A0A3L8DF49 A0A2S2P1X6 A0A0J7KBZ0 A0A0A9WBL7 A0A3L8D6N4 A0A0J7N314 J9JUM9 A0A2W1BJC9 A0A0A9WAJ3 A0A0A9ZG52 A0A355B931 A0A1Y1LIC8 A0A1B6MUZ9 X1WTX8 A0A1Y1LBZ3 A0A1B6KZV2 A0A0A9W8L8 A0A0N0PAG9 A0A0J7N158 A0A0A9Z275 A0A0A9ZHU0 A0A087T9T6 A0A0A9YYJ5 A0A146KRI6 A0A0A9WP39 A0A3Q0J7N1 A0A146L5E8 A0A0A9WHD6 A0A3Q0J7K9 W8AH72 A0A0J7N189 A0A0J7KBS8 A0A0J7K8M4 A0A3Q0JFQ3 A0A0J7N2B8 J9K0A1 A0A0J7KCI2 A0A023F0J1 A0A0A9WE49 A0A146M9H0 X1WVF9 A0A226E0X4 X1WNM7 A0A146LBU6 A0A226EPI5 A0A2S2NNC8 A0A146L7F7 A0A2S2PLU2 A0A0A9WW84 A0A0J7KMJ0 X1WZF3 A0A023F096
X1WRA3 J9JNP4 J9LPA9 J9JLG6 A0A0J7K7U0 A0A2S2PM34 A0A1S3DIZ2 X1WY64 A0A1S3DJC0 V5GC07 A0A2S2NCY5 A0A1S3DJJ0 A0A1S3DJ34 J9JVD6 A0A182HA33 A0A1B6KIX6 A0A226D417 A0A0J7KDN6 A0A1B6LLT3 A0A0J7ND98 A0A1B6MT91 J9KD60 A0A1U8N896 X1WZY4 X1WYQ8 A0A1B6GGX5 A0A2S2R0Y5 A0A1I8M1N7 A0A2S2P2U4 A0A0A9Z001 A0A1S3DMK8 A0A1S4EIX5 A0A0K8SX96 A0A1Y1LM33 A0A0A9VYL4 A0A0J7N465 A0A034WA88 A0A0A9VSF7 A0A1Y1LIG5 A0A0A9X2P1 A0A0A9ZJJ6 A0A3L8DF49 A0A2S2P1X6 A0A0J7KBZ0 A0A0A9WBL7 A0A3L8D6N4 A0A0J7N314 J9JUM9 A0A2W1BJC9 A0A0A9WAJ3 A0A0A9ZG52 A0A355B931 A0A1Y1LIC8 A0A1B6MUZ9 X1WTX8 A0A1Y1LBZ3 A0A1B6KZV2 A0A0A9W8L8 A0A0N0PAG9 A0A0J7N158 A0A0A9Z275 A0A0A9ZHU0 A0A087T9T6 A0A0A9YYJ5 A0A146KRI6 A0A0A9WP39 A0A3Q0J7N1 A0A146L5E8 A0A0A9WHD6 A0A3Q0J7K9 W8AH72 A0A0J7N189 A0A0J7KBS8 A0A0J7K8M4 A0A3Q0JFQ3 A0A0J7N2B8 J9K0A1 A0A0J7KCI2 A0A023F0J1 A0A0A9WE49 A0A146M9H0 X1WVF9 A0A226E0X4 X1WNM7 A0A146LBU6 A0A226EPI5 A0A2S2NNC8 A0A146L7F7 A0A2S2PLU2 A0A0A9WW84 A0A0J7KMJ0 X1WZF3 A0A023F096
Ontologies
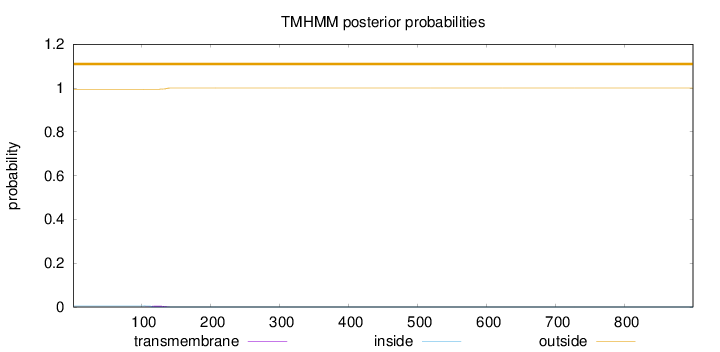
Topology
Length:
899
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.11593
Exp number, first 60 AAs:
0.00026
Total prob of N-in:
0.00505
outside
1 - 899
Population Genetic Test Statistics
Pi
52.287312
Theta
31.718257
Tajima's D
-0.660249
CLR
1.437505
CSRT
0.207839608019599
Interpretation
Uncertain
