Gene
KWMTBOMO00572
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 2.347
Sequence
CDS
ATGAGCAGGTCTTATAAGAGTAAAAGTGTGCGTTGCGATCCGAGTAGCGACCGGCGCCCTAAGTTGTCCGGGAACCGGGGTGTGAAATTGGTTACCGGCCATGGGGGAGCAGGCGACCAGGCACCTCATGGAAATTATGCCACTGGAAGAAGAACAACAAAACTCCAATTACGTAATGATATCCGCATCGGAACGTGGAATGTGAGAGGCCTCATCGATGAAGGAAAGCTCCACGTACTGGATCACGAACTCGAGCGATGTAATACGGTTATTACGGGACTAAGTGAGACGCATTGGAAAGAGAGTGGGCACAGAAACCTGGAGAACCACACCATCTACTTCTCAGGAAATGAATTATCTAGCTTTGCTGGAGTGGGTATCGCAATACCCAAGCTCTGGACTGAGTCGGTCCTGGGCTATAATCCTGTCAGCGACAGGATCATCACCATGAAACTCAGTGCTTCACCATGTGCACTGAATATCATTCAGGTCTACGCCCCTACTAGCGCTGCTACTGAGGAAAAAATTGAGAGCTTCTATAATCAACTCGAGGCTTGTATAAACAACATCCCAAAAAGAGAGATACTGATGATTATCGGCGACTTTAACGCAAAGGTTGGAACCACTGTTTTGGATGTAGGGTTACGAAATATCGTGGGTCACTACGGTCTTGGTTGCAGGAACAGTAGAGGAGAACGGTTGATTCAGTTTGCGGCAGATAATAACTTAACAATAATGAATACAGCCTTCAAACACCACCCCAGAAGACTGTATACTTGGACATCCCCAAACGGAGAGCACCGCAACCAAATCGACTATATACTGATCAAGACAAGATGGAGAAGCTCAATAACCAACGCTCATACACTACCGGGAGCTGATGTCACGTCGGATCACCAGCTACTAATATGCAATATGAGGATCAGACTGCGTAGACCTATAATTAGAAAAACTGTCAAACGCCTAGAAGTCATTGATAGAAACTCGTTTGTCCAGTCTCTTCAGGATATGGAACAGTGCTGGTACGAAAACTGCAGCAAGGAAGTGAGCGTTGATCACCTCTGGCAATCTACCGTGCAATTTATAAAAGATGTCGTGCACGAATCACAGCCCAAAAAGAAATCCAAGAAAAGGCAGCATTGGATGTCTACCGAGACCTGGGACTTAATTGAGAAGCGTAGAGAGCTTAAGGCTGGCGGTGTCAGAATGCAGGAGTTGAACGTCATGAGCTCCGCCATACAGGCTGCTTGCCGTCGTGACCGAAACACAGCGCTACAATCAATATGTGAAGAGGTTGAAAGGCATTCTCAAAATTTTGAAACAAAAGATTTACACATGAAAATCAGACTTATAACCCGTCAATTTAAGCCAAAGACTTGGGCTATTGAGAATGCGTCTGGAGATACTGTCACTGAGATCAAAAAGATTGTTAGTGCTTGGAAAGACTATTGCAAGTCTCTGTTTACTGACACCACAACAAAGATCTCAATGACCTACAGTGAACCCTCGGAACTGGAGCCGATGCCTTTGAAGGACGAGGTGCGAGCAGCAATCAACCACCTTAAACGGAATAAGGCTGTAGGTTTTGATGAAATCCCCATCGAGACAATAAAAGCCATGGGGGAAATCGGAGTGGATATACTGCACATCATCTGTTGTCGAATTTGGGTCACTGGAGTTTGGCCGAGCGATTGGTCACATTCGATATTCATTCCACTTCACAAAAAGGGATCAACGAAGAAGTGCAATAACTACAGACTAATTTCTCTCGTATCCCACGCCAGTAAAGTCATGCTGCACATAATTAACACGAGGCTCCAAGGTTACTTGTCCCGAGAGATCGCCCCCGAGCAAGCAGGATTTGTAAAAGGTCGAGGCACAAGAGAGCAACTACTCGTAATGCGGCAAATTGTAGAAAAGGCCAGAGAATTTAATATTAGCCTATATGTGTGCTTCGTGGATTTCCGAAAGGCTTTTGATACCGTCAAGTGGTGGAAGCTGTGGTTGGTACTGACGGAAATGGGTGTACCGCAACATCTTGTACACACCATCAGACGTCTGTACGAAGATGGCACGGCCGCTGTTCGTGTTGACAGTATTGACTCCGAACGCTTCAGCACACAGGCAGGCGTTAGGCAGGGCTGCATACTATCTCCTCTTTTATTTAATATATACACAGAGTATATAATGCGGATAGTTCTTGATGATTGGGATAAAGGAATATCCGTTGGTGGTCGGAAAATATCAAACTTGAGATACGCCGATGATACTACCCTCTTGGCGTCAACCAGAGACGAAATCGAGGTGCTTCTTCGCCGGCTGGAGACTACAGCGTTGGATTTTGGACTTGCGATCAATCGTGATAAGACCAAAATGATGATCGTAGATCGTGCTAACATCAACCAACCGGAAGTCCAACATATAGCGGGATGCGAGGTAGTTAATAGCTATGTATATCTAGGCTCCACTATCACCAACGCTGGAGGCTGCGAAGATGAGATCCGAAGACGTTGCGCCGTGACAAGATCCTCAGTTGAGAGGCTAACAAAGATTTGGAGAGACCGCAGAATAACCAAAAATACAAAAGTTCGGTTAATGAGATGCCTGGTCTTTCCAATCTTTCTGTATGGGGCTGAGACGTGGACGCTGAGGATGCAAGACCGTCGTAAGATCGACGCACTAGAAATGTGGTGTTGGAGACGTCTCCTCAGAATTCCGTGGACCGCGCACCGTACCAACATTTCGATCCTGAAAGAGCTCAACATCAAGGAGAGACTATCGTCAACAGTCCAATTACGAATCCTCAAATTCTTCGGGCACATCACTCGTAATGAGGACTCCATTGAGCGGTTGGTGGTGCAAGGGAAAGTCGAGGGCAAAAGATCTCGCGGACGATCTCCAACACGATGGACCGACCTCATAAAATCTGTTACTCACTCCAACTTAAATGCTTGCTCTCACTCTGCGAAACATCGTGCCACATGGCGTCGCATCGCGAGAGCATCGGTATCGCAGGATCCGACTGATGCATGA
Protein
MSRSYKSKSVRCDPSSDRRPKLSGNRGVKLVTGHGGAGDQAPHGNYATGRRTTKLQLRNDIRIGTWNVRGLIDEGKLHVLDHELERCNTVITGLSETHWKESGHRNLENHTIYFSGNELSSFAGVGIAIPKLWTESVLGYNPVSDRIITMKLSASPCALNIIQVYAPTSAATEEKIESFYNQLEACINNIPKREILMIIGDFNAKVGTTVLDVGLRNIVGHYGLGCRNSRGERLIQFAADNNLTIMNTAFKHHPRRLYTWTSPNGEHRNQIDYILIKTRWRSSITNAHTLPGADVTSDHQLLICNMRIRLRRPIIRKTVKRLEVIDRNSFVQSLQDMEQCWYENCSKEVSVDHLWQSTVQFIKDVVHESQPKKKSKKRQHWMSTETWDLIEKRRELKAGGVRMQELNVMSSAIQAACRRDRNTALQSICEEVERHSQNFETKDLHMKIRLITRQFKPKTWAIENASGDTVTEIKKIVSAWKDYCKSLFTDTTTKISMTYSEPSELEPMPLKDEVRAAINHLKRNKAVGFDEIPIETIKAMGEIGVDILHIICCRIWVTGVWPSDWSHSIFIPLHKKGSTKKCNNYRLISLVSHASKVMLHIINTRLQGYLSREIAPEQAGFVKGRGTREQLLVMRQIVEKAREFNISLYVCFVDFRKAFDTVKWWKLWLVLTEMGVPQHLVHTIRRLYEDGTAAVRVDSIDSERFSTQAGVRQGCILSPLLFNIYTEYIMRIVLDDWDKGISVGGRKISNLRYADDTTLLASTRDEIEVLLRRLETTALDFGLAINRDKTKMMIVDRANINQPEVQHIAGCEVVNSYVYLGSTITNAGGCEDEIRRRCAVTRSSVERLTKIWRDRRITKNTKVRLMRCLVFPIFLYGAETWTLRMQDRRKIDALEMWCWRRLLRIPWTAHRTNISILKELNIKERLSSTVQLRILKFFGHITRNEDSIERLVVQGKVEGKRSRGRSPTRWTDLIKSVTHSNLNACSHSAKHRATWRRIARASVSQDPTDA
Summary
Uniprot
D7F162
D7F175
D7F167
D7F161
D7F172
D7F168
+ More
D7F163 D5LB39 A0A2W1BNH4 D7F174 A0A0J7KGF7 H2YWZ5 D5LB38 A0A2A4JBK9 A0A2V9X8L4 W5NGM3 A0A076N1B6 W5MYT3 H3J6U9 A0A2B7ZXS2 A0A2J7RP47 A0A2J7PMJ3 A0A2J7QEJ0 A0A2J7RLI7 A0A2J7PQA3 A0A2J7QI56 A0A2J7RGX2 A0A2J7Q3T8 A0A2J7QKB0 A0A2J7PHM9 A0A2J7QH51 A0A2J7Q0C6 A0A2J7QGJ2 A0A2J7R4R4 A0A2J7QUN7 A0A2J7QGX8 A0A2J7Q2Z3 A0A2J7Q5D5 A0A2J7QGT4 A0A2J7Q6A8 A0A1W7RAL1 A0A076N3Z4 A0A2J7Q6P7 A0A2J7PTY0 A0A2J7QR01 A0A2J7PW81 A0A2J7REE7 A0A146LUY8 W5MT28 A0A2S2PMS6 W5MT59 A0A2A4JXP9 A0A2B4RLZ4 A0A2B4R8E3 A0A2J7PSH5 H3ABE5 A0A2P8ZLT6 A0A2P8YWA2 A0A2J7QCG2 A0A2P8Z488 A0A023F0F3 A0A060Q6A2 A0A2J7PT27 A0A2J7PFC5 A0A0U4HM41 A0A2J7Q6T5 A0A2J7QJJ9 A0A2P8Y2S1 A0A2P8ZFZ0 W4XQE1 A0A2P8Z313 H3A4I6 A0A2J7R8P5 A0A2J7QMC5 A0A2J7R901 A0A2J7QUD0 A0A2J7R8G1 A0A2J7RIL8 A0A2J7PX03 A0A0V0G8V1 A0A2S2PT92 Q4QQE8 A0A2J7PSI0 D7F173 A0A2J7Q7C6 A0A023EZ18 A0A0J7KMY2 D7F170 A0A2M4ACI4 C7C207 J9LFP1 A0A1B0DRK3
D7F163 D5LB39 A0A2W1BNH4 D7F174 A0A0J7KGF7 H2YWZ5 D5LB38 A0A2A4JBK9 A0A2V9X8L4 W5NGM3 A0A076N1B6 W5MYT3 H3J6U9 A0A2B7ZXS2 A0A2J7RP47 A0A2J7PMJ3 A0A2J7QEJ0 A0A2J7RLI7 A0A2J7PQA3 A0A2J7QI56 A0A2J7RGX2 A0A2J7Q3T8 A0A2J7QKB0 A0A2J7PHM9 A0A2J7QH51 A0A2J7Q0C6 A0A2J7QGJ2 A0A2J7R4R4 A0A2J7QUN7 A0A2J7QGX8 A0A2J7Q2Z3 A0A2J7Q5D5 A0A2J7QGT4 A0A2J7Q6A8 A0A1W7RAL1 A0A076N3Z4 A0A2J7Q6P7 A0A2J7PTY0 A0A2J7QR01 A0A2J7PW81 A0A2J7REE7 A0A146LUY8 W5MT28 A0A2S2PMS6 W5MT59 A0A2A4JXP9 A0A2B4RLZ4 A0A2B4R8E3 A0A2J7PSH5 H3ABE5 A0A2P8ZLT6 A0A2P8YWA2 A0A2J7QCG2 A0A2P8Z488 A0A023F0F3 A0A060Q6A2 A0A2J7PT27 A0A2J7PFC5 A0A0U4HM41 A0A2J7Q6T5 A0A2J7QJJ9 A0A2P8Y2S1 A0A2P8ZFZ0 W4XQE1 A0A2P8Z313 H3A4I6 A0A2J7R8P5 A0A2J7QMC5 A0A2J7R901 A0A2J7QUD0 A0A2J7R8G1 A0A2J7RIL8 A0A2J7PX03 A0A0V0G8V1 A0A2S2PT92 Q4QQE8 A0A2J7PSI0 D7F173 A0A2J7Q7C6 A0A023EZ18 A0A0J7KMY2 D7F170 A0A2M4ACI4 C7C207 J9LFP1 A0A1B0DRK3
EMBL
FJ265547
ADI61815.1
FJ265560
ADI61828.1
FJ265552
ADI61820.1
+ More
FJ265546 ADI61814.1 FJ265557 ADI61825.1 FJ265553 ADI61821.1 FJ265548 ADI61816.1 GU815090 ADF18553.1 KZ150025 PZC74827.1 FJ265559 ADI61827.1 LBMM01007694 KMQ89503.1 GU815089 ADF18552.1 NWSH01001998 PCG69477.1 QIAP01000240 PYX61602.1 AHAT01008485 KF881087 AIJ27486.1 AHAT01017643 AAGJ04118689 AAGJ04161747 PDHN01000949 PGH37547.1 NEVH01002141 PNF42599.1 NEVH01023985 PNF17546.1 NEVH01015309 PNF27009.1 NEVH01002684 PNF41704.1 NEVH01022640 NEVH01008218 NEVH01005885 PNF18513.1 PNF34430.1 PNF38481.1 NEVH01013959 PNF28275.1 NEVH01003750 PNF40091.1 NEVH01019064 PNF23230.1 NEVH01013275 PNF29009.1 NEVH01025135 PNF15828.1 NEVH01014358 PNF27914.1 NEVH01019971 PNF22037.1 NEVH01018372 NEVH01014371 NEVH01011208 NEVH01002697 NEVH01002552 PNF23678.1 PNF27673.1 PNF31697.1 PNF41592.1 PNF42118.1 NEVH01007402 PNF35815.1 NEVH01023960 NEVH01010501 PNF17732.1 PNF32290.1 NEVH01014359 PNF27813.1 NEVH01019075 PNF22955.1 NEVH01017570 PNF23798.1 PNF27789.1 NEVH01017534 PNF24114.1 GFAH01000222 JAV48167.1 KF881086 AIJ27485.1 NEVH01017460 PNF24248.1 NEVH01021219 PNF19787.1 NEVH01012082 PNF31006.1 NEVH01020937 PNF20581.1 NEVH01005277 PNF39212.1 GDHC01007098 GDHC01001590 JAQ11531.1 JAQ17039.1 AHAT01016400 GGMR01018096 MBY30715.1 AHAT01010273 NWSH01000469 PCG76200.1 LSMT01000478 PFX17357.1 LSMT01001244 PFX12648.1 NEVH01021931 PNF19256.1 AFYH01236276 PYGN01000020 PSN57453.1 PYGN01000324 PSN48495.1 NEVH01016289 PNF26277.1 PYGN01000203 PSN51319.1 GBBI01004261 JAC14451.1 HE799691 CCH14900.1 NEVH01021921 PNF19488.1 NEVH01025668 PNF15041.1 KU311054 ALX81665.1 NEVH01017452 PNF24293.1 NEVH01013553 PNF28765.1 PYGN01001004 PSN38555.1 PYGN01000070 PSN55417.1 AAGJ04045270 PYGN01000219 PSN50892.1 AFYH01196306 NEVH01006721 PNF37211.1 NEVH01013215 PNF29703.1 PNF37310.1 NEVH01011186 PNF32183.1 NEVH01016335 PNF25462.1 PNF37115.1 NEVH01003500 PNF40666.1 NEVH01020865 PNF20863.1 GECL01001585 JAP04539.1 GGMR01020071 MBY32690.1 BN000794 CAJ00236.1 NEVH01021928 PNF19298.1 FJ265558 ADI61826.1 NEVH01017443 PNF24485.1 GBBI01004262 JAC14450.1 LBMM01005351 KMQ91586.1 FJ265555 ADI61823.1 GGFK01005184 MBW38505.1 FN356221 CAX83710.1 ABLF02066124 AJVK01001578
FJ265546 ADI61814.1 FJ265557 ADI61825.1 FJ265553 ADI61821.1 FJ265548 ADI61816.1 GU815090 ADF18553.1 KZ150025 PZC74827.1 FJ265559 ADI61827.1 LBMM01007694 KMQ89503.1 GU815089 ADF18552.1 NWSH01001998 PCG69477.1 QIAP01000240 PYX61602.1 AHAT01008485 KF881087 AIJ27486.1 AHAT01017643 AAGJ04118689 AAGJ04161747 PDHN01000949 PGH37547.1 NEVH01002141 PNF42599.1 NEVH01023985 PNF17546.1 NEVH01015309 PNF27009.1 NEVH01002684 PNF41704.1 NEVH01022640 NEVH01008218 NEVH01005885 PNF18513.1 PNF34430.1 PNF38481.1 NEVH01013959 PNF28275.1 NEVH01003750 PNF40091.1 NEVH01019064 PNF23230.1 NEVH01013275 PNF29009.1 NEVH01025135 PNF15828.1 NEVH01014358 PNF27914.1 NEVH01019971 PNF22037.1 NEVH01018372 NEVH01014371 NEVH01011208 NEVH01002697 NEVH01002552 PNF23678.1 PNF27673.1 PNF31697.1 PNF41592.1 PNF42118.1 NEVH01007402 PNF35815.1 NEVH01023960 NEVH01010501 PNF17732.1 PNF32290.1 NEVH01014359 PNF27813.1 NEVH01019075 PNF22955.1 NEVH01017570 PNF23798.1 PNF27789.1 NEVH01017534 PNF24114.1 GFAH01000222 JAV48167.1 KF881086 AIJ27485.1 NEVH01017460 PNF24248.1 NEVH01021219 PNF19787.1 NEVH01012082 PNF31006.1 NEVH01020937 PNF20581.1 NEVH01005277 PNF39212.1 GDHC01007098 GDHC01001590 JAQ11531.1 JAQ17039.1 AHAT01016400 GGMR01018096 MBY30715.1 AHAT01010273 NWSH01000469 PCG76200.1 LSMT01000478 PFX17357.1 LSMT01001244 PFX12648.1 NEVH01021931 PNF19256.1 AFYH01236276 PYGN01000020 PSN57453.1 PYGN01000324 PSN48495.1 NEVH01016289 PNF26277.1 PYGN01000203 PSN51319.1 GBBI01004261 JAC14451.1 HE799691 CCH14900.1 NEVH01021921 PNF19488.1 NEVH01025668 PNF15041.1 KU311054 ALX81665.1 NEVH01017452 PNF24293.1 NEVH01013553 PNF28765.1 PYGN01001004 PSN38555.1 PYGN01000070 PSN55417.1 AAGJ04045270 PYGN01000219 PSN50892.1 AFYH01196306 NEVH01006721 PNF37211.1 NEVH01013215 PNF29703.1 PNF37310.1 NEVH01011186 PNF32183.1 NEVH01016335 PNF25462.1 PNF37115.1 NEVH01003500 PNF40666.1 NEVH01020865 PNF20863.1 GECL01001585 JAP04539.1 GGMR01020071 MBY32690.1 BN000794 CAJ00236.1 NEVH01021928 PNF19298.1 FJ265558 ADI61826.1 NEVH01017443 PNF24485.1 GBBI01004262 JAC14450.1 LBMM01005351 KMQ91586.1 FJ265555 ADI61823.1 GGFK01005184 MBW38505.1 FN356221 CAX83710.1 ABLF02066124 AJVK01001578
Proteomes
Interpro
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
D7F162
D7F175
D7F167
D7F161
D7F172
D7F168
+ More
D7F163 D5LB39 A0A2W1BNH4 D7F174 A0A0J7KGF7 H2YWZ5 D5LB38 A0A2A4JBK9 A0A2V9X8L4 W5NGM3 A0A076N1B6 W5MYT3 H3J6U9 A0A2B7ZXS2 A0A2J7RP47 A0A2J7PMJ3 A0A2J7QEJ0 A0A2J7RLI7 A0A2J7PQA3 A0A2J7QI56 A0A2J7RGX2 A0A2J7Q3T8 A0A2J7QKB0 A0A2J7PHM9 A0A2J7QH51 A0A2J7Q0C6 A0A2J7QGJ2 A0A2J7R4R4 A0A2J7QUN7 A0A2J7QGX8 A0A2J7Q2Z3 A0A2J7Q5D5 A0A2J7QGT4 A0A2J7Q6A8 A0A1W7RAL1 A0A076N3Z4 A0A2J7Q6P7 A0A2J7PTY0 A0A2J7QR01 A0A2J7PW81 A0A2J7REE7 A0A146LUY8 W5MT28 A0A2S2PMS6 W5MT59 A0A2A4JXP9 A0A2B4RLZ4 A0A2B4R8E3 A0A2J7PSH5 H3ABE5 A0A2P8ZLT6 A0A2P8YWA2 A0A2J7QCG2 A0A2P8Z488 A0A023F0F3 A0A060Q6A2 A0A2J7PT27 A0A2J7PFC5 A0A0U4HM41 A0A2J7Q6T5 A0A2J7QJJ9 A0A2P8Y2S1 A0A2P8ZFZ0 W4XQE1 A0A2P8Z313 H3A4I6 A0A2J7R8P5 A0A2J7QMC5 A0A2J7R901 A0A2J7QUD0 A0A2J7R8G1 A0A2J7RIL8 A0A2J7PX03 A0A0V0G8V1 A0A2S2PT92 Q4QQE8 A0A2J7PSI0 D7F173 A0A2J7Q7C6 A0A023EZ18 A0A0J7KMY2 D7F170 A0A2M4ACI4 C7C207 J9LFP1 A0A1B0DRK3
D7F163 D5LB39 A0A2W1BNH4 D7F174 A0A0J7KGF7 H2YWZ5 D5LB38 A0A2A4JBK9 A0A2V9X8L4 W5NGM3 A0A076N1B6 W5MYT3 H3J6U9 A0A2B7ZXS2 A0A2J7RP47 A0A2J7PMJ3 A0A2J7QEJ0 A0A2J7RLI7 A0A2J7PQA3 A0A2J7QI56 A0A2J7RGX2 A0A2J7Q3T8 A0A2J7QKB0 A0A2J7PHM9 A0A2J7QH51 A0A2J7Q0C6 A0A2J7QGJ2 A0A2J7R4R4 A0A2J7QUN7 A0A2J7QGX8 A0A2J7Q2Z3 A0A2J7Q5D5 A0A2J7QGT4 A0A2J7Q6A8 A0A1W7RAL1 A0A076N3Z4 A0A2J7Q6P7 A0A2J7PTY0 A0A2J7QR01 A0A2J7PW81 A0A2J7REE7 A0A146LUY8 W5MT28 A0A2S2PMS6 W5MT59 A0A2A4JXP9 A0A2B4RLZ4 A0A2B4R8E3 A0A2J7PSH5 H3ABE5 A0A2P8ZLT6 A0A2P8YWA2 A0A2J7QCG2 A0A2P8Z488 A0A023F0F3 A0A060Q6A2 A0A2J7PT27 A0A2J7PFC5 A0A0U4HM41 A0A2J7Q6T5 A0A2J7QJJ9 A0A2P8Y2S1 A0A2P8ZFZ0 W4XQE1 A0A2P8Z313 H3A4I6 A0A2J7R8P5 A0A2J7QMC5 A0A2J7R901 A0A2J7QUD0 A0A2J7R8G1 A0A2J7RIL8 A0A2J7PX03 A0A0V0G8V1 A0A2S2PT92 Q4QQE8 A0A2J7PSI0 D7F173 A0A2J7Q7C6 A0A023EZ18 A0A0J7KMY2 D7F170 A0A2M4ACI4 C7C207 J9LFP1 A0A1B0DRK3
PDB
5HHL
E-value=0.000866784,
Score=105
Ontologies
PANTHER
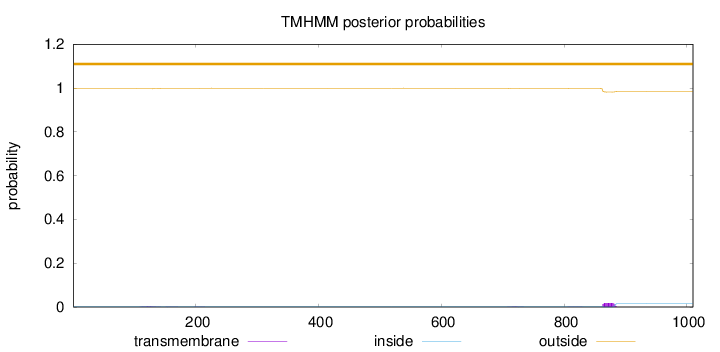
Topology
Length:
1010
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.47318
Exp number, first 60 AAs:
0.00018
Total prob of N-in:
0.00374
outside
1 - 1010
Population Genetic Test Statistics
Pi
1.025671
Theta
1.94266
Tajima's D
-1.686282
CLR
0
CSRT
0.0264486775661217
Interpretation
Uncertain
